+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-0844 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Signal substraction of TcdB1-TccC2 and part of TcdA1 | |||||||||
 Map data Map data | B1C1 and part of A1 | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | protein transportation / TcdA1 / TcdB1 / TccC2 / TOXIN-PROTEIN TRANSPORT complex | |||||||||
| Function / homology |  Function and homology information Function and homology information | |||||||||
| Biological species |  Photorhabdus luminescens (bacteria) Photorhabdus luminescens (bacteria) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 2.9 Å | |||||||||
 Authors Authors | Yan ZF / Yang GW | |||||||||
 Citation Citation |  Journal: To Be Published Journal: To Be PublishedTitle: Signal substraction of TcdB1-TccC2 and part of TcdA1 Authors: Yan ZF / Zhang N / Yang GW | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_0844.map.gz emd_0844.map.gz | 12.8 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-0844-v30.xml emd-0844-v30.xml emd-0844.xml emd-0844.xml | 13 KB 13 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_0844.png emd_0844.png | 183.4 KB | ||
| Filedesc metadata |  emd-0844.cif.gz emd-0844.cif.gz | 6.2 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-0844 http://ftp.pdbj.org/pub/emdb/structures/EMD-0844 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-0844 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-0844 | HTTPS FTP |
-Related structure data
| Related structure data |  6l7iMC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_0844.map.gz / Format: CCP4 / Size: 209.3 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_0844.map.gz / Format: CCP4 / Size: 209.3 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | B1C1 and part of A1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.338 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
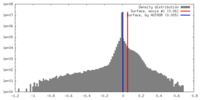
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : Toxin complex TcdA1-TcdB1-TccC2
| Entire | Name: Toxin complex TcdA1-TcdB1-TccC2 |
|---|---|
| Components |
|
-Supramolecule #1: Toxin complex TcdA1-TcdB1-TccC2
| Supramolecule | Name: Toxin complex TcdA1-TcdB1-TccC2 / type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  Photorhabdus luminescens (bacteria) Photorhabdus luminescens (bacteria) |
-Macromolecule #1: TcdA1
| Macromolecule | Name: TcdA1 / type: protein_or_peptide / ID: 1 / Number of copies: 5 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Photorhabdus luminescens (bacteria) Photorhabdus luminescens (bacteria) |
| Molecular weight | Theoretical: 20.255887 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: RALEVERTVS LAEVYAGLPK DNGPFSLAQE IDKLVSQGSG SAGSGNNNLA FGAGTDTKTS LQASVSFADL KIREDYPASL GKIRRIKQI SVTLPALLGP YQDVQAILSY GDKAGLANGC EALAVSHGMN DSGQFQLDFN DGKFLPFEGI AIDQGTLTLS F PNASMPEK GKQATMLKTL NDIILHIRYT IK UniProtKB: TcdA1 |
-Macromolecule #2: TcdB1
| Macromolecule | Name: TcdB1 / type: protein_or_peptide / ID: 2 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Photorhabdus luminescens (bacteria) Photorhabdus luminescens (bacteria) |
| Molecular weight | Theoretical: 165.296453 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MQNSQTFSVT ELSLPKGGGA ITGMGEALTP AGPDGMAALS LPLPISAGRG YAPSLTLNYN SGTGNSPFGL GWDCGVMAIR RRTSTGVPN YDETDTFLGP EGEVLVVALN EAGQADIRSE SSLQGINLGA TFTVTCYRSR LESHFNRLEY WQPQTTGATD F WLIYSPDG ...String: MQNSQTFSVT ELSLPKGGGA ITGMGEALTP AGPDGMAALS LPLPISAGRG YAPSLTLNYN SGTGNSPFGL GWDCGVMAIR RRTSTGVPN YDETDTFLGP EGEVLVVALN EAGQADIRSE SSLQGINLGA TFTVTCYRSR LESHFNRLEY WQPQTTGATD F WLIYSPDG QVHLLGKNPQ ARISNPLNVN QTAQWLLEAS ISSHSEQIYY QYRAEDEAGC ETDELAAHPS ATVQRYLQTV HY GNLTASD VFPTLNGDDP LKSGWMFCLV FDYGERKNSL SEMPLFKATG NWLCRKDRFS RYEYGFELRT RRLCRQILMF HRL QTLSGQ AKGDDEPALV SRLILDYDEN AMVSTLVSVR RVGHEDNNTV TALPPLELAY QPFEPEQTAL WQSMDVLANF NTIQ RWQLL DLKGEGVPGI LYQDRNGWWY RSAQRQAGEE MNAVTWGKMQ LLPITPAVQD NASLMDINGD GQLDWVITGP GLRGY HSQH PDGSWTRFTP LHALPIEYSH PRAQLADLMG AGLSDLVLIG PKSVRLYVNN RDGFTEGRDV VQSGDITLPL PGADAR KLV AFSDVLGSGQ AHLVEVSATQ VTCWPNLGHG RFGQPIVLPG FSQSAASFNP DRVHLADLDG SGPADLIYVH ADRLDIF SN ESGNGFAKPF TLSFPDGLRF DDTCQLQVAD VQGLGVVSLI LSVPHMAPHH WRCDLTNAKP WLLSETNNNM GANHTLHY R SSVQFWLDEK AAALATGQTP VCYLPFPVHT LWQTETEDEI SGNKLVTTLR YAHGAWDGRE REFRGFGYVE QTDSHQLAQ GNAPERTPPA LTKSWYATGL PAVDNALSAG YWRGDKQAFA GFTPRFTLWK EGKDVPLTPE DDHNLYWLNR ALKGQPLRSE LYGLDGSAQ QQIPYTVTES RPQVRQLQDG ATVSPVLWAS VVESRSYHYE RIISDPQCNQ DITLSSDLFG QPLKQVSVQY P RRNKPTTN PYPDTLPDTL FASSYDDQQQ LLRLTCRQSS WHHLIGNELR VLGLPDGTRS DAFTYDAKQV PVDGLNLETL CA ENSLIAD DKPREYLNQQ RTFYTDGKNQ TPLKTPTRQA LIAFTETAVL TESLLSAFDG GITPDELPGI LTQAGYQQEP YLF PRTGEN KVWVARQGYT DYGTEAQFWR PVAQRNSLLT GKMTLKWDTH YCVITQTQDA AGLTVSANYD WRFLTPTQLT DIND NVHLI TLDALGRPVT QRFWGIESGV ATGYSSSEEK PFSPPNDIDT AINLTGPLPV AQCLVYAPDS WMPLFSQETF NTLTQ EEQE TLRDSRIITE DWRICALTRR RWLQSQKIST PLVKLLTNSI GLPPHNLTLT TDRYDRDSEQ QIRQQVAFSD GFGRLL QAS VRHEAGEAWQ RNQDGSLVTK VENTKTRWAV TGRTEYDNKG QTIRTYQPYF LNDWRYVSDD SARKEAYADT HIYDPIG RE IRVITAKGWL RQSQYFPWFT VSEDENDTAA DALV UniProtKB: TcdB1 |
-Macromolecule #3: TccC2
| Macromolecule | Name: TccC2 / type: protein_or_peptide / ID: 3 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Photorhabdus luminescens (bacteria) Photorhabdus luminescens (bacteria) |
| Molecular weight | Theoretical: 74.759125 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MSSYNSAIDQ KTPSIKVLDN RKLNVRTLEY LRTQADENSD ELITFYEFNI PGFQVKSTDP RKNKNQSGPN FIRVFNLAGQ VLREESVDA GRTITLNDIE SRPVLIINAT GVRQNHRYED NTLPGRLLAI TEQVQAGEKT TERLIWAGNT PQEKDYNLAG Q CVRHYDTA ...String: MSSYNSAIDQ KTPSIKVLDN RKLNVRTLEY LRTQADENSD ELITFYEFNI PGFQVKSTDP RKNKNQSGPN FIRVFNLAGQ VLREESVDA GRTITLNDIE SRPVLIINAT GVRQNHRYED NTLPGRLLAI TEQVQAGEKT TERLIWAGNT PQEKDYNLAG Q CVRHYDTA GLTQLNSLSL AGVVLSQSQQ LLVDDKNADW TGEDQSLWQQ KLSSDVYTTQ NKADATGALL TQTDAKGNIQ RL AYDVAGQ LKGCWLTLKG QAEQVIIKSL TYSAAGQKLR EEHGNGVITE YSYEPETQRL IGIATRRPSD AKVLQDLRYQ YDP VGNVIN IRNDAEATRF WRNQKVVPEN SYTYDSLYQL ISATGREMAN IGQQNNQLPS PALPSDNNTY TNYTRSYSYD HSGN LTQIR HSSPATQNNY TVAITLSNRS NRGVLSTLTT DPNQVDTLFD AGGHQTSLLP GQTLIWTPRG ELKQVNNGPG NEWYR YDSN GMRQLKVSEQ PTQNTTQQQR VIYLPGLELR TTQSNATTTE ELHVITLGEA GRAQVRVLHW ESGKPEDVNN NQLRYS YDN LIGSSQLELD NQGQIISEEE YYPFGGTALW AANSQTEASY KTIRYSGKER DATGLYYYGY RYYQPWAGRW LSADPAG TI DGLNLYRMVR NNPVSLQDEN GL UniProtKB: TccC2 |
-Macromolecule #4: TccC2
| Macromolecule | Name: TccC2 / type: protein_or_peptide / ID: 4 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Photorhabdus luminescens (bacteria) Photorhabdus luminescens (bacteria) |
| Molecular weight | Theoretical: 1.905087 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: APEKGKYTKE VNFFDE UniProtKB: TccC2 |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 8 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI/PHILIPS CM300FEG/HE |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Average electron dose: 50.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: SPOT SCAN / Imaging mode: BRIGHT FIELD |
- Image processing
Image processing
| Startup model | Type of model: OTHER |
|---|---|
| Final reconstruction | Resolution.type: BY AUTHOR / Resolution: 2.9 Å / Resolution method: FSC 0.143 CUT-OFF / Number images used: 124828 |
| Initial angle assignment | Type: MAXIMUM LIKELIHOOD |
| Final angle assignment | Type: MAXIMUM LIKELIHOOD |
 Movie
Movie Controller
Controller














 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)