+Search query
-Structure paper
| Title | Maturation Pathway from Germline to Broad HIV-1 Neutralizer of a CD4-Mimic Antibody. |
|---|---|
| Journal, issue, pages | Cell, Vol. 165, Issue 2, Page 449-463, Year 2016 |
| Publish date | Apr 7, 2016 |
 Authors Authors | Mattia Bonsignori / Tongqing Zhou / Zizhang Sheng / Lei Chen / Feng Gao / M Gordon Joyce / Gabriel Ozorowski / Gwo-Yu Chuang / Chaim A Schramm / Kevin Wiehe / S Munir Alam / Todd Bradley / Morgan A Gladden / Kwan-Ki Hwang / Sheelah Iyengar / Amit Kumar / Xiaozhi Lu / Kan Luo / Michael C Mangiapani / Robert J Parks / Hongshuo Song / Priyamvada Acharya / Robert T Bailer / Allen Cao / Aliaksandr Druz / Ivelin S Georgiev / Young D Kwon / Mark K Louder / Baoshan Zhang / Anqi Zheng / Brenna J Hill / Rui Kong / Cinque Soto / / James C Mullikin / Daniel C Douek / David C Montefiori / Michael A Moody / George M Shaw / Beatrice H Hahn / Garnett Kelsoe / Peter T Hraber / Bette T Korber / Scott D Boyd / Andrew Z Fire / Thomas B Kepler / Lawrence Shapiro / Andrew B Ward / John R Mascola / Hua-Xin Liao / Peter D Kwong / Barton F Haynes /  |
| PubMed Abstract | Antibodies with ontogenies from VH1-2 or VH1-46-germline genes dominate the broadly neutralizing response against the CD4-binding site (CD4bs) on HIV-1. Here, we define with longitudinal sampling ...Antibodies with ontogenies from VH1-2 or VH1-46-germline genes dominate the broadly neutralizing response against the CD4-binding site (CD4bs) on HIV-1. Here, we define with longitudinal sampling from time-of-infection the development of a VH1-46-derived antibody lineage that matured to neutralize 90% of HIV-1 isolates. Structures of lineage antibodies CH235 (week 41 from time-of-infection, 18% breadth), CH235.9 (week 152, 77%), and CH235.12 (week 323, 90%) demonstrated the maturing epitope to focus on the conformationally invariant portion of the CD4bs. Similarities between CH235 lineage and five unrelated CD4bs lineages in epitope focusing, length-of-time to develop breadth, and extraordinary level of somatic hypermutation suggested commonalities in maturation among all CD4bs antibodies. Fortunately, the required CH235-lineage hypermutation appeared substantially guided by the intrinsic mutability of the VH1-46 gene, which closely resembled VH1-2. We integrated our CH235-lineage findings with a second broadly neutralizing lineage and HIV-1 co-evolution to suggest a vaccination strategy for inducing both lineages. |
 External links External links |  Cell / Cell /  PubMed:26949186 / PubMed:26949186 /  PubMed Central PubMed Central |
| Methods | EM (single particle) / X-ray diffraction |
| Resolution | 1.86 - 25.0 Å |
| Structure data |  EMDB-8078:  EMDB-8079:  EMDB-8080:  EMDB-8081:  EMDB-8082:  EMDB-8083:  PDB-5f96:  PDB-5f9o:  PDB-5f9w: |
| Chemicals |  ChemComp-NAG:  ChemComp-EPE:  ChemComp-HOH: 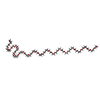 ChemComp-15P:  ChemComp-PO4: |
| Source |
|
 Keywords Keywords | IMMUNE SYSTEM / HIV-1 / Antibody / CH235 lineage / VH1-46 germline / antibody evolution HIV-1 broadly neutralizing CD4 binding site / neutralizing antibody / CH235 |
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About Yorodumi Papers
About Yorodumi Papers




 human immunodeficiency virus 1
human immunodeficiency virus 1 homo sapiens (human)
homo sapiens (human)