+Search query
-Structure paper
| Title | Structure of the ATP-driven methyl-coenzyme M reductase activation complex. |
|---|---|
| Journal, issue, pages | Nature, Vol. 642, Issue 8068, Page 814-821, Year 2025 |
| Publish date | Apr 16, 2025 |
 Authors Authors | Fidel Ramírez-Amador / Sophia Paul / Anuj Kumar / Christian Lorent / Sebastian Keller / Stefan Bohn / Thinh Nguyen / Stefano Lometto / Dennis Vlegels / Jörg Kahnt / Darja Deobald / Frank Abendroth / Olalla Vázquez / Georg Hochberg / Silvan Scheller / Sven T Stripp / Jan Michael Schuller /   |
| PubMed Abstract | Methyl-coenzyme M reductase (MCR) is the enzyme responsible for nearly all biologically generated methane. Its active site comprises coenzyme F, a porphyrin-based cofactor with a central nickel ion ...Methyl-coenzyme M reductase (MCR) is the enzyme responsible for nearly all biologically generated methane. Its active site comprises coenzyme F, a porphyrin-based cofactor with a central nickel ion that is active exclusively in the Ni(I) state. How methanogenic archaea perform the reductive activation of F represents a major gap in our understanding of one of the most ancient bioenergetic systems in nature. Here we purified and characterized the MCR activation complex from Methanococcus maripaludis. McrC, a small subunit encoded in the mcr operon, co-purifies with the methanogenic marker proteins Mmp7, Mmp17, Mmp3 and the A2 component. We demonstrated that this complex can activate MCR in vitro in a strictly ATP-dependent manner, enabling the formation of methane. In addition, we determined the cryo-electron microscopy structure of the MCR activation complex exhibiting different functional states with local resolutions reaching 1.8-2.1 Å. Our data revealed three complex iron-sulfur clusters that formed an electron transfer pathway towards F. Topology and electron paramagnetic resonance spectroscopy analyses indicate that these clusters are similar to the [8Fe-9S-C] cluster, a maturation intermediate of the catalytic cofactor in nitrogenase. Altogether, our findings offer insights into the activation mechanism of MCR and prospects on the early evolution of nitrogenase. |
 External links External links |  Nature / Nature /  PubMed:40240609 / PubMed:40240609 /  PubMed Central PubMed Central |
| Methods | EM (single particle) |
| Resolution | 2.14 - 2.78 Å |
| Structure data | EMDB-19787, PDB-8s7v: EMDB-19788, PDB-8s7x: EMDB-51767, PDB-9h1l: |
| Chemicals | 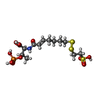 ChemComp-SHT: 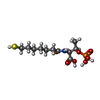 ChemComp-TP7: 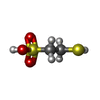 ChemComp-COM: 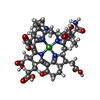 ChemComp-F43:  ChemComp-S5Q:  ChemComp-ZN:  ChemComp-ATP:  ChemComp-MG: |
| Source |
|
 Keywords Keywords | OXIDOREDUCTASE / Methyl-coenzyme M reductase / activation complex / ATPase / Iron-sulfur clusters |
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About Yorodumi Papers
About Yorodumi Papers









 methanococcus maripaludis (archaea)
methanococcus maripaludis (archaea)