登録情報 データベース : EMDB / ID : EMD-11091タイトル Transcription termination intermediate complex 5 複合体 : Transcription termination complex機能・相同性 分子機能 ドメイン・相同性 構成要素
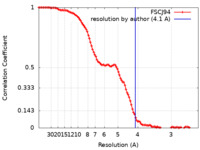
/ / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / 生物種 Escherichia coli (大腸菌)手法 / / 解像度 : 4.1 Å Said N / Hilal T / Buerger J / Mielke T / Loll B / Wahl MC 資金援助 Organization Grant number 国 German Research Foundation (DFG) HA 2549/15-2 German Research Foundation (DFG) RTG 2473-1 German Research Foundation (DFG) WA 1126/11-1 National Institutes of Health/National Center for Complementary and Integrative Health (NIH/NCCIH) GM067153
ジャーナル : Science / 年 : 2021タイトル : Steps toward translocation-independent RNA polymerase inactivation by terminator ATPase ρ.著者 : Nelly Said / Tarek Hilal / Nicholas D Sunday / Ajay Khatri / Jörg Bürger / Thorsten Mielke / Georgiy A Belogurov / Bernhard Loll / Ranjan Sen / Irina Artsimovitch / Markus C Wahl / 要旨 : Factor-dependent transcription termination mechanisms are poorly understood. We determined a series of cryo-electron microscopy structures portraying the hexameric adenosine triphosphatase (ATPase) ... Factor-dependent transcription termination mechanisms are poorly understood. We determined a series of cryo-electron microscopy structures portraying the hexameric adenosine triphosphatase (ATPase) ρ on a pathway to terminating NusA/NusG-modified elongation complexes. An open ρ ring contacts NusA, NusG, and multiple regions of RNA polymerase, trapping and locally unwinding proximal upstream DNA. NusA wedges into the ρ ring, initially sequestering RNA. Upon deflection of distal upstream DNA over the RNA polymerase zinc-binding domain, NusA rotates underneath one capping ρ subunit, which subsequently captures RNA. After detachment of NusG and clamp opening, RNA polymerase loses its grip on the RNA:DNA hybrid and is inactivated. Our structural and functional analyses suggest that ρ, and other termination factors across life, may use analogous strategies to allosterically trap transcription complexes in a moribund state. 履歴 登録 2020年5月28日 - ヘッダ(付随情報) 公開 2021年1月13日 - マップ公開 2021年1月13日 - 更新 2021年7月28日 - 現状 2021年7月28日 処理サイト : PDBe / 状態 : 公開
すべて表示 表示を減らす
 データを開く
データを開く 基本情報
基本情報 マップデータ
マップデータ 試料
試料 機能・相同性情報
機能・相同性情報
 データ登録者
データ登録者 ドイツ,
ドイツ,  米国, 4件
米国, 4件  引用
引用 ジャーナル: Science / 年: 2021
ジャーナル: Science / 年: 2021



 構造の表示
構造の表示 ムービービューア
ムービービューア SurfView
SurfView Molmil
Molmil Jmol/JSmol
Jmol/JSmol ダウンロードとリンク
ダウンロードとリンク emd_11091.map.gz
emd_11091.map.gz EMDBマップデータ形式
EMDBマップデータ形式 emd-11091-v30.xml
emd-11091-v30.xml emd-11091.xml
emd-11091.xml EMDBヘッダ
EMDBヘッダ emd_11091_fsc.xml
emd_11091_fsc.xml FSCデータファイル
FSCデータファイル emd_11091.png
emd_11091.png http://ftp.pdbj.org/pub/emdb/structures/EMD-11091
http://ftp.pdbj.org/pub/emdb/structures/EMD-11091 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-11091
ftp://ftp.pdbj.org/pub/emdb/structures/EMD-11091 emd_11091_validation.pdf.gz
emd_11091_validation.pdf.gz EMDB検証レポート
EMDB検証レポート emd_11091_full_validation.pdf.gz
emd_11091_full_validation.pdf.gz emd_11091_validation.xml.gz
emd_11091_validation.xml.gz emd_11091_validation.cif.gz
emd_11091_validation.cif.gz https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-11091
https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-11091 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-11091
ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-11091 リンク
リンク EMDB (EBI/PDBe) /
EMDB (EBI/PDBe) /  EMDataResource
EMDataResource マップ
マップ ダウンロード / ファイル: emd_11091.map.gz / 形式: CCP4 / 大きさ: 103 MB / タイプ: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES)
ダウンロード / ファイル: emd_11091.map.gz / 形式: CCP4 / 大きさ: 103 MB / タイプ: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) 試料の構成要素
試料の構成要素

 解析
解析 試料調製
試料調製 電子顕微鏡法
電子顕微鏡法 FIELD EMISSION GUN
FIELD EMISSION GUN
 ムービー
ムービー コントローラー
コントローラー




































 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)