+Search query
-Structure paper
| Title | Mechanisms of OCT4-SOX2 motif readout on nucleosomes. |
|---|---|
| Journal, issue, pages | Science, Vol. 368, Issue 6498, Page 1460-1465, Year 2020 |
| Publish date | Jun 26, 2020 |
 Authors Authors | Alicia K Michael / Ralph S Grand / Luke Isbel / Simone Cavadini / Zuzanna Kozicka / Georg Kempf / Richard D Bunker / Andreas D Schenk / Alexandra Graff-Meyer / Ganesh R Pathare / Joscha Weiss / Syota Matsumoto / Lukas Burger / Dirk Schübeler / Nicolas H Thomä /  |
| PubMed Abstract | Transcription factors (TFs) regulate gene expression through chromatin where nucleosomes restrict DNA access. To study how TFs bind nucleosome-occupied motifs, we focused on the reprogramming factors ...Transcription factors (TFs) regulate gene expression through chromatin where nucleosomes restrict DNA access. To study how TFs bind nucleosome-occupied motifs, we focused on the reprogramming factors OCT4 and SOX2 in mouse embryonic stem cells. We determined TF engagement throughout a nucleosome at base-pair resolution in vitro, enabling structure determination by cryo-electron microscopy at two preferred positions. Depending on motif location, OCT4 and SOX2 differentially distort nucleosomal DNA. At one position, OCT4-SOX2 removes DNA from histone H2A and histone H3; however, at an inverted motif, the TFs only induce local DNA distortions. OCT4 uses one of its two DNA-binding domains to engage DNA in both structures, reading out a partial motif. These findings explain site-specific nucleosome engagement by the pluripotency factors OCT4 and SOX2, and they reveal how TFs distort nucleosomes to access chromatinized motifs. |
 External links External links |  Science / Science /  PubMed:32327602 PubMed:32327602 |
| Methods | EM (single particle) |
| Resolution | 3.05 - 3.49 Å |
| Structure data | EMDB-10406, PDB-6t90: EMDB-10408, PDB-6t93: EMDB-10864, PDB-6yov: |
| Chemicals | 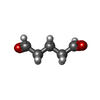 ChemComp-PTD: |
| Source |
|
 Keywords Keywords | TRANSCRIPTION / nucleosome / OCT4 / SOX2 / transcription factor |
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About Yorodumi Papers
About Yorodumi Papers









 homo sapiens (human)
homo sapiens (human)
