+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-11187 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | hSARM1 GraFix-ed | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | NADase / ARM domain / SAM domain / TIR domain / HYDROLASE | |||||||||
| Function / homology |  Function and homology information Function and homology informationnegative regulation of MyD88-independent toll-like receptor signaling pathway / extrinsic component of synaptic membrane / MyD88-independent TLR4 cascade / NADP+ nucleosidase activity / Toll Like Receptor 3 (TLR3) Cascade / NAD+ nucleosidase activity / regulation of synapse pruning / modification of postsynaptic structure / NAD+ catabolic process / ADP-ribosyl cyclase/cyclic ADP-ribose hydrolase ...negative regulation of MyD88-independent toll-like receptor signaling pathway / extrinsic component of synaptic membrane / MyD88-independent TLR4 cascade / NADP+ nucleosidase activity / Toll Like Receptor 3 (TLR3) Cascade / NAD+ nucleosidase activity / regulation of synapse pruning / modification of postsynaptic structure / NAD+ catabolic process / ADP-ribosyl cyclase/cyclic ADP-ribose hydrolase / NAD+ nucleosidase activity, cyclic ADP-ribose generating / protein localization to mitochondrion / nervous system process / Hydrolases; Glycosylases; Hydrolysing N-glycosyl compounds / regulation of dendrite morphogenesis / response to axon injury / response to glucose / signaling adaptor activity / regulation of neuron apoptotic process / TRAF6-mediated induction of TAK1 complex within TLR4 complex / Activation of IRF3, IRF7 mediated by TBK1, IKKε (IKBKE) / IKK complex recruitment mediated by RIP1 / neuromuscular junction / nervous system development / microtubule / cell differentiation / mitochondrial outer membrane / innate immune response / axon / synapse / dendrite / glutamatergic synapse / cell surface / signal transduction / protein-containing complex / mitochondrion / identical protein binding / cytoplasm / cytosol Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 2.88 Å | |||||||||
 Authors Authors | Sporny M / Guez-Haddad J | |||||||||
| Funding support |  Israel, 2 items Israel, 2 items
| |||||||||
 Citation Citation |  Journal: Elife / Year: 2020 Journal: Elife / Year: 2020Title: Structural basis for SARM1 inhibition and activation under energetic stress. Authors: Michael Sporny / Julia Guez-Haddad / Tami Khazma / Avraham Yaron / Moshe Dessau / Yoel Shkolnisky / Carsten Mim / Michail N Isupov / Ran Zalk / Michael Hons / Yarden Opatowsky /     Abstract: SARM1, an executor of axonal degeneration, displays NADase activity that depletes the key cellular metabolite, NAD+, in response to nerve injury. The basis of SARM1 inhibition and its activation ...SARM1, an executor of axonal degeneration, displays NADase activity that depletes the key cellular metabolite, NAD+, in response to nerve injury. The basis of SARM1 inhibition and its activation under stress conditions are still unknown. Here, we present cryo-EM maps of SARM1 at 2.9 and 2.7 Å resolutions. These indicate that SARM1 homo-octamer avoids premature activation by assuming a packed conformation, with ordered inner and peripheral rings, that prevents dimerization and activation of the catalytic domains. This inactive conformation is stabilized by binding of SARM1's own substrate NAD+ in an allosteric location, away from the catalytic sites. This model was validated by mutagenesis of the allosteric site, which led to constitutively active SARM1. We propose that the reduction of cellular NAD+ concentration contributes to the disassembly of SARM1's peripheral ring, which allows formation of active NADase domain dimers, thereby further depleting NAD+ to cause an energetic catastrophe and cell death. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_11187.map.gz emd_11187.map.gz | 80.5 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-11187-v30.xml emd-11187-v30.xml emd-11187.xml emd-11187.xml | 10.2 KB 10.2 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_11187.png emd_11187.png | 147 KB | ||
| Filedesc metadata |  emd-11187.cif.gz emd-11187.cif.gz | 5.4 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-11187 http://ftp.pdbj.org/pub/emdb/structures/EMD-11187 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-11187 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-11187 | HTTPS FTP |
-Related structure data
| Related structure data |  6zfxMC  6zg0C  6zg1C  7anwC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_11187.map.gz / Format: CCP4 / Size: 163.6 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_11187.map.gz / Format: CCP4 / Size: 163.6 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.827 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : hSARM1
| Entire | Name: hSARM1 |
|---|---|
| Components |
|
-Supramolecule #1: hSARM1
| Supramolecule | Name: hSARM1 / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Macromolecule #1: NAD(+) hydrolase SARM1
| Macromolecule | Name: NAD(+) hydrolase SARM1 / type: protein_or_peptide / ID: 1 / Number of copies: 8 / Enantiomer: LEVO / EC number: ADP-ribosyl cyclase/cyclic ADP-ribose hydrolase |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 79.971258 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MSYHHHHHHD YDIPTTENLY FQGAMGSERL AVPGPDGGGG TGPWWAAGGR GPREVSPGAG TEVQDALERA LPELQQALSA LKQAGGARA VGAGLAEVFQ LVEEAWLLPA VGREVAQGLC DAIRLDGGLD LLLRLLQAPE LETRVQAARL LEQILVAENR D RVARIGLG ...String: MSYHHHHHHD YDIPTTENLY FQGAMGSERL AVPGPDGGGG TGPWWAAGGR GPREVSPGAG TEVQDALERA LPELQQALSA LKQAGGARA VGAGLAEVFQ LVEEAWLLPA VGREVAQGLC DAIRLDGGLD LLLRLLQAPE LETRVQAARL LEQILVAENR D RVARIGLG VILNLAKERE PVELARSVAG ILEHMFKHSE ETCQRLVAAG GLDAVLYWCR RTDPALLRHC ALALGNCALH GG QAVQRRM VEKRAAEWLF PLAFSKEDEL LRLHACLAVA VLATNKEVER EVERSGTLAL VEPLVASLDP GRFARCLVDA SDT SQGRGP DDLQRLVPLL DSNRLEAQCI GAFYLCAEAA IKSLQGKTKV FSDIGAIQSL KRLVSYSTNG TKSALAKRAL RLLG EEVPR PILPSVPSWK EAEVQTWLQQ IGFSKYCESF REQQVDGDLL LRLTEEELQT DLGMKSGITR KRFFRELTEL KTFAN YSTC DRSNLADWLG SLDPRFRQYT YGLVSCGLDR SLLHRVSEQQ LLEDCGIHLG VHRARILTAA REMLHSPLPC TGGKPS GDT PDVFISYRRN SGSQLASLLK VHLQLHGFSV FIDVEKLEAG KFEDKLIQSV MGARNFVLVL SPGALDKCMQ DHDCKDW VH KQIVTALSCG KNIVPIIDGF EWPEPQVLPE DMQAVLTFNG IKWSHEYQEA TIEKIIRFLQ GRSSRDSSAG SDTSLEGA A PMGPT UniProtKB: NAD(+) hydrolase SARM1 |
-Macromolecule #2: (~{E})-4-methylnon-4-enedial
| Macromolecule | Name: (~{E})-4-methylnon-4-enedial / type: ligand / ID: 2 / Number of copies: 16 / Formula: S1N |
|---|---|
| Molecular weight | Theoretical: 168.233 Da |
| Chemical component information | 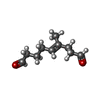 ChemComp-S1N: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 8.8 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | #0 - Image recording ID: 1 / #0 - Film or detector model: GATAN K2 SUMMIT (4k x 4k) / #0 - Average electron dose: 50.0 e/Å2 / #1 - Image recording ID: 2 / #1 - Film or detector model: GATAN K2 SUMMIT (4k x 4k) / #1 - Average electron dose: 50.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)





















