+ Open data
Open data
- Basic information
Basic information
| Entry |  | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | STING oligomer bound to PI(3,5)P2 | ||||||||||||
 Map data Map data | |||||||||||||
 Sample Sample |
| ||||||||||||
 Keywords Keywords | STING / lipid / oligomerization / SIGNALING PROTEIN / SIGNALING PROTEIN-ACTIVATOR complex | ||||||||||||
| Function / homology |  Function and homology information Function and homology informationSTING complex / STAT6-mediated induction of chemokines / serine/threonine protein kinase complex / protein localization to endoplasmic reticulum / 2',3'-cyclic GMP-AMP binding / cyclic-di-GMP binding / STING mediated induction of host immune responses / positive regulation of type I interferon-mediated signaling pathway / IRF3-mediated induction of type I IFN / proton channel activity ...STING complex / STAT6-mediated induction of chemokines / serine/threonine protein kinase complex / protein localization to endoplasmic reticulum / 2',3'-cyclic GMP-AMP binding / cyclic-di-GMP binding / STING mediated induction of host immune responses / positive regulation of type I interferon-mediated signaling pathway / IRF3-mediated induction of type I IFN / proton channel activity / reticulophagy / cGAS/STING signaling pathway / pattern recognition receptor signaling pathway / cellular response to exogenous dsRNA / cytoplasmic pattern recognition receptor signaling pathway / protein complex oligomerization / autophagosome membrane / positive regulation of macroautophagy / autophagosome assembly / cellular response to interferon-beta / positive regulation of defense response to virus by host / positive regulation of type I interferon production / endoplasmic reticulum-Golgi intermediate compartment membrane / signaling adaptor activity / activation of innate immune response / positive regulation of interferon-beta production / autophagosome / secretory granule membrane / cytoplasmic vesicle membrane / antiviral innate immune response / Regulation of innate immune responses to cytosolic DNA / protein serine/threonine kinase binding / SARS-CoV-1 activates/modulates innate immune responses / peroxisome / regulation of inflammatory response / defense response to virus / RNA polymerase II-specific DNA-binding transcription factor binding / mitochondrial outer membrane / transcription coactivator activity / endosome / cilium / ciliary basal body / Golgi membrane / innate immune response / Neutrophil degranulation / ubiquitin protein ligase binding / endoplasmic reticulum membrane / protein kinase binding / SARS-CoV-2 activates/modulates innate and adaptive immune responses / perinuclear region of cytoplasm / protein homodimerization activity / positive regulation of transcription by RNA polymerase II / nucleoplasm / identical protein binding / plasma membrane / cytosol Similarity search - Function | ||||||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | ||||||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 2.9 Å | ||||||||||||
 Authors Authors | Li J / Zhang X / Bai X | ||||||||||||
| Funding support |  United States, 3 items United States, 3 items
| ||||||||||||
 Citation Citation |  Journal: Nature / Year: 2026 Journal: Nature / Year: 2026Title: Regulation of STING activation by phosphoinositide and cholesterol. Authors: Jie Li / Jay Xiaojun Tan / Zhijian J Chen / Xuewu Zhang / Xiao-Chen Bai /  Abstract: Stimulator of interferon genes (STING) is an essential adaptor in the cytosolic DNA-sensing innate immune pathway. STING is activated by cyclic GMP-AMP (cGAMP) produced by the DNA sensor cGAMP ...Stimulator of interferon genes (STING) is an essential adaptor in the cytosolic DNA-sensing innate immune pathway. STING is activated by cyclic GMP-AMP (cGAMP) produced by the DNA sensor cGAMP synthase (cGAS). cGAMP-induced high-order oligomerization and translocation of STING from the endoplasmic reticulum to the Golgi and post-Golgi vesicles are critical for STING activation. Other studies have shown that phosphatidylinositol phosphates (PtdInsPs) and cholesterol also have important roles in STING activation, but the underlying mechanisms remain unclear. Here we demonstrate that cGAMP-induced high-order oligomerization of STING is enhanced strongly by phosphatidylinositol 3,5-bisphosphate (PtdIns(3,5)P and PtdIns(4,5)P, and by PtdIns4P to a lesser extent. Our cryo-electron microscopy structures reveal that PtdInsPs together with cholesterol bind at the interface between STING dimers, directly promoting the high-order oligomerization. The structures also provide an explanation for the preference of the STING oligomer to different PtdInsPs. Mutational and biochemical analyses confirm the binding modes of PtdInsPs and cholesterol and their roles in STING activation. Our findings shed light on the regulatory mechanisms of STING mediated by specific lipids, which may underlie the role of intracellular trafficking in dictating STING signalling. | ||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_46697.map.gz emd_46697.map.gz | 13.1 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-46697-v30.xml emd-46697-v30.xml emd-46697.xml emd-46697.xml | 22.7 KB 22.7 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_46697_fsc.xml emd_46697_fsc.xml | 11.7 KB | Display |  FSC data file FSC data file |
| Images |  emd_46697.png emd_46697.png | 102.9 KB | ||
| Filedesc metadata |  emd-46697.cif.gz emd-46697.cif.gz | 6.5 KB | ||
| Others |  emd_46697_half_map_1.map.gz emd_46697_half_map_1.map.gz emd_46697_half_map_2.map.gz emd_46697_half_map_2.map.gz | 104.6 MB 104.5 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-46697 http://ftp.pdbj.org/pub/emdb/structures/EMD-46697 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-46697 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-46697 | HTTPS FTP |
-Related structure data
| Related structure data |  9datMC  9danC  9davC  9dawC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_46697.map.gz / Format: CCP4 / Size: 134.6 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_46697.map.gz / Format: CCP4 / Size: 134.6 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.788 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: #2
| File | emd_46697_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #1
| File | emd_46697_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : STING oligomer bound to its ligand cGAMP, a synthetic activator n...
| Entire | Name: STING oligomer bound to its ligand cGAMP, a synthetic activator named STG2, Cholesteryl hemisuccinate and PI(4,5)P2 |
|---|---|
| Components |
|
-Supramolecule #1: STING oligomer bound to its ligand cGAMP, a synthetic activator n...
| Supramolecule | Name: STING oligomer bound to its ligand cGAMP, a synthetic activator named STG2, Cholesteryl hemisuccinate and PI(4,5)P2 type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Macromolecule #1: Stimulator of interferon genes protein
| Macromolecule | Name: Stimulator of interferon genes protein / type: protein_or_peptide / ID: 1 / Number of copies: 4 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 39.553398 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MPHSSLHPSI PCPRGHGAQK AALVLLSACL VTLWGLGEPP EHTLRYLVLH LASLQLGLLL NGVCSLAEEL RHIHSRYRGS YWRTVRACL GCPLRRGALL LLSIYFYYSL PNAVGPPFTW MLALLGLSQA LNILLGLKGL APAEISAVCE KGNFNVAHGL A WSYYIGYL ...String: MPHSSLHPSI PCPRGHGAQK AALVLLSACL VTLWGLGEPP EHTLRYLVLH LASLQLGLLL NGVCSLAEEL RHIHSRYRGS YWRTVRACL GCPLRRGALL LLSIYFYYSL PNAVGPPFTW MLALLGLSQA LNILLGLKGL APAEISAVCE KGNFNVAHGL A WSYYIGYL RLILPELQAR IRTYNQHYNN LLRGAVSQRL YILLPLDCGV PDNLSMADPN IRFLDKLPQQ TGDRAGIKDR VY SNSIYEL LENGQRAGTC VLEYATPLQT LFAMSQYSQA GFSREDRLEQ AKLFCRTLED ILADAPESQN NCRLIAYQEP ADD SSFSLS QEVLRHLRQE EKEEVTVGTS SGLEVLFQ UniProtKB: Stimulator of interferon genes protein |
-Macromolecule #2: 4-({[4-(2-tert-butyl-5,5-dimethyl-1,3-dioxan-2-yl)phenyl]methyl}a...
| Macromolecule | Name: 4-({[4-(2-tert-butyl-5,5-dimethyl-1,3-dioxan-2-yl)phenyl]methyl}amino)-3-methoxybenzoic acid type: ligand / ID: 2 / Number of copies: 2 / Formula: Y6H |
|---|---|
| Molecular weight | Theoretical: 427.533 Da |
| Chemical component information | 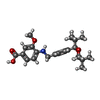 ChemComp-Y6H: |
-Macromolecule #3: (2R)-3-{[(R)-hydroxy{[(1S,2R,3R,4S,5S,6R)-2,4,6-trihydroxy-3,5-bi...
| Macromolecule | Name: (2R)-3-{[(R)-hydroxy{[(1S,2R,3R,4S,5S,6R)-2,4,6-trihydroxy-3,5-bis(phosphonooxy)cyclohexyl]oxy}phosphoryl]oxy}propane-1,2-diyl di[(9Z)-octadec-9-enoate] type: ligand / ID: 3 / Number of copies: 2 / Formula: A1BBH |
|---|---|
| Molecular weight | Theoretical: 1.023066 KDa |
-Macromolecule #4: CHOLESTEROL HEMISUCCINATE
| Macromolecule | Name: CHOLESTEROL HEMISUCCINATE / type: ligand / ID: 4 / Number of copies: 2 / Formula: Y01 |
|---|---|
| Molecular weight | Theoretical: 486.726 Da |
| Chemical component information |  ChemComp-Y01: |
-Macromolecule #5: cGAMP
| Macromolecule | Name: cGAMP / type: ligand / ID: 5 / Number of copies: 2 / Formula: 1SY |
|---|---|
| Molecular weight | Theoretical: 674.411 Da |
| Chemical component information | 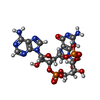 ChemComp-1SY: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.4 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Average electron dose: 1.2 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.5 µm / Nominal defocus min: 1.0 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller











 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)






































