+Search query
-Structure paper
| Title | Isolation and escape mapping of broadly neutralizing antibodies against emerging delta-coronaviruses. |
|---|---|
| Journal, issue, pages | Immunity, Vol. 57, Issue 12, Page 2914-22927.e7, Year 2024 |
| Publish date | Dec 10, 2024 |
 Authors Authors | Megi Rexhepaj / Daniel Asarnow / Lisa Perruzza / Young-Jun Park / Barbara Guarino / Mathew Mccallum / Katja Culap / Christian Saliba / Giada Leoni / Alessio Balmelli / Courtney N Yoshiyama / Miles S Dickinson / Joel Quispe / Jack T Brown / M Alejandra Tortorici / Kaitlin R Sprouse / Ashley L Taylor / Davide Corti / Tyler N Starr / Fabio Benigni / David Veesler /   |
| PubMed Abstract | Porcine delta-coronavirus (PDCoV) spillovers were recently detected in febrile children, underscoring the recurrent zoonoses of divergent CoVs. To date, no vaccines or specific therapeutics are ...Porcine delta-coronavirus (PDCoV) spillovers were recently detected in febrile children, underscoring the recurrent zoonoses of divergent CoVs. To date, no vaccines or specific therapeutics are approved for use in humans against PDCoV. To prepare for possible future PDCoV epidemics, we isolated PDCoV spike (S)-directed monoclonal antibodies (mAbs) from humanized mice and found that two, designated PD33 and PD41, broadly neutralized a panel of PDCoV variants. Cryoelectron microscopy (cryo-EM) structures of PD33 and PD41 in complex with the S receptor-binding domain (RBD) and ectodomain trimer revealed the epitopes recognized by these mAbs, rationalizing their broad inhibitory activity. We show that both mAbs competitively interfere with host aminopeptidase N binding to neutralize PDCoV and used deep-mutational scanning epitope mapping to associate RBD antigenic sites with mAb-mediated neutralization potency. Our results indicate a PD33-PD41 mAb cocktail may heighten the barrier to escape. PD33 and PD41 are candidates for clinical advancement against future PDCoV outbreaks. |
 External links External links |  Immunity / Immunity /  PubMed:39488210 / PubMed:39488210 /  PubMed Central PubMed Central |
| Methods | EM (single particle) |
| Resolution | 2.6 - 3.0 Å |
| Structure data | EMDB-44103, PDB-9b2c: EMDB-46804, PDB-9dez: EMDB-46805, PDB-9df0:  EMDB-46806: PDCoV S SD2018/300 Apo (Class I)  EMDB-46814: PDCoV S SD2018/300 with one PD41 Fab bound (Class II)  EMDB-46815: PDCoV S SD2018/300 with one PD41 Fab bound (Class III)  EMDB-46816: PDCoV S SD2018/300 with two PD41 Fabs bound (Class IV) |
| Chemicals |  ChemComp-NAG: 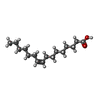 ChemComp-PAM: |
| Source |
|
 Keywords Keywords | VIRAL PROTEIN/IMMUNE SYSTEM / Spike glycoprotein / fusion protein / Structural Genomics / Seattle Structural Genomics Center for Infectious Disease / SSGCID / viral neutralization / inhibitor / VIRAL PROTEIN-IMMUNE SYSTEM complex / VIRAL PROTEIN / coronavirus / spike / antibody / PDCoV / PD41 / complex / Fab / antigen |
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About Yorodumi Papers
About Yorodumi Papers









 Homo sapiens (human)
Homo sapiens (human) porcine deltacoronavirus
porcine deltacoronavirus
