+Search query
-Structure paper
| Title | Probing the mechanism by which the retinal G protein transducin activates its biological effector PDE6. |
|---|---|
| Journal, issue, pages | J Biol Chem, Vol. 300, Issue 2, Page 105608, Year 2024 |
| Publish date | Dec 28, 2023 |
 Authors Authors | Cody Aplin / Richard A Cerione /  |
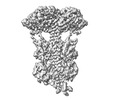
| PubMed Abstract | Phototransduction in retinal rods occurs when the G protein-coupled photoreceptor rhodopsin triggers the activation of phosphodiesterase 6 (PDE6) by GTP-bound alpha subunits of the G protein ...Phototransduction in retinal rods occurs when the G protein-coupled photoreceptor rhodopsin triggers the activation of phosphodiesterase 6 (PDE6) by GTP-bound alpha subunits of the G protein transducin (Gα). Recently, we presented a cryo-EM structure for a complex between two GTP-bound recombinant Gα subunits and native PDE6, that included a bivalent antibody bound to the C-terminal ends of Gα and the inhibitor vardenafil occupying the active sites on the PDEα and PDEβ subunits. We proposed Gα-activated PDE6 by inducing a striking reorientation of the PDEγ subunits away from the catalytic sites. However, questions remained including whether in the absence of the antibody Gα binds to PDE6 in a similar manner as observed when the antibody is present, does Gα activate PDE6 by enabling the substrate cGMP to access the catalytic sites, and how does the lipid membrane enhance PDE6 activation? Here, we demonstrate that 2:1 Gα-PDE6 complexes form with either recombinant or retinal Gα in the absence of the Gα antibody. We show that Gα binding is not necessary for cGMP nor competitive inhibitors to access the active sites; instead, occupancy of the substrate binding sites enables Gα to bind and reposition the PDE6γ subunits to promote catalytic activity. Moreover, we demonstrate by reconstituting Gα-stimulated PDE6 activity in lipid bilayer nanodiscs that the membrane-induced enhancement results from an increase in the apparent binding affinity of Gα for PDE6. These findings provide new insights into how the retinal G protein stimulates rapid catalytic turnover by PDE6 required for dim light vision. |
 External links External links |  J Biol Chem / J Biol Chem /  PubMed:38159849 / PubMed:38159849 /  PubMed Central PubMed Central |
| Methods | EM (single particle) |
| Resolution | 3.0 - 4.44 Å |
| Structure data | EMDB-42208, PDB-8ufi: EMDB-42220, PDB-8ugb: EMDB-42234, PDB-8ugs:  EMDB-42235: Bovine rod phosphodiesterase 6 bound to one retinal transducin alpha subunit  EMDB-42237: Bovine rod phosphodiesterase 6 bound to two retinal transducin alpha subunits  EMDB-42238: Bovine rod phosphodiesterase 6 bound to two chimera transducin alpha subunits without a stabilizing antibody EMDB-42358, PDB-8ulg: |
| Chemicals |  ChemComp-PCG:  ChemComp-ZN:  ChemComp-MG: 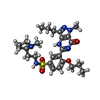 ChemComp-ZUD:  ChemComp-IBM: |
| Source |
|
 Keywords Keywords | SIGNALING PROTEIN / phosphodiesterase / GPCR effector enzyme / SIGNALING PROTEIN/INHIBITOR / inhibitor / SIGNALING PROTEIN-INHIBITOR complex |
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About Yorodumi Papers
About Yorodumi Papers