[English] 日本語
 Yorodumi
Yorodumi- EMDB-10298: Human endogenous retrovirus (HML2) mature capsid assembly, T=3 ic... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-10298 | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
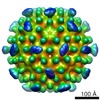
| Title | Human endogenous retrovirus (HML2) mature capsid assembly, T=3 icosahedron | ||||||||||||
 Map data Map data | None | ||||||||||||
 Sample Sample |
| ||||||||||||
| Function / homology |  Function and homology information Function and homology informationviral process / nucleic acid binding / structural molecule activity / zinc ion binding / plasma membrane Similarity search - Function | ||||||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | ||||||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 4.34 Å | ||||||||||||
 Authors Authors | Acton OJH / Taylor IA / Rosenthal PB | ||||||||||||
| Funding support |  United Kingdom, 3 items United Kingdom, 3 items
| ||||||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2019 Journal: Nat Commun / Year: 2019Title: Structural basis for Fullerene geometry in a human endogenous retrovirus capsid. Authors: Oliver Acton / Tim Grant / Giuseppe Nicastro / Neil J Ball / David C Goldstone / Laura E Robertson / Kasim Sader / Andrea Nans / Andres Ramos / Jonathan P Stoye / Ian A Taylor / Peter B Rosenthal /     Abstract: The HML2 (HERV-K) group constitutes the most recently acquired family of human endogenous retroviruses, with many proviruses less than one million years old. Many maintain intact open reading frames ...The HML2 (HERV-K) group constitutes the most recently acquired family of human endogenous retroviruses, with many proviruses less than one million years old. Many maintain intact open reading frames and provirus expression together with HML2 particle formation are observed in early stage human embryo development and are associated with pluripotency as well as inflammatory disease, cancers and HIV-1 infection. Here, we reconstruct the core structural protein (CA) of an HML2 retrovirus, assemble particles in vitro and employ single particle cryogenic electron microscopy (cryo-EM) to determine structures of four classes of CA Fullerene shell assemblies. These icosahedral and capsular assemblies reveal at high-resolution the molecular interactions that allow CA to form both pentamers and hexamers and show how invariant pentamers and structurally plastic hexamers associate to form the unique polyhedral structures found in retroviral cores. | ||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_10298.map.gz emd_10298.map.gz | 677.3 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-10298-v30.xml emd-10298-v30.xml emd-10298.xml emd-10298.xml | 15.4 KB 15.4 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_10298.png emd_10298.png | 91.8 KB | ||
| Others |  emd_10298_additional.map.gz emd_10298_additional.map.gz | 677.2 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-10298 http://ftp.pdbj.org/pub/emdb/structures/EMD-10298 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-10298 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-10298 | HTTPS FTP |
-Validation report
| Summary document |  emd_10298_validation.pdf.gz emd_10298_validation.pdf.gz | 388.4 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_10298_full_validation.pdf.gz emd_10298_full_validation.pdf.gz | 387.5 KB | Display | |
| Data in XML |  emd_10298_validation.xml.gz emd_10298_validation.xml.gz | 8.7 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-10298 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-10298 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-10298 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-10298 | HTTPS FTP |
-Related structure data
| Related structure data |  6ssmMC  6sa9C  6saiC  6ssjC  6sskC  6sslC C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_10298.map.gz / Format: CCP4 / Size: 729 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_10298.map.gz / Format: CCP4 / Size: 729 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | None | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.09 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Additional map: Unsharpened map
| File | emd_10298_additional.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Unsharpened map | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Human endogenous retrovirus (HML2) mature capsid assembly, T=3 ic...
| Entire | Name: Human endogenous retrovirus (HML2) mature capsid assembly, T=3 icosahedron |
|---|---|
| Components |
|
-Supramolecule #1: Human endogenous retrovirus (HML2) mature capsid assembly, T=3 ic...
| Supramolecule | Name: Human endogenous retrovirus (HML2) mature capsid assembly, T=3 icosahedron type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Recombinant expression | Organism:  |
| Molecular weight | Experimental: 5.1 MDa |
-Macromolecule #1: Endogenous retrovirus group K member 24 Gag polyprotein
| Macromolecule | Name: Endogenous retrovirus group K member 24 Gag polyprotein type: protein_or_peptide / ID: 1 / Number of copies: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 27.570488 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: PVTLEPMPPG EGAQEGEPPT VEARYKSFSI KMLKDMKEGV KQYGPNSPYM RTLLDSIAHG HRLIPYDWEI LAKSSLSPSQ FLQFKTWWI DGVQEQVRRN RAANPPVNID ADQLLGIGQN WSTISQQALM QNEAIEQVRA ICLRAWEKIQ DPGSACPSFN T VRQGSKEP ...String: PVTLEPMPPG EGAQEGEPPT VEARYKSFSI KMLKDMKEGV KQYGPNSPYM RTLLDSIAHG HRLIPYDWEI LAKSSLSPSQ FLQFKTWWI DGVQEQVRRN RAANPPVNID ADQLLGIGQN WSTISQQALM QNEAIEQVRA ICLRAWEKIQ DPGSACPSFN T VRQGSKEP YPDFVARLQD VAQKSIADEK ARKVIVELMA YENANPECQS AIKPLKGKVP AGSDVISEYV KACDGIGGAM HK AMLMAQL E |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 16 mg/mL |
|---|---|
| Buffer | pH: 8 |
| Grid | Model: Quantifoil R2/2 / Material: COPPER / Mesh: 200 / Support film - Material: CARBON / Support film - topology: HOLEY ARRAY / Pretreatment - Type: GLOW DISCHARGE |
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: FEI FALCON III (4k x 4k) / Detector mode: COUNTING / Number grids imaged: 1 / Number real images: 10935 / Average exposure time: 60.0 sec. / Average electron dose: 30.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| CTF correction | Software - Name: CTFFIND (ver. 4.1) |
|---|---|
| Final reconstruction | Applied symmetry - Point group: I (icosahedral) / Resolution.type: BY AUTHOR / Resolution: 4.34 Å / Resolution method: FSC 0.143 CUT-OFF / Software - Name: cisTEM / Number images used: 348 |
| Initial angle assignment | Type: MAXIMUM LIKELIHOOD / Software - Name: RELION (ver. 2.1) |
| Final angle assignment | Type: MAXIMUM LIKELIHOOD / Software - Name: RELION (ver. 2.1) |
-Atomic model buiding 1
| Refinement | Space: REAL / Protocol: OTHER / Overall B value: 90 / Target criteria: map vs model FSC |
|---|---|
| Output model |  PDB-6ssm: |
 Movie
Movie Controller
Controller

















 Z
Z Y
Y X
X