[English] 日本語
 Yorodumi
Yorodumi- EMDB-75858: Azotobacter vinelandii MoFeP (C1 symmetry) determined using the S... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | ||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Azotobacter vinelandii MoFeP (C1 symmetry) determined using the SPT Labtech chameleon in the presence of 1x SurfACT | ||||||||||||||||||
 Map data Map data | Azotobacter vinelandii MoFeP (C1 symmetry) determined using the SPT Labtech chameleon in the presence of 1x SurfACT | ||||||||||||||||||
 Sample Sample |
| ||||||||||||||||||
 Keywords Keywords | Nitrogenase / FeMoCo / nitrogen / P-cluster / METAL BINDING PROTEIN / OXIDOREDUCTASE | ||||||||||||||||||
| Function / homology |  Function and homology information Function and homology informationnitrogen fixation / molybdenum-iron nitrogenase complex / nitrogenase / nitrogenase activity / iron-sulfur cluster binding / ATP binding / metal ion binding Similarity search - Function | ||||||||||||||||||
| Biological species |  Azotobacter vinelandii (bacteria) Azotobacter vinelandii (bacteria) | ||||||||||||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 2.49 Å | ||||||||||||||||||
 Authors Authors | Enos SE / Cook BD / Rahmani H / Narehood SM / Li Y / Kuschnerus IC / Redford HT / Dukakis P / Ji D / Bachochin MJ ...Enos SE / Cook BD / Rahmani H / Narehood SM / Li Y / Kuschnerus IC / Redford HT / Dukakis P / Ji D / Bachochin MJ / Grotjahn DA / Herzik MA | ||||||||||||||||||
| Funding support |  United States, 5 items United States, 5 items
| ||||||||||||||||||
 Citation Citation |  Journal: bioRxiv / Year: 2026 Journal: bioRxiv / Year: 2026Title: A Surfactant Cocktail Overcomes Air-Water Interface Artifacts in Single-Particle CryoEM. Authors: Suzanne E Enos / Brian D Cook / Hamidreza Rahmani / Sarah M Narehood / Yizhou Li / Inga C Kuschnerus / Trevor H Redford / Peter Dukakis / Daniel Ji / Maxwell J Bachochin / Danielle A Grotjahn / Mark A Herzik /  Abstract: Single-particle cryogenic electron microscopy (cryoEM) is a widely used technique for structure determination of biomacromolecules to near-atomic resolution. Random distributions of these molecules ...Single-particle cryogenic electron microscopy (cryoEM) is a widely used technique for structure determination of biomacromolecules to near-atomic resolution. Random distributions of these molecules in vitrified ice are necessary to accumulate enough two-dimensional views to generate a complete three-dimensional (3-D) reconstruction. However, interactions between the sample and the air-water interface (AWI) that occur during vitrification often bias the views of the sample, a phenomenon termed preferred orientation, limiting our ability to obtain 3-D reconstructions. Surfactants are often used as sample additives to prevent AWI-induced deterioration, but no general strategy exists for surfactant choice, requiring laborious screening for each sample. To circumvent these issues, we developed SurfACT, a cocktail of diverse surfactants with distinct physicochemical properties that limits AWI-dependent sample denaturation and orientation bias, while mitigating individual surfactant-specific drawbacks. Here we demonstrate SurfACT's effectiveness with four proteins plagued by AWI-induced issues, including two species of hemagglutinin (HA), molybdenum-iron protein (MoFeP) from the nitrogenase enzyme, and aldolase. All four samples show drastically improved viewing distribution and map completeness when SurfACT is applied. Cryogenic electron tomography demonstrates that SurfACT redistributes particles from the AWI into the bulk ice, driving signal recovery and inhibiting denaturation. This versatile sample additive minimizes sample-specific screening and expands the capabilities and range of suitable samples for cryoEM. | ||||||||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_75858.map.gz emd_75858.map.gz | 204.1 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-75858-v30.xml emd-75858-v30.xml emd-75858.xml emd-75858.xml | 32.1 KB 32.1 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_75858_fsc.xml emd_75858_fsc.xml | 12.6 KB | Display |  FSC data file FSC data file |
| Images |  emd_75858.png emd_75858.png | 154.5 KB | ||
| Masks |  emd_75858_msk_1.map emd_75858_msk_1.map | 216 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-75858.cif.gz emd-75858.cif.gz | 8.1 KB | ||
| Others |  emd_75858_additional_1.map.gz emd_75858_additional_1.map.gz emd_75858_additional_2.map.gz emd_75858_additional_2.map.gz emd_75858_half_map_1.map.gz emd_75858_half_map_1.map.gz emd_75858_half_map_2.map.gz emd_75858_half_map_2.map.gz | 7.4 MB 108.2 MB 200.4 MB 200.4 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-75858 http://ftp.pdbj.org/pub/emdb/structures/EMD-75858 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-75858 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-75858 | HTTPS FTP |
-Related structure data
| Related structure data |  11neMC  11msC  11mtC  11muC  11mvC  11mxC  11mzC  11naC  11nbC  11ncC  11ndC  11nfC  11nhC  11niC  11njC  11nkC  11nlC  11nmC  11nnC  11noC  11npC  11nrC  11ntC  11nuC  11nwC  11nxC  11obC  11ocC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_75858.map.gz / Format: CCP4 / Size: 216 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_75858.map.gz / Format: CCP4 / Size: 216 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Azotobacter vinelandii MoFeP (C1 symmetry) determined using the SPT Labtech chameleon in the presence of 1x SurfACT | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.735 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_75858_msk_1.map emd_75858_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Additional map: Phenix RESOLVE modified EM map of Azotobacter vinelandii...
| File | emd_75858_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Phenix RESOLVE modified EM map of Azotobacter vinelandii MoFeP (C1 symmetry) determined using the SPT Labtech chameleon in the presence of 1x SurfACT | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Additional map: Unsharpened map of Azotobacter vinelandii MoFeP (C1 symmetry)...
| File | emd_75858_additional_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Unsharpened map of Azotobacter vinelandii MoFeP (C1 symmetry) determined using the SPT Labtech chameleon in the presence of 1x SurfACT | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: Half map A of Azotobacter vinelandii MoFeP determined...
| File | emd_75858_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half map A of Azotobacter vinelandii MoFeP determined using the SPT Labtech chameleon in the presence of 1x SurfACT; C1 symmetry | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: Half map B of Azotobacter vinelandii MoFeP determined...
| File | emd_75858_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half map B of Azotobacter vinelandii MoFeP determined using the SPT Labtech chameleon in the presence of 1x SurfACT; C1 symmetry | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Azotobacter vinelandii MoFeP (C1 symmetry) determined using the S...
| Entire | Name: Azotobacter vinelandii MoFeP (C1 symmetry) determined using the SPT Labtech chameleon in the presence of 1x SurfACT |
|---|---|
| Components |
|
-Supramolecule #1: Azotobacter vinelandii MoFeP (C1 symmetry) determined using the S...
| Supramolecule | Name: Azotobacter vinelandii MoFeP (C1 symmetry) determined using the SPT Labtech chameleon in the presence of 1x SurfACT type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#2 |
|---|---|
| Source (natural) | Organism:  Azotobacter vinelandii (bacteria) Azotobacter vinelandii (bacteria) |
| Molecular weight | Theoretical: 230 KDa |
-Macromolecule #1: Nitrogenase molybdenum-iron protein alpha chain
| Macromolecule | Name: Nitrogenase molybdenum-iron protein alpha chain / type: protein_or_peptide / ID: 1 / Number of copies: 2 / Enantiomer: LEVO / EC number: nitrogenase |
|---|---|
| Source (natural) | Organism:  Azotobacter vinelandii (bacteria) Azotobacter vinelandii (bacteria) |
| Molecular weight | Theoretical: 54.000559 KDa |
| Sequence | String: MSREEVESLI QEVLEVYPEK ARKDRNKHLA VNDPAVTQSK KCIISNKKSQ PGLMTIRGCA YAGSKGVVWG PIKDMIHISH GPVGCGQYS RAGRRNYYIG TTGVNAFVTM NFTSDFQEKD IVFGGDKKLA KLIDEVETLF PLNKGISVQS ECPIGLIGDD I ESVSKVKG ...String: MSREEVESLI QEVLEVYPEK ARKDRNKHLA VNDPAVTQSK KCIISNKKSQ PGLMTIRGCA YAGSKGVVWG PIKDMIHISH GPVGCGQYS RAGRRNYYIG TTGVNAFVTM NFTSDFQEKD IVFGGDKKLA KLIDEVETLF PLNKGISVQS ECPIGLIGDD I ESVSKVKG AELSKTIVPV RCEGFRGVSQ SLGHHIANDA VRDWVLGKRD EDTTFASTPY DVAIIGDYNI GGDAWSSRIL LE EMGLRCV AQWSGDGSIS EIELTPKVKL NLVHCYRSMN YISRHMEEKY GIPWMEYNFF GPTKTIESLR AIAAKFDESI QKK CEEVIA KYKPEWEAVV AKYRPRLEGK RVMLYIGGLR PRHVIGAYED LGMEVVGTGY EFAHNDDYDR TMKEMGDSTL LYDD VTGYE FEEFVKRIKP DLIGSGIKEK FIFQKMGIPF REMHSWDYSG PYHGFDGFAI FARDMDMTLN NPCWKKLQAP WE UniProtKB: Nitrogenase molybdenum-iron protein alpha chain |
-Macromolecule #2: Nitrogenase molybdenum-iron protein beta chain
| Macromolecule | Name: Nitrogenase molybdenum-iron protein beta chain / type: protein_or_peptide / ID: 2 / Number of copies: 2 / Enantiomer: LEVO / EC number: nitrogenase |
|---|---|
| Source (natural) | Organism:  Azotobacter vinelandii (bacteria) Azotobacter vinelandii (bacteria) |
| Molecular weight | Theoretical: 59.404684 KDa |
| Sequence | String: SQQVDKIKAS YPLFLDQDYK DMLAKKRDGF EEKYPQDKID EVFQWTTTKE YQELNFQREA LTVNPAKACQ PLGAVLCALG FEKTMPYVH GSQGCVAYFR SYFNRHFREP VSCVSDSMTE DAAVFGGQQN MKDGLQNCKA TYKPDMIAVS TTCMAEVIGD D LNAFINNS ...String: SQQVDKIKAS YPLFLDQDYK DMLAKKRDGF EEKYPQDKID EVFQWTTTKE YQELNFQREA LTVNPAKACQ PLGAVLCALG FEKTMPYVH GSQGCVAYFR SYFNRHFREP VSCVSDSMTE DAAVFGGQQN MKDGLQNCKA TYKPDMIAVS TTCMAEVIGD D LNAFINNS KKEGFIPDEF PVPFAHTPSF VGSHVTGWDN MFEGIARYFT LKSMDDKVVG SNKKINIVPG FETYLGNFRV IK RMLSEMG VGYSLLSDPE EVLDTPADGQ FRMYAGGTTQ EEMKDAPNAL NTVLLQPWHL EKTKKFVEGT WKHEVPKLNI PMG LDWTDE FLMKVSEISG QPIPASLTKE RGRLVDMMTD SHTWLHGKRF ALWGDPDFVM GLVKFLLELG CEPVHILCHN GNKR WKKAV DAILAASPYG KNATVYIGKD LWHLRSLVFT DKPDFMIGNS YGKFIQRDTL HKGKEFEVPL IRIGFPIFDR HHLHR STTL GYEGAMQILT TLVNSILERL DEETRGMQAT DYNHDLVR UniProtKB: Nitrogenase molybdenum-iron protein beta chain |
-Macromolecule #3: 3-HYDROXY-3-CARBOXY-ADIPIC ACID
| Macromolecule | Name: 3-HYDROXY-3-CARBOXY-ADIPIC ACID / type: ligand / ID: 3 / Number of copies: 2 / Formula: HCA |
|---|---|
| Molecular weight | Theoretical: 206.15 Da |
| Chemical component information | 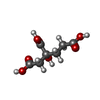 ChemComp-HCA: |
-Macromolecule #4: iron-sulfur-molybdenum cluster with interstitial carbon
| Macromolecule | Name: iron-sulfur-molybdenum cluster with interstitial carbon type: ligand / ID: 4 / Number of copies: 2 / Formula: ICS |
|---|---|
| Molecular weight | Theoretical: 787.451 Da |
| Chemical component information | 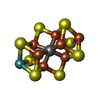 ChemComp-ICE: |
-Macromolecule #5: FE (III) ION
| Macromolecule | Name: FE (III) ION / type: ligand / ID: 5 / Number of copies: 2 / Formula: FE |
|---|---|
| Molecular weight | Theoretical: 55.845 Da |
-Macromolecule #6: FE(8)-S(7) CLUSTER
| Macromolecule | Name: FE(8)-S(7) CLUSTER / type: ligand / ID: 6 / Number of copies: 2 / Formula: CLF |
|---|---|
| Molecular weight | Theoretical: 671.215 Da |
| Chemical component information |  ChemComp-CLF: |
-Macromolecule #7: water
| Macromolecule | Name: water / type: ligand / ID: 7 / Number of copies: 1680 / Formula: HOH |
|---|---|
| Molecular weight | Theoretical: 18.015 Da |
| Chemical component information |  ChemComp-HOH: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 3.6 mg/mL | |||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 8 Component:
| |||||||||||||||||||||
| Grid | Model: Quantifoil Active R1.2/0.8 / Material: COPPER / Support film - Material: CARBON / Support film - topology: HOLEY / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 20 sec. | |||||||||||||||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 75 % / Chamber temperature: 298.15 K / Instrument: SPT LABTECH CHAMELEON Details: Samples were frozen with the SPT Labtech chameleon. |
- Electron microscopy
Electron microscopy
| Microscope | TFS KRIOS |
|---|---|
| Specialist optics | Energy filter - Name: TFS Selectris X / Energy filter - Slit width: 10 eV |
| Image recording | Film or detector model: TFS FALCON 4i (4k x 4k) / Digitization - Dimensions - Width: 4096 pixel / Digitization - Dimensions - Height: 4096 pixel / Number grids imaged: 1 / Number real images: 1080 / Average exposure time: 5.0 sec. / Average electron dose: 65.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 2.5 µm / Nominal defocus min: 1.0 µm / Nominal magnification: 165000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Initial model | PDB ID: Chain - Source name: PDB / Chain - Initial model type: experimental model |
|---|---|
| Refinement | Space: REAL / Protocol: RIGID BODY FIT / Overall B value: 49.9 |
| Output model |  PDB-11ne: |
 Movie
Movie Controller
Controller











































 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)






























































