[English] 日本語
 Yorodumi
Yorodumi- PDB-1dd6: IMP-1 METALLO BETA-LACTAMASE FROM PSEUDOMONAS AERUGINOSA IN COMPL... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 1dd6 | ||||||
|---|---|---|---|---|---|---|---|
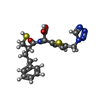
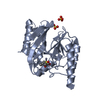
| Title | IMP-1 METALLO BETA-LACTAMASE FROM PSEUDOMONAS AERUGINOSA IN COMPLEX WITH A MERCAPTOCARBOXYLATE INHIBITOR | ||||||
 Components Components | IMP-1 METALLO BETA-LACTAMASE | ||||||
 Keywords Keywords | HYDROLASE / METALLO BETA-LACTAMASE INHIBITOR / MERCAPTOCARBOXYLATE INHIBITOR / IMP-1 METALLO BETA-LACTAMASE | ||||||
| Function / homology |  Function and homology information Function and homology informationantibiotic catabolic process / beta-lactamase activity / beta-lactamase / periplasmic space / response to antibiotic / zinc ion binding Similarity search - Function | ||||||
| Biological species |  | ||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / Resolution: 2 Å SYNCHROTRON / Resolution: 2 Å | ||||||
 Authors Authors | Concha, N.O. / Janson, C.A. / Rowling, P. / Pearson, S. / Cheever, C.A. / Clarke, B.P. / Lewis, C. / Galleni, M. / Frere, J.M. / Payne, D.J. ...Concha, N.O. / Janson, C.A. / Rowling, P. / Pearson, S. / Cheever, C.A. / Clarke, B.P. / Lewis, C. / Galleni, M. / Frere, J.M. / Payne, D.J. / Bateson, J.H. / Abdel-Meguid, S.S. | ||||||
 Citation Citation |  Journal: Biochemistry / Year: 2000 Journal: Biochemistry / Year: 2000Title: Crystal structure of the IMP-1 metallo beta-lactamase from Pseudomonas aeruginosa and its complex with a mercaptocarboxylate inhibitor: binding determinants of a potent, broad-spectrum inhibitor. Authors: Concha, N.O. / Janson, C.A. / Rowling, P. / Pearson, S. / Cheever, C.A. / Clarke, B.P. / Lewis, C. / Galleni, M. / Frere, J.M. / Payne, D.J. / Bateson, J.H. / Abdel-Meguid, S.S. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  1dd6.cif.gz 1dd6.cif.gz | 107 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb1dd6.ent.gz pdb1dd6.ent.gz | 82.5 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  1dd6.json.gz 1dd6.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Summary document |  1dd6_validation.pdf.gz 1dd6_validation.pdf.gz | 549.7 KB | Display |  wwPDB validaton report wwPDB validaton report |
|---|---|---|---|---|
| Full document |  1dd6_full_validation.pdf.gz 1dd6_full_validation.pdf.gz | 562.3 KB | Display | |
| Data in XML |  1dd6_validation.xml.gz 1dd6_validation.xml.gz | 13 KB | Display | |
| Data in CIF |  1dd6_validation.cif.gz 1dd6_validation.cif.gz | 20 KB | Display | |
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/dd/1dd6 https://data.pdbj.org/pub/pdb/validation_reports/dd/1dd6 ftp://data.pdbj.org/pub/pdb/validation_reports/dd/1dd6 ftp://data.pdbj.org/pub/pdb/validation_reports/dd/1dd6 | HTTPS FTP |
-Related structure data
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 | 
| ||||||||
| 2 | 
| ||||||||
| Unit cell |
|
- Components
Components
| #1: Protein | Mass: 25146.676 Da / Num. of mol.: 2 Source method: isolated from a genetically manipulated source Source: (gene. exp.)   #2: Chemical | ChemComp-SO4 / | #3: Chemical | ChemComp-ZN / #4: Chemical | #5: Water | ChemComp-HOH / | |
|---|
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 2 X-RAY DIFFRACTION / Number of used crystals: 2 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.55 Å3/Da / Density % sol: 51.76 % | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Crystal grow | Temperature: 293 K / Method: vapor diffusion, sitting drop / pH: 6.5 Details: SITTING DROPS WERE PREPARED BY MIXING EQUAL VOLUMES OF PROTEIN WHICH WAS 14 MG/ ML IN 20MM HEPES, PH 7.5 AND TO WHICH AN EXCESS OF SOLID INHIBITOR HAD BEEN ADDED AND RESERVOIR SOLUTION (30% ...Details: SITTING DROPS WERE PREPARED BY MIXING EQUAL VOLUMES OF PROTEIN WHICH WAS 14 MG/ ML IN 20MM HEPES, PH 7.5 AND TO WHICH AN EXCESS OF SOLID INHIBITOR HAD BEEN ADDED AND RESERVOIR SOLUTION (30% PEG 2000 MONOMETHYLETHER, 0.1M SODIUM ACETATE, PH 5.0 AND 0.2M AMMONIUM SULFATE). THIS MIXTURE WAS INCUBATED OVERNIGHT AT 4C AND CENTRIFUGED TO REMOVE PRECIPITATE BEFORE SETTING UP CRYSTALLIZATION DROPS. CO-CRYSTALS WERE GROWN FROM 6 ML SITTING DROPS OF THE PROTEIN-RESERVOIR SOLUTION AND 0.3 ML OF THE RESERVOIR SOLUTION AT EITHER ROOM TEMPERATURE OR 4C, pH 6.5, VAPOR DIFFUSION, SITTING DROP, temperature 293K Temp details: 293-295 | ||||||||||||||||||||||||||||||||||||
| Crystal grow | *PLUS pH: 7.5 | ||||||||||||||||||||||||||||||||||||
| Components of the solutions | *PLUS
|
-Data collection
| Diffraction |
| ||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Diffraction source |
| ||||||||||||||||||
| Detector |
| ||||||||||||||||||
| Radiation |
| ||||||||||||||||||
| Radiation wavelength |
| ||||||||||||||||||
| Reflection | Resolution: 2→20 Å / Num. all: 35656 / Num. obs: 545270 / Observed criterion σ(F): 0 / Observed criterion σ(I): 2 / Redundancy: 16 % / Biso Wilson estimate: 25 Å2 / Rmerge(I) obs: 0.103 / Net I/σ(I): 21 | ||||||||||||||||||
| Reflection shell | Resolution: 2→2.07 Å / Redundancy: 7 % / Rmerge(I) obs: 0.314 / % possible all: 97.6 | ||||||||||||||||||
| Reflection | *PLUS Num. obs: 34187 / % possible obs: 95.9 % / Num. measured all: 545270 | ||||||||||||||||||
| Reflection shell | *PLUS % possible obs: 97.6 % |
- Processing
Processing
| Software |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Resolution: 2→20 Å / σ(F): 0 / σ(I): 0 / Stereochemistry target values: ENGH & HUBER
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 2→20 Å
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Software | *PLUS Name: CNS / Classification: refinement | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement | *PLUS σ(F): 2 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Solvent computation | *PLUS | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | *PLUS | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints | *PLUS Type: c_angle_deg / Dev ideal: 2 |
 Movie
Movie Controller
Controller












 PDBj
PDBj