+Search query
-Structure paper
| Title | Structures of the AMPA receptor in complex with its auxiliary subunit cornichon. |
|---|---|
| Journal, issue, pages | Science, Vol. 366, Issue 6470, Page 1259-1263, Year 2019 |
| Publish date | Dec 6, 2019 |
 Authors Authors | Terunaga Nakagawa /  |
| PubMed Abstract | In the brain, AMPA-type glutamate receptors (AMPARs) form complexes with their auxiliary subunits and mediate the majority of fast excitatory neurotransmission. Signals transduced by these complexes ...In the brain, AMPA-type glutamate receptors (AMPARs) form complexes with their auxiliary subunits and mediate the majority of fast excitatory neurotransmission. Signals transduced by these complexes are critical for synaptic plasticity, learning, and memory. The two major categories of AMPAR auxiliary subunits are transmembrane AMPAR regulatory proteins (TARPs) and cornichon homologs (CNIHs); these subunits share little homology and play distinct roles in controlling ion channel gating and trafficking of AMPAR. Here, I report high-resolution cryo-electron microscopy structures of AMPAR in complex with CNIH3. Contrary to its predicted membrane topology, CNIH3 lacks an extracellular domain and instead contains four membrane-spanning helices. The protein-protein interaction interface that dictates channel modulation and the lipids surrounding the complex are revealed. These structures provide insights into the molecular mechanism for ion channel modulation and assembly of AMPAR/CNIH3 complexes. |
 External links External links |  Science / Science /  PubMed:31806817 / PubMed:31806817 /  PubMed Central PubMed Central |
| Methods | EM (single particle) |
| Resolution | 2.97 - 4.4 Å |
| Structure data | EMDB-20330, PDB-6peq:  EMDB-20332: EMDB-20654: NTD of GluA2 in complex with CNIH3 - with antagonist ZK200775 - in pseudo-symmetric global conformation. EMDB-20666, PDB-6u6i:  EMDB-20717: EMDB-20727, PDB-6ucb:  EMDB-20732: EMDB-20733, PDB-6ud4: EMDB-20734, PDB-6ud8: |
| Chemicals | 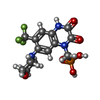 ChemComp-ZK1:  ChemComp-OLC: 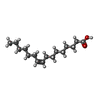 ChemComp-PAM:  ChemComp-CLR:  ChemComp-NAG: |
| Source |
|
 Keywords Keywords | TRANSPORT PROTEIN / ionotropic glutamate receptor / AMPA receptor / cornichon / auxiliary subunit / ion channel / ligand gated ion channel / synaptic transmission / excitatory synaptic transmission / neurotransmitter receptor / stargazin / TARP / ZK200775 / lipid / MPQX |
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About Yorodumi Papers
About Yorodumi Papers
















