[English] 日本語
 Yorodumi
Yorodumi- EMDB-12630: RNA polymerase II pre-initiation complex with intermediary promot... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-12630 | ||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | RNA polymerase II pre-initiation complex with intermediary promoter DNA | ||||||||||||||||||
 Map data Map data | Local resolution filtered and sharpened map | ||||||||||||||||||
 Sample Sample |
| ||||||||||||||||||
| Function / homology |  Function and homology information Function and homology informationMMXD complex / core TFIIH complex portion of holo TFIIH complex / : / positive regulation of core promoter binding / Cytosolic iron-sulfur cluster assembly / RNA polymerase II core complex assembly / central nervous system myelin formation / positive regulation of mitotic recombination / transcription factor TFIIE complex / RNA polymerase transcription factor SL1 complex ...MMXD complex / core TFIIH complex portion of holo TFIIH complex / : / positive regulation of core promoter binding / Cytosolic iron-sulfur cluster assembly / RNA polymerase II core complex assembly / central nervous system myelin formation / positive regulation of mitotic recombination / transcription factor TFIIE complex / RNA polymerase transcription factor SL1 complex / hair cell differentiation / meiotic sister chromatid cohesion / nucleotide-excision repair factor 3 complex / phosphatase activator activity / nucleotide-excision repair, preincision complex assembly / hair follicle maturation / ventricular system development / RNA polymerase III general transcription initiation factor activity / TFIIF-class transcription factor complex binding / transcription factor TFIIK complex / transcriptional start site selection at RNA polymerase II promoter / RNA polymerase I core promoter sequence-specific DNA binding / CAK-ERCC2 complex / transcription factor TFIIF complex / female germ cell nucleus / RNA Polymerase III Transcription Initiation From Type 1 Promoter / RNA Polymerase III Transcription Initiation From Type 2 Promoter / RNA Polymerase III Transcription Initiation From Type 3 Promoter / embryonic cleavage / Formation of RNA Pol II elongation complex / Formation of the Early Elongation Complex / Transcriptional regulation by small RNAs / RNA Polymerase II Pre-transcription Events / TP53 Regulates Transcription of DNA Repair Genes / FGFR2 alternative splicing / RNA polymerase II transcribes snRNA genes / mRNA Capping / mRNA Splicing - Minor Pathway / Processing of Capped Intron-Containing Pre-mRNA / RNA Polymerase II Promoter Escape / RNA Polymerase II Transcription Pre-Initiation And Promoter Opening / RNA Polymerase II Transcription Initiation / RNA Polymerase II Transcription Elongation / RNA Polymerase II Transcription Initiation And Promoter Clearance / RNA Pol II CTD phosphorylation and interaction with CE / Estrogen-dependent gene expression / transcription factor TFIIA complex / Formation of TC-NER Pre-Incision Complex / Dual incision in TC-NER / Gap-filling DNA repair synthesis and ligation in TC-NER / mRNA Splicing - Major Pathway / RNA Polymerase III Abortive And Retractive Initiation / UV protection / regulation of cyclin-dependent protein serine/threonine kinase activity / male pronucleus / female pronucleus / transcription factor TFIIH core complex / transcription factor TFIIH holo complex / cyclin-dependent protein serine/threonine kinase activator activity / RNA polymerase II general transcription initiation factor binding / germinal vesicle / DNA 5'-3' helicase / G protein-coupled receptor internalization / adult heart development / Abortive elongation of HIV-1 transcript in the absence of Tat / nuclear thyroid hormone receptor binding / FGFR2 alternative splicing / transcription preinitiation complex / RNA Polymerase I Transcription Termination / Viral Messenger RNA Synthesis / protein acetylation / Signaling by FGFR2 IIIa TM / cell division site / transcription factor TFIID complex / RNA polymerase II general transcription initiation factor activity / erythrocyte maturation / acetyltransferase activity / regulation of mitotic cell cycle phase transition / hematopoietic stem cell proliferation / positive regulation of nuclear-transcribed mRNA poly(A) tail shortening / RNA Pol II CTD phosphorylation and interaction with CE during HIV infection / RNA Pol II CTD phosphorylation and interaction with CE / Formation of the Early Elongation Complex / Formation of the HIV-1 Early Elongation Complex / mRNA Capping / organelle membrane / bone mineralization / HIV Transcription Initiation / RNA Polymerase II HIV Promoter Escape / Transcription of the HIV genome / RNA Polymerase II Promoter Escape / RNA Polymerase II Transcription Pre-Initiation And Promoter Opening / RNA Polymerase II Transcription Initiation / RNA Polymerase II Transcription Initiation And Promoter Clearance / spinal cord development / mRNA Splicing - Minor Pathway / aryl hydrocarbon receptor binding / viral transcription / TFIIB-class transcription factor binding / RNA polymerase II complex binding Similarity search - Function | ||||||||||||||||||
| Biological species |  | ||||||||||||||||||
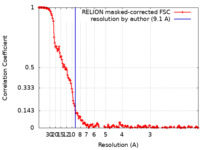
| Method | single particle reconstruction / cryo EM / Resolution: 9.1 Å | ||||||||||||||||||
 Authors Authors | Aibara S / Schilbach S / Cramer P | ||||||||||||||||||
| Funding support |  Germany, 5 items Germany, 5 items
| ||||||||||||||||||
 Citation Citation |  Journal: Nature / Year: 2021 Journal: Nature / Year: 2021Title: Structures of mammalian RNA polymerase II pre-initiation complexes. Authors: Shintaro Aibara / Sandra Schilbach / Patrick Cramer /  Abstract: The initiation of transcription is a focal point for the regulation of gene activity during mammalian cell differentiation and development. To initiate transcription, RNA polymerase II (Pol II) ...The initiation of transcription is a focal point for the regulation of gene activity during mammalian cell differentiation and development. To initiate transcription, RNA polymerase II (Pol II) assembles with general transcription factors into a pre-initiation complex (PIC) that opens promoter DNA. Previous work provided the molecular architecture of the yeast and human PIC and a topological model for DNA opening by the general transcription factor TFIIH. Here we report the high-resolution cryo-electron microscopy structure of PIC comprising human general factors and Sus scrofa domesticus Pol II, which is 99.9% identical to human Pol II. We determine the structures of PIC with closed and opened promoter DNA at 2.5-2.8 Å resolution, and resolve the structure of TFIIH at 2.9-4.0 Å resolution. We capture the TFIIH translocase XPB in the pre- and post-translocation states, and show that XPB induces and propagates a DNA twist to initiate the opening of DNA approximately 30 base pairs downstream of the TATA box. We also provide evidence that DNA opening occurs in two steps and leads to the detachment of TFIIH from the core PIC, which may stop DNA twisting and enable RNA chain initiation. | ||||||||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_12630.map.gz emd_12630.map.gz | 325 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-12630-v30.xml emd-12630-v30.xml emd-12630.xml emd-12630.xml | 18.2 KB 18.2 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_12630_fsc.xml emd_12630_fsc.xml | 16.1 KB | Display |  FSC data file FSC data file |
| Images |  emd_12630.png emd_12630.png | 135.8 KB | ||
| Masks |  emd_12630_msk_1.map emd_12630_msk_1.map | 347.6 MB |  Mask map Mask map | |
| Others |  emd_12630_additional_1.map.gz emd_12630_additional_1.map.gz emd_12630_half_map_1.map.gz emd_12630_half_map_1.map.gz emd_12630_half_map_2.map.gz emd_12630_half_map_2.map.gz | 205.6 MB 277 MB 277 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-12630 http://ftp.pdbj.org/pub/emdb/structures/EMD-12630 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-12630 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-12630 | HTTPS FTP |
-Related structure data
| Related structure data |  7nvrC  7nvsC  7nvtC  7nvuC  7nvvC  7nvwC  7nvxC  7nvyC  7nvzC  7nw0C C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_12630.map.gz / Format: CCP4 / Size: 347.6 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_12630.map.gz / Format: CCP4 / Size: 347.6 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Local resolution filtered and sharpened map | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.05 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Mask #1
| File |  emd_12630_msk_1.map emd_12630_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
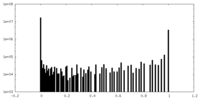
| Density Histograms |
-Additional map: Local resolution filtered map, unsharpened
| File | emd_12630_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Local resolution filtered map, unsharpened | ||||||||||||
| Projections & Slices |
| ||||||||||||
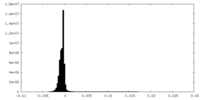
| Density Histograms |
-Half map: Unfiltered half map 1
| File | emd_12630_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Unfiltered half map 1 | ||||||||||||
| Projections & Slices |
| ||||||||||||
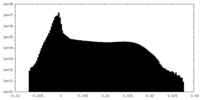
| Density Histograms |
-Half map: Unfiltered half map 2
| File | emd_12630_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Unfiltered half map 2 | ||||||||||||
| Projections & Slices |
| ||||||||||||
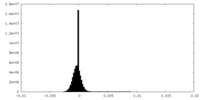
| Density Histograms |
- Sample components
Sample components
-Entire : RNA polymerase II pre-initiation complex with intermediary promot...
| Entire | Name: RNA polymerase II pre-initiation complex with intermediary promoter DNA |
|---|---|
| Components |
|
-Supramolecule #1: RNA polymerase II pre-initiation complex with intermediary promot...
| Supramolecule | Name: RNA polymerase II pre-initiation complex with intermediary promoter DNA type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#22 |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 1.47 MDa |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Average electron dose: 43.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller






















































 X (Sec.)
X (Sec.) Y (Row.)
Y (Row.) Z (Col.)
Z (Col.)