[English] 日本語
 Yorodumi
Yorodumi- EMDB-20717: GluA2 in complex with complex full-length in AS, bound to antagon... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-20717 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | GluA2 in complex with complex full-length in AS, bound to antagonist ZK200775 | |||||||||
 Map data Map data | GluA2/CNIH3 complex full-length in AS, bound to antagonist ZK200775 | |||||||||
 Sample Sample |
| |||||||||
| Function / homology |  Function and homology information Function and homology informationCargo concentration in the ER / COPII-mediated vesicle transport / regulation of AMPA receptor activity / channel regulator activity / spine synapse / dendritic spine neck / dendritic spine cytoplasm / dendritic spine head / cellular response to amine stimulus / Activation of AMPA receptors ...Cargo concentration in the ER / COPII-mediated vesicle transport / regulation of AMPA receptor activity / channel regulator activity / spine synapse / dendritic spine neck / dendritic spine cytoplasm / dendritic spine head / cellular response to amine stimulus / Activation of AMPA receptors / neurotransmitter receptor localization to postsynaptic specialization membrane / ligand-gated monoatomic cation channel activity / perisynaptic space / Trafficking of GluR2-containing AMPA receptors / response to lithium ion / AMPA glutamate receptor activity / AMPA glutamate receptor clustering / kainate selective glutamate receptor activity / immunoglobulin binding / AMPA glutamate receptor complex / regulation of receptor recycling / extracellularly glutamate-gated ion channel activity / cellular response to glycine / ionotropic glutamate receptor complex / asymmetric synapse / Unblocking of NMDA receptors, glutamate binding and activation / glutamate receptor binding / conditioned place preference / positive regulation of synaptic transmission / response to fungicide / regulation of synaptic transmission, glutamatergic / extracellular ligand-gated monoatomic ion channel activity / cytoskeletal protein binding / vesicle-mediated transport / glutamate-gated receptor activity / cellular response to brain-derived neurotrophic factor stimulus / regulation of long-term synaptic depression / somatodendritic compartment / glutamate-gated calcium ion channel activity / presynaptic active zone membrane / excitatory synapse / ionotropic glutamate receptor signaling pathway / ionotropic glutamate receptor binding / dendrite membrane / dendrite cytoplasm / ligand-gated monoatomic ion channel activity involved in regulation of presynaptic membrane potential / positive regulation of excitatory postsynaptic potential / dendritic shaft / SNARE binding / synaptic membrane / PDZ domain binding / establishment of protein localization / protein tetramerization / transmitter-gated monoatomic ion channel activity involved in regulation of postsynaptic membrane potential / synaptic transmission, glutamatergic / regulation of membrane potential / cerebral cortex development / receptor internalization / postsynaptic density membrane / modulation of chemical synaptic transmission / Schaffer collateral - CA1 synapse / terminal bouton / synaptic vesicle / long-term synaptic potentiation / amyloid-beta binding / synaptic vesicle membrane / growth cone / presynapse / signaling receptor activity / scaffold protein binding / presynaptic membrane / dendritic spine / chemical synaptic transmission / perikaryon / postsynaptic membrane / neuron projection / postsynaptic density / signaling receptor binding / external side of plasma membrane / axon / neuronal cell body / synapse / dendrite / protein kinase binding / protein-containing complex binding / glutamatergic synapse / cell surface / endoplasmic reticulum / protein-containing complex / membrane / identical protein binding / plasma membrane Similarity search - Function | |||||||||
| Biological species |   | |||||||||
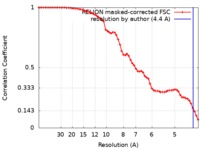
| Method | single particle reconstruction / cryo EM / Resolution: 4.4 Å | |||||||||
 Authors Authors | Nakagawa T | |||||||||
| Funding support |  United States, 1 items United States, 1 items
| |||||||||
 Citation Citation |  Journal: Science / Year: 2019 Journal: Science / Year: 2019Title: Structures of the AMPA receptor in complex with its auxiliary subunit cornichon. Authors: Terunaga Nakagawa /  Abstract: In the brain, AMPA-type glutamate receptors (AMPARs) form complexes with their auxiliary subunits and mediate the majority of fast excitatory neurotransmission. Signals transduced by these complexes ...In the brain, AMPA-type glutamate receptors (AMPARs) form complexes with their auxiliary subunits and mediate the majority of fast excitatory neurotransmission. Signals transduced by these complexes are critical for synaptic plasticity, learning, and memory. The two major categories of AMPAR auxiliary subunits are transmembrane AMPAR regulatory proteins (TARPs) and cornichon homologs (CNIHs); these subunits share little homology and play distinct roles in controlling ion channel gating and trafficking of AMPAR. Here, I report high-resolution cryo-electron microscopy structures of AMPAR in complex with CNIH3. Contrary to its predicted membrane topology, CNIH3 lacks an extracellular domain and instead contains four membrane-spanning helices. The protein-protein interaction interface that dictates channel modulation and the lipids surrounding the complex are revealed. These structures provide insights into the molecular mechanism for ion channel modulation and assembly of AMPAR/CNIH3 complexes. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_20717.map.gz emd_20717.map.gz | 20.7 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-20717-v30.xml emd-20717-v30.xml emd-20717.xml emd-20717.xml | 15.6 KB 15.6 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_20717_fsc.xml emd_20717_fsc.xml | 6.4 KB | Display |  FSC data file FSC data file |
| Images |  emd_20717.png emd_20717.png | 203.1 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-20717 http://ftp.pdbj.org/pub/emdb/structures/EMD-20717 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-20717 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-20717 | HTTPS FTP |
-Related structure data
| Related structure data |  6peqC  6u5sC  6u6iC  6ucbC  6ud4C  6ud8C C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_20717.map.gz / Format: CCP4 / Size: 22.2 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_20717.map.gz / Format: CCP4 / Size: 22.2 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | GluA2/CNIH3 complex full-length in AS, bound to antagonist ZK200775 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 2.132 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : GluA2 in complex with CNIH3 full length at 4:4 stoichiometry
| Entire | Name: GluA2 in complex with CNIH3 full length at 4:4 stoichiometry |
|---|---|
| Components |
|
-Supramolecule #1: GluA2 in complex with CNIH3 full length at 4:4 stoichiometry
| Supramolecule | Name: GluA2 in complex with CNIH3 full length at 4:4 stoichiometry type: complex / ID: 1 / Parent: 0 / Macromolecule list: all Details: Bound to antagonist ZK200775 (MPQX). Lipid densities are observed. |
|---|---|
| Molecular weight | Theoretical: 470 KDa |
-Supramolecule #2: Glutamate receptor 2
| Supramolecule | Name: Glutamate receptor 2 / type: complex / ID: 2 / Parent: 1 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:  |
| Recombinant expression | Organism:  Homo sapiens (human) / Recombinant strain: HEK Homo sapiens (human) / Recombinant strain: HEK |
-Supramolecule #3: Protein cornichon homolog 3
| Supramolecule | Name: Protein cornichon homolog 3 / type: complex / ID: 3 / Parent: 1 / Macromolecule list: #2 |
|---|---|
| Source (natural) | Organism:  |
| Recombinant expression | Organism:  Homo sapiens (human) / Recombinant strain: HEK Homo sapiens (human) / Recombinant strain: HEK |
-Macromolecule #1: Glutamate receptor 2
| Macromolecule | Name: Glutamate receptor 2 / type: protein_or_peptide / ID: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MQKIMHISVL LSPVLWGLIF GVSSNSIQIG GLFPRGADQE YSAFRVGMVQ FSTSEFRLTP HIDNLEVANS FAVTNAFCSQ FSRGVYAIFG FYDKKSVNTI TSFCGTLHVS FITPSFPTDG THPFVIQMRP DLKGALLSLI EYYQWDKFAY LYDSDRGLST LQAVLDSAAE ...String: MQKIMHISVL LSPVLWGLIF GVSSNSIQIG GLFPRGADQE YSAFRVGMVQ FSTSEFRLTP HIDNLEVANS FAVTNAFCSQ FSRGVYAIFG FYDKKSVNTI TSFCGTLHVS FITPSFPTDG THPFVIQMRP DLKGALLSLI EYYQWDKFAY LYDSDRGLST LQAVLDSAAE KKWQVTAINV GNINNDKKDE TYRSLFQDLE LKKERRVILD CERDKVNDIV DQVITIGKHV KGYHYIIANL GFTDGDLLKI QFGGANVSGF QIVDYDDSLV SKFIERWSTL EEKEYPGAHT ATIKYTSALT YDAVQVMTEA FRNLRKQRIE ISRRGNAGDC LANPAVPWGQ GVEIERALKQ VQVEGLSGNI KFDQNGKRIN YTINIMELKT NGPRKIGYWS EVDKMVVTLT ELPSGNDTSG LENKTVVVTT ILESPYVMMK KNHEMLEGNE RYEGYCVDLA AEIAKHCGFK YKLTIVGDGK YGARDADTKI WNGMVGELVY GKADIAIAPL TITLVREEVI DFSKPFMSLG ISIMIKKPQK SKPGVFSFLD PLAYEIWMCI VFAYIGVSVV LFLVSRFSPY EWHTEEFEDG RETQSSESTN EFGIFNSLWF SLGAFMRQGC DISPRSLSGR IVGGVWWFFT LIIISSYTAN LAAFLTVERM VSPIESAEDL SKQTEIAYGT LDSGSTKEFF RRSKIAVFDK MWTYMRSAEP SVFVRTTAEG VARVRKSKGK YAYLLESTMN EYIEQRKPCD TMKVGGNLDS KGYGIATPKG SSLGNAVNLA VLKLNEQGLL DKLKNKWWYD KGECGSGGGD SKEKTSALSL SNVAGVFYIL VGGLGLAMLV ALIEFCYKSR AEAKRMKVAK NPQNINPSSS QNSQNFATDY KDDDDKEGYN VYGIESVKI |
-Macromolecule #2: Protein cornichon homolog 3
| Macromolecule | Name: Protein cornichon homolog 3 / type: protein_or_peptide / ID: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MAFTFAAFCY MLSLVLCAAL IFFAIWHIIA FDELRTDFKS PIDQCNPVHA RERLRNIERI CFLLRKLVLP EYSIHSLFCI MFLCAQEWLT LGLNVPLLFY HFWRYFHCPA DSSELAYDPP VVMNADTLSY CQKEAWCKLA FYLLSFFYYL YCMIYTLVSS GGRGGTETSQ VAPA |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 8 Component:
| ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277.15 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Specialist optics | Energy filter - Name: GIF Bioquantum / Energy filter - Slit width: 20 eV |
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Number grids imaged: 1 / Number real images: 11340 / Average exposure time: 6.0 sec. / Average electron dose: 58.5 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 2.0 µm / Nominal defocus min: 0.8 µm / Nominal magnification: 81000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Refinement | Space: REAL |
|---|
 Movie
Movie Controller
Controller




















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)