+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-0360 | |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Saccharomyces cerevisiae spliceosomal E complex (UBC4) | |||||||||||||||
 Map data Map data | Spliceosomal E complex (UBC4) | |||||||||||||||
 Sample Sample |
| |||||||||||||||
 Keywords Keywords | pre-mRNA splicing / spliceosome / E complex / RNA BINDING PROTEIN | |||||||||||||||
| Function / homology |  Function and homology information Function and homology informationProcessing of Intronless Pre-mRNAs / nuclear cap binding complex / pre-mRNA 5'-splice site binding / RNA cap binding / Transport of Mature mRNA derived from an Intron-Containing Transcript / primary miRNA processing / mRNA splice site recognition / U4/U6 snRNP / 7-methylguanosine cap hypermethylation / pICln-Sm protein complex ...Processing of Intronless Pre-mRNAs / nuclear cap binding complex / pre-mRNA 5'-splice site binding / RNA cap binding / Transport of Mature mRNA derived from an Intron-Containing Transcript / primary miRNA processing / mRNA splice site recognition / U4/U6 snRNP / 7-methylguanosine cap hypermethylation / pICln-Sm protein complex / snRNP binding / positive regulation of mRNA splicing, via spliceosome / small nuclear ribonucleoprotein complex / splicing factor binding / mRNA 3'-end processing / SMN-Sm protein complex / spliceosomal tri-snRNP complex / commitment complex / mRNA cis splicing, via spliceosome / U2-type prespliceosome assembly / U4 snRNP / U2 snRNP / nuclear-transcribed mRNA catabolic process, nonsense-mediated decay / U1 snRNP / poly(U) RNA binding / response to osmotic stress / U2-type prespliceosome / Processing of Capped Intron-Containing Pre-mRNA / Formation of the Early Elongation Complex / mRNA Capping / precatalytic spliceosome / RNA Polymerase II Pre-transcription Events / spliceosomal complex assembly / mRNA 5'-splice site recognition / Nonsense Mediated Decay (NMD) independent of the Exon Junction Complex (EJC) / Nonsense Mediated Decay (NMD) enhanced by the Exon Junction Complex (EJC) / nuclear-transcribed mRNA catabolic process / Prp19 complex / 7-methylguanosine mRNA capping / U5 snRNP / spliceosomal snRNP assembly / mRNA export from nucleus / U1 snRNA binding / U4/U6 x U5 tri-snRNP complex / catalytic step 2 spliceosome / spliceosomal complex / mRNA transcription by RNA polymerase II / mRNA splicing, via spliceosome / mRNA binding / perinuclear region of cytoplasm / RNA binding / zinc ion binding / nucleus / cytosol / cytoplasm Similarity search - Function | |||||||||||||||
| Biological species |    | |||||||||||||||
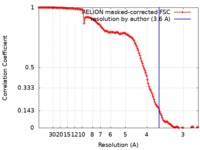
| Method | single particle reconstruction / cryo EM / Resolution: 3.6 Å | |||||||||||||||
 Authors Authors | Liu S / Li X | |||||||||||||||
| Funding support |  United States, 4 items United States, 4 items
| |||||||||||||||
 Citation Citation |  Journal: Nature / Year: 2019 Journal: Nature / Year: 2019Title: A unified mechanism for intron and exon definition and back-splicing. Authors: Xueni Li / Shiheng Liu / Lingdi Zhang / Aaron Issaian / Ryan C Hill / Sara Espinosa / Shasha Shi / Yanxiang Cui / Kalli Kappel / Rhiju Das / Kirk C Hansen / Z Hong Zhou / Rui Zhao /  Abstract: The molecular mechanisms of exon definition and back-splicing are fundamental unanswered questions in pre-mRNA splicing. Here we report cryo-electron microscopy structures of the yeast spliceosomal E ...The molecular mechanisms of exon definition and back-splicing are fundamental unanswered questions in pre-mRNA splicing. Here we report cryo-electron microscopy structures of the yeast spliceosomal E complex assembled on introns, providing a view of the earliest event in the splicing cycle that commits pre-mRNAs to splicing. The E complex architecture suggests that the same spliceosome can assemble across an exon, and that it either remodels to span an intron for canonical linear splicing (typically on short exons) or catalyses back-splicing to generate circular RNA (on long exons). The model is supported by our experiments, which show that an E complex assembled on the middle exon of yeast EFM5 or HMRA1 can be chased into circular RNA when the exon is sufficiently long. This simple model unifies intron definition, exon definition, and back-splicing through the same spliceosome in all eukaryotes and should inspire experiments in many other systems to understand the mechanism and regulation of these processes. | |||||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_0360.map.gz emd_0360.map.gz | 199.6 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-0360-v30.xml emd-0360-v30.xml emd-0360.xml emd-0360.xml | 37.2 KB 37.2 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_0360_fsc.xml emd_0360_fsc.xml | 13.6 KB | Display |  FSC data file FSC data file |
| Images |  emd_0360.png emd_0360.png | 46.3 KB | ||
| Filedesc metadata |  emd-0360.cif.gz emd-0360.cif.gz | 10.7 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-0360 http://ftp.pdbj.org/pub/emdb/structures/EMD-0360 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-0360 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-0360 | HTTPS FTP |
-Validation report
| Summary document |  emd_0360_validation.pdf.gz emd_0360_validation.pdf.gz | 594.4 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_0360_full_validation.pdf.gz emd_0360_full_validation.pdf.gz | 593.9 KB | Display | |
| Data in XML |  emd_0360_validation.xml.gz emd_0360_validation.xml.gz | 13.5 KB | Display | |
| Data in CIF |  emd_0360_validation.cif.gz emd_0360_validation.cif.gz | 18.3 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-0360 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-0360 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-0360 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-0360 | HTTPS FTP |
-Related structure data
| Related structure data |  6n7pMC  0361C  6n7rC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_0360.map.gz / Format: CCP4 / Size: 216 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_0360.map.gz / Format: CCP4 / Size: 216 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Spliceosomal E complex (UBC4) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.36 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
+Entire : Saccharomyces cerevisiae spliceosomal E complex (EBC4)
+Supramolecule #1: Saccharomyces cerevisiae spliceosomal E complex (EBC4)
+Macromolecule #1: U1 small nuclear ribonucleoprotein 70 kDa homolog
+Macromolecule #2: U1 small nuclear ribonucleoprotein C
+Macromolecule #3: U1 small nuclear ribonucleoprotein A
+Macromolecule #4: U1 small nuclear ribonucleoprotein component PRP42
+Macromolecule #5: Pre-mRNA-processing factor 39
+Macromolecule #6: Protein NAM8
+Macromolecule #7: 56 kDa U1 small nuclear ribonucleoprotein component
+Macromolecule #8: U1 small nuclear ribonucleoprotein component SNU71,U1 small nucle...
+Macromolecule #9: Protein LUC7
+Macromolecule #10: Pre-mRNA-processing protein PRP40
+Macromolecule #11: Small nuclear ribonucleoprotein-associated protein B
+Macromolecule #12: Small nuclear ribonucleoprotein Sm D1
+Macromolecule #13: Small nuclear ribonucleoprotein Sm D2
+Macromolecule #14: Small nuclear ribonucleoprotein Sm D3
+Macromolecule #15: Small nuclear ribonucleoprotein E
+Macromolecule #16: Small nuclear ribonucleoprotein F
+Macromolecule #17: Small nuclear ribonucleoprotein G
+Macromolecule #20: Nuclear cap-binding protein complex subunit 1
+Macromolecule #21: Nuclear cap-binding protein subunit 2
+Macromolecule #18: U1 snRNA
+Macromolecule #19: UBC4 pre-mRNA
+Macromolecule #22: ZINC ION
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.9 |
|---|---|
| Grid | Details: unspecified |
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Detector mode: COUNTING / Average electron dose: 30.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller























 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)