+Search query
-Structure paper
| Title | Regulation of STING activation by phosphoinositide and cholesterol. |
|---|---|
| Journal, issue, pages | Nature, Vol. 652, Issue 8109, Page 499-507, Year 2026 |
| Publish date | Feb 4, 2026 |
 Authors Authors | Jie Li / Jay Xiaojun Tan / Zhijian J Chen / Xuewu Zhang / Xiao-Chen Bai /  |
| PubMed Abstract | Stimulator of interferon genes (STING) is an essential adaptor in the cytosolic DNA-sensing innate immune pathway. STING is activated by cyclic GMP-AMP (cGAMP) produced by the DNA sensor cGAMP ...Stimulator of interferon genes (STING) is an essential adaptor in the cytosolic DNA-sensing innate immune pathway. STING is activated by cyclic GMP-AMP (cGAMP) produced by the DNA sensor cGAMP synthase (cGAS). cGAMP-induced high-order oligomerization and translocation of STING from the endoplasmic reticulum to the Golgi and post-Golgi vesicles are critical for STING activation. Other studies have shown that phosphatidylinositol phosphates (PtdInsPs) and cholesterol also have important roles in STING activation, but the underlying mechanisms remain unclear. Here we demonstrate that cGAMP-induced high-order oligomerization of STING is enhanced strongly by phosphatidylinositol 3,5-bisphosphate (PtdIns(3,5)P and PtdIns(4,5)P, and by PtdIns4P to a lesser extent. Our cryo-electron microscopy structures reveal that PtdInsPs together with cholesterol bind at the interface between STING dimers, directly promoting the high-order oligomerization. The structures also provide an explanation for the preference of the STING oligomer to different PtdInsPs. Mutational and biochemical analyses confirm the binding modes of PtdInsPs and cholesterol and their roles in STING activation. Our findings shed light on the regulatory mechanisms of STING mediated by specific lipids, which may underlie the role of intracellular trafficking in dictating STING signalling. |
 External links External links |  Nature / Nature /  PubMed:41639452 PubMed:41639452 |
| Methods | EM (single particle) |
| Resolution | 2.6 - 2.9 Å |
| Structure data | EMDB-46694, PDB-9dan: EMDB-46697, PDB-9dat: EMDB-46699, PDB-9dav: EMDB-46700, PDB-9daw: |
| Chemicals | 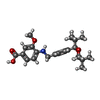 ChemComp-Y6H:  PDB-1bbg:  ChemComp-Y01: 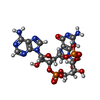 ChemComp-1SY:  PDB-1bbh: 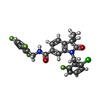 ChemComp-9IM: |
| Source |
|
 Keywords Keywords | SIGNALING PROTEIN/ACTIVATOR / STING / lipid / oligomerization / SIGNALING PROTEIN / SIGNALING PROTEIN-ACTIVATOR complex / oligomer / PI(3 / 5)P2 |
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About Yorodumi Papers
About Yorodumi Papers











 homo sapiens (human)
homo sapiens (human)