+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Neck of phage 812 after tail contraction (C6) | |||||||||
 Map data Map data | Neck of phage 812 after tail contraction (C6) | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | phage / neck / connector / VIRUS | |||||||||
| Function / homology | Major tail sheath protein / Neck protein / Capsid protein / Baseplate hub assembly protein / Capsid protein Function and homology information Function and homology information | |||||||||
| Biological species |  Staphylococcus phage 812 (virus) Staphylococcus phage 812 (virus) | |||||||||
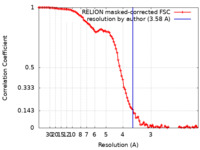
| Method | single particle reconstruction / cryo EM / Resolution: 3.58 Å | |||||||||
 Authors Authors | Cienikova Z / Siborova M / Fuzik T / Plevka P | |||||||||
| Funding support |  Czech Republic, 1 items Czech Republic, 1 items
| |||||||||
 Citation Citation |  Journal: To Be Published Journal: To Be PublishedTitle: Genome anchoring, retention, and release by neck proteins of Herelleviridae phage 812 Authors: Cienikova Z / Novacek J / Siborova M / Popelarova B / Fuzik T / Benesik M / Bardy P / Plevka P | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_18048.map.gz emd_18048.map.gz | 35.9 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-18048-v30.xml emd-18048-v30.xml emd-18048.xml emd-18048.xml | 23.4 KB 23.4 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_18048_fsc.xml emd_18048_fsc.xml | 10.3 KB | Display |  FSC data file FSC data file |
| Images |  emd_18048.png emd_18048.png | 95.9 KB | ||
| Masks |  emd_18048_msk_1.map emd_18048_msk_1.map | 91.1 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-18048.cif.gz emd-18048.cif.gz | 7.4 KB | ||
| Others |  emd_18048_half_map_1.map.gz emd_18048_half_map_1.map.gz emd_18048_half_map_2.map.gz emd_18048_half_map_2.map.gz | 69.7 MB 69.6 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-18048 http://ftp.pdbj.org/pub/emdb/structures/EMD-18048 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-18048 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-18048 | HTTPS FTP |
-Validation report
| Summary document |  emd_18048_validation.pdf.gz emd_18048_validation.pdf.gz | 1.1 MB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_18048_full_validation.pdf.gz emd_18048_full_validation.pdf.gz | 1.1 MB | Display | |
| Data in XML |  emd_18048_validation.xml.gz emd_18048_validation.xml.gz | 17.5 KB | Display | |
| Data in CIF |  emd_18048_validation.cif.gz emd_18048_validation.cif.gz | 23.1 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-18048 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-18048 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-18048 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-18048 | HTTPS FTP |
-Related structure data
| Related structure data |  8q01MC  8q1iC  8q7dC  8qekC  8qemC  8qgrC  8qjeC  8qkhC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_18048.map.gz / Format: CCP4 / Size: 91.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_18048.map.gz / Format: CCP4 / Size: 91.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Neck of phage 812 after tail contraction (C6) | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.057 Å | ||||||||||||||||||||||||||||||||||||
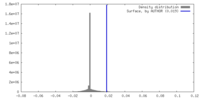
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_18048_msk_1.map emd_18048_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
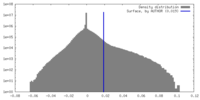
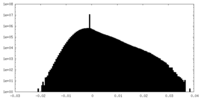
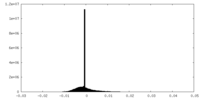
| Density Histograms |
-Half map: half-map 1 for neck of phage 812 after tail contraction (C6)
| File | emd_18048_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | half-map 1 for neck of phage 812 after tail contraction (C6) | ||||||||||||
| Projections & Slices |
| ||||||||||||
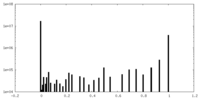
| Density Histograms |
-Half map: half-map 2 for neck of phage 812 after...
| File | emd_18048_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | half-map 2 for neck of phage 812 after tail contraction (C6), box 288 px | ||||||||||||
| Projections & Slices |
| ||||||||||||
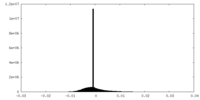
| Density Histograms |
- Sample components
Sample components
-Entire : Staphylococcus phage 812
| Entire | Name:  Staphylococcus phage 812 (virus) Staphylococcus phage 812 (virus) |
|---|---|
| Components |
|
-Supramolecule #1: Staphylococcus phage 812
| Supramolecule | Name: Staphylococcus phage 812 / type: virus / ID: 1 / Parent: 0 / Macromolecule list: #1-#5 / NCBI-ID: 307898 / Sci species name: Staphylococcus phage 812 / Sci species strain: K1-420 / Virus type: VIRION / Virus isolate: STRAIN / Virus enveloped: No / Virus empty: Yes |
|---|---|
| Host (natural) | Organism:  |
| Virus shell | Shell ID: 1 / Diameter: 1100.0 Å |
-Macromolecule #1: Tail tube protein
| Macromolecule | Name: Tail tube protein / type: protein_or_peptide / ID: 1 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Staphylococcus phage 812 (virus) / Strain: K1-420 Staphylococcus phage 812 (virus) / Strain: K1-420 |
| Molecular weight | Theoretical: 15.94297 KDa |
| Sequence | String: MASEAKQTVH TGNTVLLMIK GKPVGRAQSA SGQREYGTTG VYEIGSIMPQ EHVYLRYEGT ITVERLRMKK ENFADLGYAS LGEEILKKD IIDILVVDNL TKQVIISYHG CSANNYNETW QTNEIVTEEI EFSYLTASDK ART UniProtKB: Capsid protein |
-Macromolecule #2: Tail terminator protein
| Macromolecule | Name: Tail terminator protein / type: protein_or_peptide / ID: 2 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Staphylococcus phage 812 (virus) / Strain: K1-420 Staphylococcus phage 812 (virus) / Strain: K1-420 |
| Molecular weight | Theoretical: 31.79968 KDa |
| Sequence | String: MAITSVDSYL LSEIKPRLNT VLENCYIIDE VLKDFDYQTR ESFKEAFCGK NAQHEVTVGF NFPKFKNNYE AHYLIQLGQG QETKNSLGS IQSSYFEATG DTLVESSTAI REDDKLVFTV SKPIGELIKV EDIEFAKYDN LQVEGNKVSF KYQTNEDYEN Y NANIIFTE ...String: MAITSVDSYL LSEIKPRLNT VLENCYIIDE VLKDFDYQTR ESFKEAFCGK NAQHEVTVGF NFPKFKNNYE AHYLIQLGQG QETKNSLGS IQSSYFEATG DTLVESSTAI REDDKLVFTV SKPIGELIKV EDIEFAKYDN LQVEGNKVSF KYQTNEDYEN Y NANIIFTE KKNDSKGLVK GFTVEEQVTV VGLSFNVDVA RCLDAVLKMI LISMRDSIEE QQTFQLQNLS FGDIAPIIED GD SMIFGRP TIIKYTSSLD LDYTITQDIN KLTFKERKDW K UniProtKB: Baseplate hub assembly protein |
-Macromolecule #3: Tail sheath protein
| Macromolecule | Name: Tail sheath protein / type: protein_or_peptide / ID: 3 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Staphylococcus phage 812 (virus) / Strain: K1-420 Staphylococcus phage 812 (virus) / Strain: K1-420 |
| Molecular weight | Theoretical: 64.559008 KDa |
| Sequence | String: MAVEPFPRRP ITRPHASIEV DTSGIGGSAG SSEKVFCLIG QAEGGEPNTV YELRNYSQAK RLFRSGELLD AIELAWGSNP NYTAGRILA MRIEDAKPAS AEIGGLKITS KIYGNVANNI QVGLEKNTLS DSLRLRVIFQ DDRFNEVYDN IGNIFTIKYK G EEANATFS ...String: MAVEPFPRRP ITRPHASIEV DTSGIGGSAG SSEKVFCLIG QAEGGEPNTV YELRNYSQAK RLFRSGELLD AIELAWGSNP NYTAGRILA MRIEDAKPAS AEIGGLKITS KIYGNVANNI QVGLEKNTLS DSLRLRVIFQ DDRFNEVYDN IGNIFTIKYK G EEANATFS VEHDEETQKA SRLVLKVGDQ EVKSYDLTGG AYDYTNAIIT DINQLPDFEA KLSPFGDKNL ESSKLDKIEN AN IKDKAVY VKAVFGDLEK QTAYNGIVSF EQLNAEGEVP SNVEVEAGEE SATVTATSPI KTIEPFELTK LKGGTNGEPP ATW ADKLDK FAHEGGYYIV PLSSKQSVHA EVASFVKERS DAGEPMRAIV GGGFNESKEQ LFGRQASLSN PRVSLVANSG TFVM DDGRK NHVPAYMVAV ALGGLASGLE IGESITFKPL RVSSLDQIYE SIDLDELNEN GIISIEFVRN RTNTFFRIVD DVTTF NDKS DPVKAEMAVG EANDFLVSEL KVQLEDQFIG TRTINTSASI IKDFIQSYLG RKKRDNEIQD FPAEDVQVIV EGNEAR ISM TVYPIRSFKK ISVSLVYKQQ TLQA UniProtKB: Major tail sheath protein |
-Macromolecule #4: Capsid protein
| Macromolecule | Name: Capsid protein / type: protein_or_peptide / ID: 4 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Staphylococcus phage 812 (virus) / Strain: K1-420 Staphylococcus phage 812 (virus) / Strain: K1-420 |
| Molecular weight | Theoretical: 33.757332 KDa |
| Sequence | String: MEKPYMIGAN SNPNVINKST TYTTTTQADE QDKPKYTTRL EFDTIDMIRF INDRGIKVLW EEAYFCPCLN PDTGHPRVDC PRCHGKGIA YLPPKETIMA IQSQEKGTNQ LDIGILDTGT AIGTTQLEKR ISYRDRFTVP EVLMPQQMIY FVNKDRIKKG I PLYYDVKE ...String: MEKPYMIGAN SNPNVINKST TYTTTTQADE QDKPKYTTRL EFDTIDMIRF INDRGIKVLW EEAYFCPCLN PDTGHPRVDC PRCHGKGIA YLPPKETIMA IQSQEKGTNQ LDIGILDTGT AIGTTQLEKR ISYRDRFTVP EVLMPQQMIY FVNKDRIKKG I PLYYDVKE ITYIATQDGT VYEEDYEIKN NRLYLNEKYE NHTVTLKILM TLRYVVSDIL KESRYQYTKF NQPKSKFENL PQ KLLLKRE DVIVLQDPYK VNDGIEEDLE IQVDDPKASA SNPSNLGGFF GGAFK UniProtKB: Capsid protein |
-Macromolecule #5: Adaptor protein
| Macromolecule | Name: Adaptor protein / type: protein_or_peptide / ID: 5 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Staphylococcus phage 812 (virus) / Strain: K1-420 Staphylococcus phage 812 (virus) / Strain: K1-420 |
| Molecular weight | Theoretical: 34.191703 KDa |
| Sequence | String: MVNSMFGGDL DPYEKSLNYE YPYHPSGNPK HIDVSEIDNL TLADYGWSPD AVKAYMFGIV VQNPDTGQPM GDEFYNHILE RAVGKAERA LDISILPDTQ HEMRDYHETE FNSYMFVHAY RKPILQVENL QLQFNGRPIY KYPANWWKVE HLAGHVQLFP T ALMQTGQS ...String: MVNSMFGGDL DPYEKSLNYE YPYHPSGNPK HIDVSEIDNL TLADYGWSPD AVKAYMFGIV VQNPDTGQPM GDEFYNHILE RAVGKAERA LDISILPDTQ HEMRDYHETE FNSYMFVHAY RKPILQVENL QLQFNGRPIY KYPANWWKVE HLAGHVQLFP T ALMQTGQS MSYDAVFNGY PQLAGVYPPS GATFAPQMIR LEYVSGMLPR KKAGRNKPWE MPPELEQLVI KYALKEIYQV WG NLIIGAG IANKTLEVDG ITETIGTTQS AMYGGASAQI LQINEDIKEL LDGLRAYFGY NMIGL UniProtKB: Neck protein |
-Macromolecule #6: ZINC ION
| Macromolecule | Name: ZINC ION / type: ligand / ID: 6 / Number of copies: 1 / Formula: ZN |
|---|---|
| Molecular weight | Theoretical: 65.409 Da |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 8 Component:
| ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Grid | Model: Quantifoil R2/1 / Material: COPPER / Mesh: 200 / Support film - Material: CARBON / Support film - topology: HOLEY / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 30 sec. | ||||||||||||
| Vitrification | Cryogen name: ETHANE-PROPANE / Chamber humidity: 100 % / Instrument: FEI VITROBOT MARK IV | ||||||||||||
| Details | Phage particle after tail contraction and genome ejection |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Specialist optics | Energy filter - Slit width: 20 eV |
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Detector mode: SUPER-RESOLUTION / Number real images: 15371 / Average exposure time: 7.0 sec. / Average electron dose: 42.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 50.0 µm / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 2.4 µm / Nominal defocus min: 1.0 µm / Nominal magnification: 130000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Initial model | Chain - Source name: AlphaFold / Chain - Initial model type: in silico model |
|---|---|
| Refinement | Space: REAL / Protocol: AB INITIO MODEL / Target criteria: Cross-correlation coefficient |
| Output model |  PDB-8q01: |
 Movie
Movie Controller
Controller















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)