+Search query
-Structure paper
| Title | Structure and function of a near fully-activated intermediate GPCR-Gαβγ complex. |
|---|---|
| Journal, issue, pages | Nat Commun, Vol. 16, Issue 1, Page 1100, Year 2025 |
| Publish date | Jan 28, 2025 |
 Authors Authors | Maxine Bi / Xudong Wang / Jinan Wang / Jun Xu / Wenkai Sun / Victor Ayo Adediwura / Yinglong Miao / Yifan Cheng / Libin Ye /  |
| PubMed Abstract | Unraveling the signaling roles of intermediate complexes is pivotal for G protein-coupled receptor (GPCR) drug development. Despite hundreds of GPCR-Gαβγ structures, these snapshots primarily ...Unraveling the signaling roles of intermediate complexes is pivotal for G protein-coupled receptor (GPCR) drug development. Despite hundreds of GPCR-Gαβγ structures, these snapshots primarily capture the fully activated complex. Consequently, the functions of intermediate GPCR-G protein complexes remain elusive. Guided by a conformational landscape visualized via F quantitative NMR and molecular dynamics (MD) simulations, we determined the structure of an intermediate GPCR-mini-Gαβγ complex at 2.6 Å using cryo-EM, by blocking its transition to the fully activated complex. Furthermore, we present direct evidence that the complex at this intermediate state initiates a rate-limited nucleotide exchange before transitioning to the fully activated complex. In this state, BODIPY-GDP/GTP based nucleotide exchange assays further indicated the α-helical domain of the Gα is partially open, allowing it to grasp a nucleotide at a non-canonical binding site, distinct from the canonical nucleotide-binding site. These advances bridge a significant gap in our understanding of the complexity of GPCR signaling. |
 External links External links |  Nat Commun / Nat Commun /  PubMed:39875358 / PubMed:39875358 /  PubMed Central PubMed Central |
| Methods | EM (single particle) |
| Resolution | 2.63 - 3.36 Å |
| Structure data | EMDB-47951, PDB-9ee8: EMDB-47952, PDB-9ee9: EMDB-47953, PDB-9eea: |
| Chemicals | 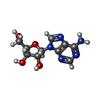 ChemComp-ADN: |
| Source |
|
 Keywords Keywords | SIGNALING PROTEIN/IMMUNE SYSTEM / GPCR / Heterotrimer / Receptor / cryo-EM / SIGNALING PROTEIN-IMMUNE SYSTEM complex |
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About Yorodumi Papers
About Yorodumi Papers









 homo sapiens (human)
homo sapiens (human)
