+Search query
-Structure paper
| Title | Uncompetitive Allosteric Inhibitor of Mitochondrial Creatine Kinase Prevents Binding and Release of Creatine by Stabilization of Loop Closure. |
|---|---|
| Journal, issue, pages | bioRxiv, Year 2026 |
| Publish date | Jan 8, 2026 |
| PubMed Abstract | Mitochondrial creatine kinase (MtCK) is a key enzyme in energy buffering and homeostasis in cells. It catalyzes transfer of phosphoryl group from ATP to creatine. Overexpression of MtCK occurs in ...Mitochondrial creatine kinase (MtCK) is a key enzyme in energy buffering and homeostasis in cells. It catalyzes transfer of phosphoryl group from ATP to creatine. Overexpression of MtCK occurs in many cancer cells to meet elevated energy demands, which is associated with poor prognosis. This suggests that MtCK may be a promising target for cancer therapeutics. We sought to discover first-in-class selective inhibitors of MtCK with diverse mechanisms of action using high-throughput screening, biochemical characterization and cryo-EM studies. Through these studies, we identified diverse types of compounds that modulate activity of MtCK , including fast-equilibrium and time-dependent orthosteric and allosteric inhibitors. Select hits were subjected to enzymatic and binding assays to assess MtCK inhibition and binding. A subset of inhibitors was advanced into structural studies using cryo-EM resulting in the molecular structure of MtCK with and without bound substrates in complex with an allosteric uncompetitive inhibitor that we discovered. These studies identified compound's unique binding pocket on MtCK and established the molecular steps manifesting in the apparent uncompetitive mode of inhibition. We demonstrate that the compound stabilizes an active site loop in a closed conformation restricting access of the creatine substrate to and the release of phosphocreatine product from its binding pocket. These findings establish chemical tools suitable for validation of MtCK as a promising breast cancer target with high therapeutic potential and build a foundation for future structure-guided optimization of the hit compounds of MtCK we identified and de novo rational design of novel MtCK inhibitors. |
 External links External links |  bioRxiv / bioRxiv /  PubMed:41542558 / PubMed:41542558 /  PubMed Central PubMed Central |
| Methods | EM (single particle) |
| Resolution | 2.59 - 2.6 Å |
| Structure data | EMDB-73766, PDB-9z2d: EMDB-73767, PDB-9z2f: |
| Chemicals |  ChemComp-ADP: 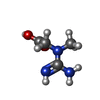 ChemComp-CRN:  PDB-1czq:  ChemComp-MG: |
| Source |
|
 Keywords Keywords | TRANSFERASE/INHIBITOR / Kinase / inhibitor / TRANSFERASE-INHIBITOR complex / transition state analog |
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About Yorodumi Papers
About Yorodumi Papers







 homo sapiens (human)
homo sapiens (human)