+検索条件
-Structure paper
| タイトル | The antibody response to SARS-CoV-2 Beta underscores the antigenic distance to other variants. |
|---|---|
| ジャーナル・号・ページ | Cell Host Microbe, Vol. 30, Issue 1, Page 53-68.e12, Year 2022 |
| 掲載日 | 2022年1月12日 |
 著者 著者 | Chang Liu / Daming Zhou / Rungtiwa Nutalai / Helen M E Duyvesteyn / Aekkachai Tuekprakhon / Helen M Ginn / Wanwisa Dejnirattisai / Piyada Supasa / Alexander J Mentzer / Beibei Wang / James Brett Case / Yuguang Zhao / Donal T Skelly / Rita E Chen / Sile Ann Johnson / Thomas G Ritter / Chris Mason / Tariq Malik / Nigel Temperton / Neil G Paterson / Mark A Williams / David R Hall / Daniel K Clare / Andrew Howe / Philip J R Goulder / Elizabeth E Fry / Michael S Diamond / Juthathip Mongkolsapaya / Jingshan Ren / David I Stuart / Gavin R Screaton /    |
| PubMed 要旨 | Alpha-B.1.1.7, Beta-B.1.351, Gamma-P.1, and Delta-B.1.617.2 variants of SARS-CoV-2 express multiple mutations in the spike protein (S). These may alter the antigenic structure of S, causing escape ...Alpha-B.1.1.7, Beta-B.1.351, Gamma-P.1, and Delta-B.1.617.2 variants of SARS-CoV-2 express multiple mutations in the spike protein (S). These may alter the antigenic structure of S, causing escape from natural or vaccine-induced immunity. Beta is particularly difficult to neutralize using serum induced by early pandemic SARS-CoV-2 strains and is most antigenically separated from Delta. To understand this, we generated 674 mAbs from Beta-infected individuals and performed a detailed structure-function analysis of the 27 most potent mAbs: one binding the spike N-terminal domain (NTD), the rest the receptor-binding domain (RBD). Two of these RBD-binding mAbs recognize a neutralizing epitope conserved between SARS-CoV-1 and -2, while 18 target mutated residues in Beta: K417N, E484K, and N501Y. There is a major response to N501Y, including a public IgVH4-39 sequence, with E484K and K417N also targeted. Recognition of these key residues underscores why serum from Beta cases poorly neutralizes early pandemic and Delta viruses. |
 リンク リンク |  Cell Host Microbe / Cell Host Microbe /  PubMed:34921776 / PubMed:34921776 /  PubMed Central PubMed Central |
| 手法 | EM (単粒子) / X線回折 |
| 解像度 | 1.7 - 4.9 Å |
| 構造データ | EMDB-13857, PDB-7q6e: EMDB-13868, PDB-7q9f: EMDB-13869, PDB-7q9g: EMDB-13870, PDB-7q9i: EMDB-13871, PDB-7q9j: EMDB-13872, PDB-7q9k: EMDB-13873, PDB-7q9m:  EMDB-13874: EMDB-13875, PDB-7q9p:  PDB-7pry:  PDB-7prz:  PDB-7ps0:  PDB-7ps1:  PDB-7ps2:  PDB-7ps3:  PDB-7ps4:  PDB-7ps5:  PDB-7ps6:  PDB-7ps7:  PDB-7q0g:  PDB-7q0h:  PDB-7q0i: |
| 化合物 |  ChemComp-SO4:  ChemComp-NAG:  ChemComp-GOL:  ChemComp-CL:  ChemComp-IOD:  ChemComp-PEG:  ChemComp-HOH: 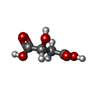 ChemComp-MLT: 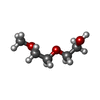 ChemComp-PG0:  ChemComp-K: 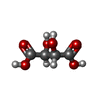 ChemComp-TAR:  ChemComp-PO4:  ChemComp-PG4: |
| 由来 |
|
 キーワード キーワード | VIRAL PROTEIN / SARS-CoV-2 alpha variant / beta variant / gamma variant / delta variant / B.1.1.7 / B.1.351 / P.1 / B.1.617.2 / antibody / receptor-binding-domain / spike / neutralisation / VIRAL PROTEIN/IMMUNE SYSTEM / SARS-COV-2 B.1.1.7 (Alpha) VARIANT / B.1.351 (Beta) VARIANT / P.1 (Gamma) VARIANT / B.1.617.2 (Delta) VARIANT / IMMUNE SYSTEM / VIRAL PROTEIN-IMMUNE SYSTEM COMPLEX / RBD / N-terminal domain / NTD / SARS-CoV2 / glycoprotein / fab / B.1.135 / Complex / neutralising / convalescent sera |
 ムービー
ムービー コントローラー
コントローラー 構造ビューア
構造ビューア 万見文献について
万見文献について




















 homo sapiens (ヒト)
homo sapiens (ヒト)