+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-12219 | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | 50S-ObgE-GMPPNP particle | ||||||||||||
 Map data Map data | |||||||||||||
 Sample Sample |
| ||||||||||||
 Keywords Keywords | RIBOSOME / pre-50S / ribosome biogenesis / ribosome assembly | ||||||||||||
| Function / homology |  Function and homology information Function and homology informationguanyl ribonucleotide binding / dormancy process / negative regulation of ribosome biogenesis / guanosine tetraphosphate binding / negative regulation of cytoplasmic translational initiation / ribosomal large subunit binding / transcriptional attenuation / endoribonuclease inhibitor activity / positive regulation of ribosome biogenesis / RNA-binding transcription regulator activity ...guanyl ribonucleotide binding / dormancy process / negative regulation of ribosome biogenesis / guanosine tetraphosphate binding / negative regulation of cytoplasmic translational initiation / ribosomal large subunit binding / transcriptional attenuation / endoribonuclease inhibitor activity / positive regulation of ribosome biogenesis / RNA-binding transcription regulator activity / negative regulation of cytoplasmic translation / DnaA-L2 complex / translation repressor activity / negative regulation of DNA-templated DNA replication initiation / mRNA regulatory element binding translation repressor activity / cytosolic ribosome assembly / response to reactive oxygen species / ribosome assembly / assembly of large subunit precursor of preribosome / chromosome segregation / translational initiation / regulation of cell growth / DNA-templated transcription termination / response to radiation / mRNA 5'-UTR binding / GDP binding / large ribosomal subunit / transferase activity / ribosome binding / 5S rRNA binding / ribosomal large subunit assembly / large ribosomal subunit rRNA binding / cytosolic large ribosomal subunit / Hydrolases; Acting on acid anhydrides; Acting on GTP to facilitate cellular and subcellular movement / cytoplasmic translation / tRNA binding / negative regulation of translation / rRNA binding / structural constituent of ribosome / ribosome / translation / response to antibiotic / negative regulation of DNA-templated transcription / mRNA binding / GTPase activity / GTP binding / magnesium ion binding / DNA binding / RNA binding / zinc ion binding / cytoplasm / cytosol Similarity search - Function | ||||||||||||
| Biological species |  | ||||||||||||
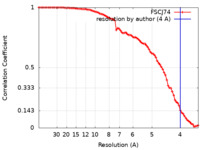
| Method | single particle reconstruction / cryo EM / Resolution: 4.0 Å | ||||||||||||
 Authors Authors | Hilal T / Nikolay R | ||||||||||||
| Funding support |  Germany, 3 items Germany, 3 items
| ||||||||||||
 Citation Citation |  Journal: Mol Cell / Year: 2021 Journal: Mol Cell / Year: 2021Title: Snapshots of native pre-50S ribosomes reveal a biogenesis factor network and evolutionary specialization. Authors: Rainer Nikolay / Tarek Hilal / Sabine Schmidt / Bo Qin / David Schwefel / Carlos H Vieira-Vieira / Thorsten Mielke / Jörg Bürger / Justus Loerke / Kazuaki Amikura / Timo Flügel / Takuya ...Authors: Rainer Nikolay / Tarek Hilal / Sabine Schmidt / Bo Qin / David Schwefel / Carlos H Vieira-Vieira / Thorsten Mielke / Jörg Bürger / Justus Loerke / Kazuaki Amikura / Timo Flügel / Takuya Ueda / Matthias Selbach / Elke Deuerling / Christian M T Spahn /   Abstract: Ribosome biogenesis is a fundamental multi-step cellular process that culminates in the formation of ribosomal subunits, whose production and modification are regulated by numerous biogenesis factors. ...Ribosome biogenesis is a fundamental multi-step cellular process that culminates in the formation of ribosomal subunits, whose production and modification are regulated by numerous biogenesis factors. In this study, we analyze physiologic prokaryotic ribosome biogenesis by isolating bona fide pre-50S subunits from an Escherichia coli strain with the biogenesis factor ObgE, affinity tagged at its native gene locus. Our integrative structural approach reveals a network of interacting biogenesis factors consisting of YjgA, RluD, RsfS, and ObgE on the immature pre-50S subunit. In addition, our study provides mechanistic insight into how the GTPase ObgE, in concert with other biogenesis factors, facilitates the maturation of the 50S functional core and reveals both conserved and divergent evolutionary features of ribosome biogenesis between prokaryotes and eukaryotes. | ||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_12219.map.gz emd_12219.map.gz | 9.4 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-12219-v30.xml emd-12219-v30.xml emd-12219.xml emd-12219.xml | 46.3 KB 46.3 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_12219_fsc.xml emd_12219_fsc.xml | 8.3 KB | Display |  FSC data file FSC data file |
| Images |  emd_12219.png emd_12219.png | 97.4 KB | ||
| Filedesc metadata |  emd-12219.cif.gz emd-12219.cif.gz | 10.3 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-12219 http://ftp.pdbj.org/pub/emdb/structures/EMD-12219 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-12219 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-12219 | HTTPS FTP |
-Related structure data
| Related structure data |  7bl6MC  7bl2C  7bl3C  7bl4C  7bl5C M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_12219.map.gz / Format: CCP4 / Size: 52.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_12219.map.gz / Format: CCP4 / Size: 52.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.74 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
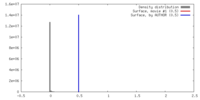
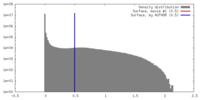
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
+Entire : in vitro reconstituted 50S-ObgE-GMPPNP
+Supramolecule #1: in vitro reconstituted 50S-ObgE-GMPPNP
+Macromolecule #1: 50S ribosomal protein L36
+Macromolecule #2: 50S ribosomal protein L2
+Macromolecule #3: 50S ribosomal protein L3
+Macromolecule #4: 50S ribosomal protein L4
+Macromolecule #5: 50S ribosomal protein L5
+Macromolecule #6: 50S ribosomal protein L6
+Macromolecule #7: 50S ribosomal protein L13
+Macromolecule #8: 50S ribosomal protein L15
+Macromolecule #9: 50S ribosomal protein L17
+Macromolecule #10: 50S ribosomal protein L18
+Macromolecule #11: 50S ribosomal protein L20
+Macromolecule #12: 50S ribosomal protein L21
+Macromolecule #13: 50S ribosomal protein L22
+Macromolecule #14: 50S ribosomal protein L23
+Macromolecule #15: 50S ribosomal protein L24
+Macromolecule #16: 50S ribosomal protein L25
+Macromolecule #17: 50S ribosomal protein L27
+Macromolecule #18: 50S ribosomal protein L28
+Macromolecule #19: 50S ribosomal protein L29
+Macromolecule #20: 50S ribosomal protein L30
+Macromolecule #21: 50S ribosomal protein L32
+Macromolecule #22: 50S ribosomal protein L33
+Macromolecule #23: 50S ribosomal protein L34
+Macromolecule #24: 50S ribosomal protein L14
+Macromolecule #25: 50S ribosomal protein L19
+Macromolecule #26: 50S ribosomal protein L16
+Macromolecule #27: 50S ribosomal protein L9
+Macromolecule #28: 50S ribosomal protein L31
+Macromolecule #31: GTPase ObgE/CgtA
+Macromolecule #32: 50S ribosomal protein L35
+Macromolecule #29: 23S ribosomal RNA
+Macromolecule #30: 5S ribosomal RNA
+Macromolecule #33: ZINC ION
+Macromolecule #34: MAGNESIUM ION
+Macromolecule #35: PHOSPHOAMINOPHOSPHONIC ACID-GUANYLATE ESTER
+Macromolecule #36: water
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TALOS ARCTICA |
|---|---|
| Image recording | Film or detector model: FEI FALCON III (4k x 4k) / Detector mode: INTEGRATING / Number real images: 2179 / Average electron dose: 40.0 e/Å2 |
| Electron beam | Acceleration voltage: 200 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.5 µm / Nominal defocus min: 1.0 µm / Nominal magnification: 120000 |
| Experimental equipment |  Model: Talos Arctica / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Refinement | Space: REAL / Protocol: RIGID BODY FIT |
|---|---|
| Output model |  PDB-7bl6: |
 Movie
Movie Controller
Controller






















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)