[English] 日本語
 Yorodumi
Yorodumi- EMDB-1176: Structure of epsilon15 bacteriophage reveals genome organization ... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-1176 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Structure of epsilon15 bacteriophage reveals genome organization and DNA packaging/injection apparatus. | |||||||||
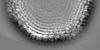
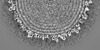
 Map data Map data | 3-fold view of icosahedral symmetry imposed capsid shell map | |||||||||
 Sample Sample |
| |||||||||
| Biological species |   Salmonella phage epsilon15 (virus) Salmonella phage epsilon15 (virus) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 9.5 Å | |||||||||
 Authors Authors | Jiang W / Chang J / Jakana J / Weigele P / King J / Chiu W | |||||||||
 Citation Citation |  Journal: Nature / Year: 2006 Journal: Nature / Year: 2006Title: Structure of epsilon15 bacteriophage reveals genome organization and DNA packaging/injection apparatus. Authors: Wen Jiang / Juan Chang / Joanita Jakana / Peter Weigele / Jonathan King / Wah Chiu /  Abstract: The critical viral components for packaging DNA, recognizing and binding to host cells, and injecting the condensed DNA into the host are organized at a single vertex of many icosahedral viruses. ...The critical viral components for packaging DNA, recognizing and binding to host cells, and injecting the condensed DNA into the host are organized at a single vertex of many icosahedral viruses. These component structures do not share icosahedral symmetry and cannot be resolved using a conventional icosahedral averaging method. Here we report the structure of the entire infectious Salmonella bacteriophage epsilon15 (ref. 1) determined from single-particle cryo-electron microscopy, without icosahedral averaging. This structure displays not only the icosahedral shell of 60 hexamers and 11 pentamers, but also the non-icosahedral components at one pentameric vertex. The densities at this vertex can be identified as the 12-subunit portal complex sandwiched between an internal cylindrical core and an external tail hub connecting to six projecting trimeric tailspikes. The viral genome is packed as coaxial coils in at least three outer layers with approximately 90 terminal nucleotides extending through the protein core and the portal complex and poised for injection. The shell protein from icosahedral reconstruction at higher resolution exhibits a similar fold to that of other double-stranded DNA viruses including herpesvirus, suggesting a common ancestor among these diverse viruses. The image reconstruction approach should be applicable to studying other biological nanomachines with components of mixed symmetries. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_1176.map.gz emd_1176.map.gz | 151.1 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-1176-v30.xml emd-1176-v30.xml emd-1176.xml emd-1176.xml | 9 KB 9 KB | Display Display |  EMDB header EMDB header |
| Images |  1176.gif 1176.gif | 51.5 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-1176 http://ftp.pdbj.org/pub/emdb/structures/EMD-1176 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-1176 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-1176 | HTTPS FTP |
-Validation report
| Summary document |  emd_1176_validation.pdf.gz emd_1176_validation.pdf.gz | 265.7 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_1176_full_validation.pdf.gz emd_1176_full_validation.pdf.gz | 264.8 KB | Display | |
| Data in XML |  emd_1176_validation.xml.gz emd_1176_validation.xml.gz | 4.8 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-1176 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-1176 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-1176 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-1176 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_1176.map.gz / Format: CCP4 / Size: 202.6 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_1176.map.gz / Format: CCP4 / Size: 202.6 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | 3-fold view of icosahedral symmetry imposed capsid shell map | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. generated in cubic-lattice coordinate | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.59 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
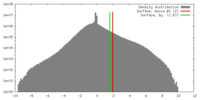
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : Bacteriophage Epsilon15
| Entire | Name: Bacteriophage Epsilon15 |
|---|---|
| Components |
|
-Supramolecule #1000: Bacteriophage Epsilon15
| Supramolecule | Name: Bacteriophage Epsilon15 / type: sample / ID: 1000 / Details: icosahedral shell structure / Oligomeric state: icosahedral / Number unique components: 1 |
|---|---|
| Molecular weight | Theoretical: 500 KDa |
-Supramolecule #1: Salmonella phage epsilon15
| Supramolecule | Name: Salmonella phage epsilon15 / type: virus / ID: 1 / NCBI-ID: 215158 / Sci species name: Salmonella phage epsilon15 / Virus type: VIRION / Virus isolate: STRAIN / Virus enveloped: No / Virus empty: No |
|---|---|
| Host (natural) | Organism:  Salmonella (bacteria) / synonym: BACTERIA(EUBACTERIA) Salmonella (bacteria) / synonym: BACTERIA(EUBACTERIA) |
| Virus shell | Shell ID: 1 / Name: shell / Diameter: 700 Å / T number (triangulation number): 7 |
| Virus shell | Shell ID: 2 / Name: shell / Diameter: 700 Å / T number (triangulation number): 7 |
| Virus shell | Shell ID: 3 / Name: shell / Diameter: 700 Å / T number (triangulation number): 7 |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 / Details: 10 mM Tris pH7.5, 25 mM NaCl and 5 mM MgCl2 |
|---|---|
| Grid | Details: Quantifoil R2/2 grid |
| Vitrification | Cryogen name: ETHANE / Instrument: OTHER / Details: Vitrification instrument: Vitrobot |
- Electron microscopy
Electron microscopy
| Microscope | JEOL 3000SFF |
|---|---|
| Temperature | Average: 4.2 K |
| Date | Oct 10, 2004 |
| Image recording | Category: FILM / Film or detector model: KODAK SO-163 FILM / Digitization - Scanner: OTHER / Digitization - Sampling interval: 6.35 µm / Number real images: 200 / Average electron dose: 25 e/Å2 / Details: Scanned by Nikon CoolScan 9000ED / Bits/pixel: 14 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 1.6 mm / Nominal defocus max: 2.0 µm / Nominal defocus min: 0.5 µm / Nominal magnification: 60000 |
| Sample stage | Specimen holder: Topentry / Specimen holder model: OTHER |
- Image processing
Image processing
| CTF correction | Details: per particle |
|---|---|
| Final reconstruction | Applied symmetry - Point group: I (icosahedral) / Algorithm: OTHER / Resolution.type: BY AUTHOR / Resolution: 9.5 Å / Resolution method: FSC 0.5 CUT-OFF / Software - Name: SAVR / Number images used: 6000 |
 Movie
Movie Controller
Controller


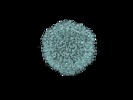






 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)