[English] 日本語
 Yorodumi
Yorodumi- EMDB-12745: Yeast RNA polymerase II transcription pre-initiation complex with... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-12745 | ||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Yeast RNA polymerase II transcription pre-initiation complex with open promoter DNA | ||||||||||||||||||
 Map data Map data | |||||||||||||||||||
 Sample Sample |
| ||||||||||||||||||
 Keywords Keywords | Pre-initiation complex / TRANSCRIPTION | ||||||||||||||||||
| Function / homology |  Function and homology information Function and homology informationregulation of mitotic recombination / RNA polymerase II promoter clearance / RNA polymerase II complex recruiting activity / TFIIA-class transcription factor complex binding / transcription factor TFIIIB complex / RNA polymerase III preinitiation complex assembly / RNA polymerase III transcription regulatory region sequence-specific DNA binding / phosphatidylinositol-5-phosphate binding / positive regulation of mitotic recombination / regulation of transcription by RNA polymerase III ...regulation of mitotic recombination / RNA polymerase II promoter clearance / RNA polymerase II complex recruiting activity / TFIIA-class transcription factor complex binding / transcription factor TFIIIB complex / RNA polymerase III preinitiation complex assembly / RNA polymerase III transcription regulatory region sequence-specific DNA binding / phosphatidylinositol-5-phosphate binding / positive regulation of mitotic recombination / regulation of transcription by RNA polymerase III / nucleotide-excision repair factor 3 complex / RNA polymerase I general transcription initiation factor binding / transcription factor TFIIE complex / DNA translocase activity / nucleotide-excision repair, preincision complex assembly / transcription factor TFIIK complex / transcription open complex formation at RNA polymerase II promoter / TFIIF-class transcription factor complex binding / transcriptional start site selection at RNA polymerase II promoter / RPB4-RPB7 complex / transcription factor TFIIF complex / DNA 5'-3' helicase / transcription factor TFIIA complex / phosphatidylinositol-3-phosphate binding / RNA polymerase I preinitiation complex assembly / transcription factor TFIIH holo complex / transcription factor TFIIH core complex / cyclin-dependent protein serine/threonine kinase activator activity / DNA 3'-5' helicase / transcription preinitiation complex / DNA binding, bending / poly(A)+ mRNA export from nucleus / RNA Polymerase I Transcription Initiation / DNA duplex unwinding / Processing of Capped Intron-Containing Pre-mRNA / nuclear-transcribed mRNA catabolic process, deadenylation-dependent decay / RNA Polymerase III Transcription Initiation From Type 2 Promoter / 3'-5' DNA helicase activity / RNA Pol II CTD phosphorylation and interaction with CE / Formation of the Early Elongation Complex / mRNA Capping / RNA polymerase II transcribes snRNA genes / TP53 Regulates Transcription of DNA Repair Genes / termination of RNA polymerase II transcription / transcription factor TFIID complex / RNA Polymerase II Promoter Escape / RNA Polymerase II Transcription Pre-Initiation And Promoter Opening / RNA Polymerase II Transcription Initiation / RNA Polymerase II Transcription Initiation And Promoter Clearance / RNA polymerase II general transcription initiation factor activity / RNA Polymerase II Pre-transcription Events / RNA-templated transcription / termination of RNA polymerase III transcription / Formation of TC-NER Pre-Incision Complex / transcription initiation at RNA polymerase III promoter / maintenance of transcriptional fidelity during transcription elongation by RNA polymerase II / termination of RNA polymerase I transcription / RNA Polymerase I Promoter Escape / nucleolar large rRNA transcription by RNA polymerase I / RNA polymerase II complex binding / protein phosphatase activator activity / Gap-filling DNA repair synthesis and ligation in TC-NER / transcription by RNA polymerase I / transcription initiation at RNA polymerase I promoter / Estrogen-dependent gene expression / nuclear-transcribed mRNA catabolic process / positive regulation of nuclear-transcribed mRNA poly(A) tail shortening / ATPase activator activity / transcription by RNA polymerase III / RNA polymerase II activity / Dual incision in TC-NER / positive regulation of transcription initiation by RNA polymerase II / transcription elongation by RNA polymerase I / RNA polymerase II core promoter sequence-specific DNA binding / transcription-coupled nucleotide-excision repair / tRNA transcription by RNA polymerase III / RNA polymerase I complex / RNA polymerase I activity / RNA polymerase III complex / positive regulation of translational initiation / translesion synthesis / ATP-dependent activity, acting on DNA / RNA polymerase II, core complex / RNA polymerase II preinitiation complex assembly / translation initiation factor binding / DNA helicase activity / TBP-class protein binding / isomerase activity / DNA-templated transcription initiation / nucleotide-excision repair / transcription initiation at RNA polymerase II promoter / positive regulation of transcription elongation by RNA polymerase II / transcription elongation by RNA polymerase II / transcription coregulator activity / P-body / ribonucleoside binding / DNA-directed 5'-3' RNA polymerase activity / DNA-directed RNA polymerase / mRNA processing / cytoplasmic stress granule Similarity search - Function | ||||||||||||||||||
| Biological species |    | ||||||||||||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.2 Å | ||||||||||||||||||
 Authors Authors | Schilbach S / Aibara S | ||||||||||||||||||
| Funding support |  Germany, 5 items Germany, 5 items
| ||||||||||||||||||
 Citation Citation |  Journal: Cell / Year: 2021 Journal: Cell / Year: 2021Title: Structure of RNA polymerase II pre-initiation complex at 2.9 Å defines initial DNA opening. Authors: Sandra Schilbach / Shintaro Aibara / Christian Dienemann / Frauke Grabbe / Patrick Cramer /  Abstract: Transcription initiation requires assembly of the RNA polymerase II (Pol II) pre-initiation complex (PIC) and opening of promoter DNA. Here, we present the long-sought high-resolution structure of ...Transcription initiation requires assembly of the RNA polymerase II (Pol II) pre-initiation complex (PIC) and opening of promoter DNA. Here, we present the long-sought high-resolution structure of the yeast PIC and define the mechanism of initial DNA opening. We trap the PIC in an intermediate state that contains half a turn of open DNA located 30-35 base pairs downstream of the TATA box. The initially opened DNA region is flanked and stabilized by the polymerase "clamp head loop" and the TFIIF "charged region" that both contribute to promoter-initiated transcription. TFIIE facilitates initiation by buttressing the clamp head loop and by regulating the TFIIH translocase. The initial DNA bubble is then extended in the upstream direction, leading to the open promoter complex and enabling start-site scanning and RNA synthesis. This unique mechanism of DNA opening may permit more intricate regulation than in the Pol I and Pol III systems. | ||||||||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
-Validation report
| Summary document |  emd_12745_validation.pdf.gz emd_12745_validation.pdf.gz | 708.1 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_12745_full_validation.pdf.gz emd_12745_full_validation.pdf.gz | 707.7 KB | Display | |
| Data in XML |  emd_12745_validation.xml.gz emd_12745_validation.xml.gz | 20.7 KB | Display | |
| Data in CIF |  emd_12745_validation.cif.gz emd_12745_validation.cif.gz | 26.6 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-12745 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-12745 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-12745 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-12745 | HTTPS FTP |
-Related structure data
| Related structure data |  7o75MC  7o4iC  7o4jC  7o4kC  7o4lC  7o72C  7o73C M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_12745.map.gz / Format: CCP4 / Size: 209.3 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_12745.map.gz / Format: CCP4 / Size: 209.3 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.05 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
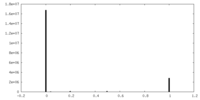
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
+Mask #1
+Additional map: Locally filtered map (non-composite) of an OC refinement...
+Additional map: Half-map 1 of focused refinement encompassing the Pol...
+Additional map: Half-map 2 of focused refinement encompassing the Pol...
+Additional map: Half-map 1 of focused refinement encompassing the Rad3/Ssl1...
+Additional map: Half-map 2 of focused refinement encompassing the Rad3/Ssl1...
+Additional map: Half-map 2 of focused refinement encompassing the Ssl1/Tfb4/Tfb1-3-helix-bundle...
+Additional map: Half-map 1 of focused refinement encompassing the Ssl1/Tfb4/Tfb1-3-helix-bundle...
+Additional map: Half-map 1 of focused refinement encompassing the Ssl2/Tfb2/Tfb5...
+Additional map: Half-map 2 of focused refinement encompassing the Ssl2/Tfb2/Tfb5...
+Additional map: Half-map 2 of focused refinement encompassing the Pol...
+Additional map: Half-map 1 of focused refinement encompassing the Pol...
+Additional map: Half-map 2 of focused refinement encompassing the TFIIE/Tfb1-PHD...
+Additional map: Half-map 1 of focused refinement encompassing the Tfb2...
+Additional map: Half-map 2 of focused refinement encompassing the Tfb2...
+Additional map: Half-map 2 of focused refinement encompassing the upstream...
+Additional map: Half-map 2 of focused refinement encompassing the Pol...
+Additional map: Half-map 1 of focused refinement encompassing the Pol...
+Additional map: Half-map 1 of focused refinement encompassing the upstream...
+Additional map: Half-map 1 of focused refinement encompassing the Pol...
+Additional map: Half-map 1 of focused refinement encompassing the TFIIE/Tfb1-PHD...
+Additional map: Half-map 2 of focused refinement encompassing the Pol...
+Additional map: Half-map 2 of focused refinement encompassing the Pol...
+Additional map: Half-map 1 of focused refinement encompassing the Pol...
+Additional map: Half-map 2 of focused refinement encompassing the Pol...
+Additional map: Half-map 1 of focused refinement encompassing the Pol...
+Half map: #2
+Half map: #1
- Sample components
Sample components
+Entire : Yeast RNA polymerase II transcription pre-initiation complex with...
+Supramolecule #1: Yeast RNA polymerase II transcription pre-initiation complex with...
+Macromolecule #1: General transcription and DNA repair factor IIH helicase subunit XPD
+Macromolecule #2: General transcription and DNA repair factor IIH subunit TFB1
+Macromolecule #3: General transcription and DNA repair factor IIH subunit TFB2
+Macromolecule #4: RNA polymerase II transcription factor B subunit 3
+Macromolecule #5: General transcription and DNA repair factor IIH subunit TFB4
+Macromolecule #6: General transcription and DNA repair factor IIH subunit TFB5
+Macromolecule #7: General transcription and DNA repair factor IIH subunit SSL1
+Macromolecule #8: General transcription and DNA repair factor IIH helicase subunit XPB
+Macromolecule #9: DNA-directed RNA polymerase II subunit RPB1
+Macromolecule #10: DNA-directed RNA polymerase II subunit RPB2
+Macromolecule #11: DNA-directed RNA polymerase II subunit RPB3
+Macromolecule #12: DNA-directed RNA polymerase II subunit RPB4
+Macromolecule #13: DNA-directed RNA polymerases I, II, and III subunit RPABC1
+Macromolecule #14: DNA-directed RNA polymerases I, II, and III subunit RPABC2
+Macromolecule #15: DNA-directed RNA polymerase II subunit RPB7
+Macromolecule #16: DNA-directed RNA polymerases I, II, and III subunit RPABC3
+Macromolecule #17: DNA-directed RNA polymerase II subunit RPB9
+Macromolecule #18: DNA-directed RNA polymerases I, II, and III subunit RPABC5
+Macromolecule #19: DNA-directed RNA polymerase II subunit RPB11
+Macromolecule #20: DNA-directed RNA polymerases I, II, and III subunit RPABC4
+Macromolecule #21: Transcription initiation factor IIB
+Macromolecule #23: TATA-box-binding protein
+Macromolecule #24: Transcription initiation factor IIF subunit alpha
+Macromolecule #25: Transcription initiation factor IIF subunit beta
+Macromolecule #27: Transcription initiation factor IIA large subunit
+Macromolecule #28: Transcription initiation factor IIA subunit 2
+Macromolecule #29: Transcription initiation factor IIE subunit alpha
+Macromolecule #30: Transcription initiation factor IIE subunit beta
+Macromolecule #22: Non-template DNA
+Macromolecule #26: Template DNA
+Macromolecule #31: IRON/SULFUR CLUSTER
+Macromolecule #32: ZINC ION
+Macromolecule #33: BERYLLIUM TRIFLUORIDE ION
+Macromolecule #34: MAGNESIUM ION
+Macromolecule #35: ADENOSINE-5'-DIPHOSPHATE
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.6 |
|---|---|
| Grid | Model: Quantifoil R3.5/1 / Material: COPPER / Support film - Material: CARBON / Support film - topology: CONTINUOUS |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Specialist optics | Energy filter - Name: GIF Quantum LS / Energy filter - Slit width: 20 eV |
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Detector mode: COUNTING / Number real images: 29670 / Average exposure time: 9.0 sec. / Average electron dose: 43.6 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 4.0 µm / Nominal defocus min: 0.5 µm / Nominal magnification: 130000 |
| Sample stage | Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller





































 X (Sec.)
X (Sec.) Y (Row.)
Y (Row.) Z (Col.)
Z (Col.)















































































































































































































































 Trichoplusia ni (cabbage looper)
Trichoplusia ni (cabbage looper)






