[English] 日本語
 Yorodumi
Yorodumi- EMDB-1165: A 9 angstroms single particle reconstruction from CCD captured im... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-1165 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | A 9 angstroms single particle reconstruction from CCD captured images on a 200 kV electron cryomicroscope. | |||||||||
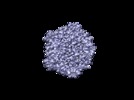
 Map data Map data | this is a half map of CPV oriented along the 5 fold axis of symmetry. Raw Data Collected on 4k x 4k Gatan US4000 CCD Camera. Reconstruction to 9 A. | |||||||||
 Sample Sample |
| |||||||||
| Biological species |   Bombyx mori cypovirus 1 Bombyx mori cypovirus 1 | |||||||||
| Method | single particle reconstruction / cryo EM / negative staining / Resolution: 9.0 Å | |||||||||
 Authors Authors | Booth C / Jiang W / Baker ML / Zhou ZH / Ludtke SJ / Chiu W | |||||||||
 Citation Citation |  Journal: J Struct Biol / Year: 2004 Journal: J Struct Biol / Year: 2004Title: A 9 angstroms single particle reconstruction from CCD captured images on a 200 kV electron cryomicroscope. Authors: Christopher R Booth / Wen Jiang / Matthew L Baker / Z Hong Zhou / Steven J Ludtke / Wah Chiu /  Abstract: Sub-nanometer resolution structure determination is becoming a common practice in electron cryomicroscopy of macromolecular assemblies. The data for these studies have until now been collected on ...Sub-nanometer resolution structure determination is becoming a common practice in electron cryomicroscopy of macromolecular assemblies. The data for these studies have until now been collected on photographic film. Using cytoplasmic polyhedrosis virus (CPV), a previously determined structure, as a test specimen, we show the feasibility of obtaining a 9 angstroms structure from images acquired from a 4 k x 4 k Gatan CCD on a 200 kV electron cryomicroscope. The match of the alpha-helices in the protein components of the CPV with the previous structure of the same virus validates the suitability of this type of camera as the recording media targeted for single particle reconstructions at sub-nanometer resolution. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_1165.map.gz emd_1165.map.gz | 117.9 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-1165-v30.xml emd-1165-v30.xml emd-1165.xml emd-1165.xml | 9.6 KB 9.6 KB | Display Display |  EMDB header EMDB header |
| Images |  1165.gif 1165.gif | 13.4 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-1165 http://ftp.pdbj.org/pub/emdb/structures/EMD-1165 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-1165 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-1165 | HTTPS FTP |
-Related structure data
| Similar structure data |
|---|
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_1165.map.gz / Format: CCP4 / Size: 231.9 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_1165.map.gz / Format: CCP4 / Size: 231.9 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | this is a half map of CPV oriented along the 5 fold axis of symmetry. Raw Data Collected on 4k x 4k Gatan US4000 CCD Camera. Reconstruction to 9 A. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. generated in cubic-lattice coordinate | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.8 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : Cytoplasmic Polyhedrosis Virus
| Entire | Name: Cytoplasmic Polyhedrosis Virus |
|---|---|
| Components |
|
-Supramolecule #1000: Cytoplasmic Polyhedrosis Virus
| Supramolecule | Name: Cytoplasmic Polyhedrosis Virus / type: sample / ID: 1000 / Number unique components: 1 |
|---|
-Supramolecule #1: Bombyx mori cypovirus 1
| Supramolecule | Name: Bombyx mori cypovirus 1 / type: virus / ID: 1 / Name.synonym: CPV / NCBI-ID: 110829 / Sci species name: Bombyx mori cypovirus 1 / Virus type: VIRION / Virus isolate: SPECIES / Virus enveloped: No / Virus empty: No / Syn species name: CPV |
|---|---|
| Host (natural) | Organism:  |
| Virus shell | Shell ID: 1 / Name: capsid / Diameter: 750 Å / T number (triangulation number): 1 |
-Experimental details
-Structure determination
| Method | negative staining, cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 1 mg/mL |
|---|---|
| Buffer | pH: 7.4 / Details: Phosphate Buffered Saline ph 7.4 |
| Staining | Type: NEGATIVE / Details: flash frozen in liquid ethane |
| Grid | Details: irradiated quantafoil R2-1 holey carbon grids |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 45 % / Chamber temperature: 80 K / Instrument: HOMEMADE PLUNGER Details: Vitrification instrument: manual plunger. homemade apparatus Method: blot on 1 side and plunge |
- Electron microscopy
Electron microscopy
| Microscope | JEOL 2010F |
|---|---|
| Temperature | Min: 90 K / Max: 90 K / Average: 90 K |
| Date | Mar 1, 2003 |
| Image recording | Category: CCD / Film or detector model: GATAN ULTRASCAN 4000 (4k x 4k) / Number real images: 856 / Average electron dose: 15 e/Å2 / Bits/pixel: 12 |
| Electron beam | Acceleration voltage: 200 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Calibrated magnification: 83062 / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 1 mm / Nominal defocus max: 4.0 µm / Nominal defocus min: 0.8 µm / Nominal magnification: 60000 |
| Sample stage | Specimen holder: Side entry liquid nitrogen-cooled cryo specimen holder Specimen holder model: GATAN LIQUID NITROGEN |
- Image processing
Image processing
| Details | particles selected using Ethan and manually verified. |
|---|---|
| CTF correction | Details: SAVR per frame |
| Final reconstruction | Applied symmetry - Point group: I (icosahedral) / Algorithm: OTHER / Resolution.type: BY AUTHOR / Resolution: 9.0 Å / Resolution method: FSC 0.5 CUT-OFF / Software - Name: SAVR / Number images used: 6465 |
-Atomic model buiding 1
| Software | Name: Foldhunter |
|---|---|
| Details | Protocol: Rigid Body. Reovirus lambda1 protein was fitted to CSPA. Helixhunter was used to identify alpha helices. |
| Refinement | Protocol: RIGID BODY FIT / Target criteria: Cross correlation |
 Movie
Movie Controller
Controller









 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)





















