+Search query
-Structure paper
| Title | A Surfactant Cocktail Overcomes Air-Water Interface Artifacts in Single-Particle CryoEM. |
|---|---|
| Journal, issue, pages | bioRxiv, Year 2026 |
| Publish date | Mar 18, 2026 |
 Authors Authors | Suzanne E Enos / Brian D Cook / Hamidreza Rahmani / Sarah M Narehood / Yizhou Li / Inga C Kuschnerus / Trevor H Redford / Peter Dukakis / Daniel Ji / Maxwell J Bachochin / Danielle A Grotjahn / Mark A Herzik /  |
| PubMed Abstract | Single-particle cryogenic electron microscopy (cryoEM) is a widely used technique for structure determination of biomacromolecules to near-atomic resolution. Random distributions of these molecules ...Single-particle cryogenic electron microscopy (cryoEM) is a widely used technique for structure determination of biomacromolecules to near-atomic resolution. Random distributions of these molecules in vitrified ice are necessary to accumulate enough two-dimensional views to generate a complete three-dimensional (3-D) reconstruction. However, interactions between the sample and the air-water interface (AWI) that occur during vitrification often bias the views of the sample, a phenomenon termed preferred orientation, limiting our ability to obtain 3-D reconstructions. Surfactants are often used as sample additives to prevent AWI-induced deterioration, but no general strategy exists for surfactant choice, requiring laborious screening for each sample. To circumvent these issues, we developed SurfACT, a cocktail of diverse surfactants with distinct physicochemical properties that limits AWI-dependent sample denaturation and orientation bias, while mitigating individual surfactant-specific drawbacks. Here we demonstrate SurfACT's effectiveness with four proteins plagued by AWI-induced issues, including two species of hemagglutinin (HA), molybdenum-iron protein (MoFeP) from the nitrogenase enzyme, and aldolase. All four samples show drastically improved viewing distribution and map completeness when SurfACT is applied. Cryogenic electron tomography demonstrates that SurfACT redistributes particles from the AWI into the bulk ice, driving signal recovery and inhibiting denaturation. This versatile sample additive minimizes sample-specific screening and expands the capabilities and range of suitable samples for cryoEM. |
 External links External links |  bioRxiv / bioRxiv /  PubMed:41889805 / PubMed:41889805 /  PubMed Central PubMed Central |
| Methods | EM (single particle) / EM (subtomogram averaging) |
| Resolution | 1.97 - 10.11 Å |
| Structure data |  EMDB-75842: Influenza A virus Hemagglutinin (A/Darwin/6/2021 H3N2) (C1 symmetry) determined using the SPT Labtech chameleon in the presence of 0x SurfACT  EMDB-75843: Influenza A virus Hemagglutinin (A/Darwin/6/2021 H3N2) (C3 symmetry) determined using the SPT Labtech chameleon in the presence of 0x SurfACT EMDB-75844, PDB-11ms: EMDB-75845, PDB-11mt: EMDB-75846, PDB-11mu: EMDB-75847, PDB-11mv:  EMDB-75848: Influenza A virus Hemagglutinin (A/California/04/2009 H1N1), E47K HA2 stabilizing mutation (C1 symmetry) determined using the SPT Labtech chameleon in the presence of 0x SurfACT  EMDB-75849: Influenza A virus Hemagglutinin (A/California/04/2009 H1N1), E47K HA2 stabilizing mutation (C3 symmetry) determined using the SPT Labtech chameleon in the presence of 0x SurfACT EMDB-75851, PDB-11mx: EMDB-75853, PDB-11mz: EMDB-75854, PDB-11na: EMDB-75855, PDB-11nb: EMDB-75856, PDB-11nc: EMDB-75857, PDB-11nd: EMDB-75858, PDB-11ne: EMDB-75859, PDB-11nf: EMDB-75860, PDB-11nh: EMDB-75861, PDB-11ni: EMDB-75862, PDB-11nj: EMDB-75863, PDB-11nk: EMDB-75864, PDB-11nl: EMDB-75865, PDB-11nm: EMDB-75866, PDB-11nn: EMDB-75867, PDB-11no: EMDB-75868, PDB-11np: EMDB-75869, PDB-11nr: EMDB-75870, PDB-11nt: EMDB-75871, PDB-11nu: EMDB-75873, PDB-11nw: EMDB-75874, PDB-11nx: EMDB-75878, PDB-11ob: EMDB-75879, PDB-11oc:  EMDB-75901: Influenza A virus Hemagglutinin (A/Darwin/6/2021 H3N2) (C3 symmetry; subtomogram averaged) determined using the SPT Labtech chameleon in the presence of 0x SurfACT  EMDB-75902: Influenza A virus Hemagglutinin (A/Darwin/6/2021 H3N2) (C3 symmetry; subtomogram averaged) determined using the SPT Labtech chameleon in the presence of 0.25x SurfACT  EMDB-75903: Rabbit muscle Aldolase (C1 symmetry; subtomogram averaged) determined using the SPT Labtech chameleon in the presence of 0x SurfACT  EMDB-75904: Rabbit muscle Aldolase (C1 symmetry; subtomogram averaged) determined using the SPT Labtech chameleon in the presence of 0.25x SurfACT  EMDB-75905: Rabbit muscle Aldolase (C1 symmetry; subtomogram averaged; top air-water interface) determined using the SPT Labtech chameleon in the presence of 0.25x SurfACT  EMDB-75906: Rabbit muscle Aldolase (C1 symmetry; subtomogram averaged; center ice) determined using the SPT Labtech chameleon in the presence of 0.25x SurfACT  EMDB-75907: Rabbit muscle Aldolase (C1 symmetry; subtomogram averaged; bottom air-water interface) determined using the SPT Labtech chameleon in the presence of 0.25x SurfACT  EMDB-75908: Rabbit muscle Aldolase (C1 symmetry; subtomogram averaged) determined using the SPT Labtech chameleon in the presence of 1x SurfACT |
| Chemicals |  ChemComp-HOH: 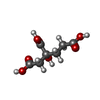 ChemComp-HCA: 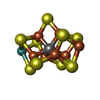 ChemComp-ICS:  ChemComp-CLF:  ChemComp-FE: |
| Source |
|
 Keywords Keywords | VIRAL PROTEIN / Hemagglutinin / Influenza A virus / H3N2 / Viral Entry / Trimer / Glycoprotein / Membrane Fusion / H1N1 / OXIDOREDUCTASE / Nitrogenase / FeMoCo / nitrogen / P-cluster / METAL BINDING PROTEIN / LYASE / Glycolysis / Fructose-bisphosphate aldolase / Carbon-carbon lyase |
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About Yorodumi Papers
About Yorodumi Papers




























































 influenza a virus
influenza a virus azotobacter vinelandii (bacteria)
azotobacter vinelandii (bacteria)
