-検索条件
-検索結果
検索 (著者・登録者: zhu & k)の結果2,456件中、1から50件目までを表示しています

EMDB-37842: 
Cryo-EM structure of noradrenaline transporter in apo state
EMDB-37843: 
Cryo-EM structure of noradrenaline transporter in complex with noradrenaline
EMDB-37844: 
Cryo-EM structure of noradrenaline transporter in complex with a x-MrlA analogue
EMDB-37845: 
Cryo-EM structure of noradrenaline transporter in complex with bupropion
EMDB-37846: 
Cryo-EM structure of noradrenaline transporter in complex with ziprasidone

PDB-8wtu: 
Cryo-EM structure of noradrenaline transporter in apo state

PDB-8wtv: 
Cryo-EM structure of noradrenaline transporter in complex with noradrenaline

PDB-8wtw: 
Cryo-EM structure of noradrenaline transporter in complex with a x-MrlA analogue

PDB-8wtx: 
Cryo-EM structure of noradrenaline transporter in complex with bupropion

PDB-8wty: 
Cryo-EM structure of noradrenaline transporter in complex with ziprasidone
EMDB-45078: 
E.coli GroEL + PBZ1587 inhibitor
EMDB-45079: 
E.coli GroEL apoenzyme
EMDB-45080: 
E.Faecium GroEL

EMDB-45098: 
E.coli GroEL + PBZ1587 inhibitor C1 reconstruction

PDB-9c0b: 
E.coli GroEL + PBZ1587 inhibitor

PDB-9c0c: 
E.coli GroEL apoenzyme

PDB-9c0d: 
E.Faecium GroEL

EMDB-37515: 
Cryo-EM structure of inward-open state human norepinephrine transporter NET bound with antidepressant desipramine in KCl condition.

EMDB-37520: 
Cryo-EM structure of inward-open state human norepinephrine transporter NET bound with norepinephrine in nanodisc.

EMDB-39729: 
Cryo-EM structure of human norepinephrine transporter NET in the presence of the antidepressant atomoxetine in an outward-open state at resolution of 3.4 angstrom.

PDB-8wgr: 
Cryo-EM structure of inward-open state human norepinephrine transporter NET bound with antidepressant desipramine in KCl condition.

PDB-8wgx: 
Cryo-EM structure of inward-open state human norepinephrine transporter NET bound with norepinephrine in nanodisc.

PDB-8z1l: 
Cryo-EM structure of human norepinephrine transporter NET in the presence of the antidepressant atomoxetine in an outward-open state at resolution of 3.4 angstrom.
EMDB-38532: 
Cryo-EM structure of human ABCC4

PDB-8xok: 
Cryo-EM structure of human ABCC4
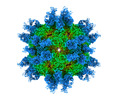

EMDB-32979: 
Cryo-EM structure of Coxsackievirus B1 A-particle in complex with nAb 8A10 (CVB1-A:8A10)

PDB-7x35: 
Cryo-EM structure of Coxsackievirus B1 A-particle in complex with nAb 8A10 (CVB1-A:8A10)

EMDB-39025: 
Structure of HCoV-HKU1A spike in the functionally anchored-3up conformation with 3TMPRSS2

EMDB-39026: 
Local structure of HCoV-HKU1A spike in complex with TMPRSS2 and glycan

EMDB-39036: 
Structure of HCoV-HKU1C spike in the functionally anchored-1up conformation with 1TMPRSS2

EMDB-39037: 
Structure of HCoV-HKU1C spike in the functionally anchored-2up conformation with 2TMPRSS2

EMDB-39038: 
Structure of HCoV-HKU1C spike in the functionally anchored-3up conformation with 2TMPRSS2

EMDB-39039: 
Structure of HCoV-HKU1C spike in the functionally anchored-3up conformation with 3TMPRSS2

EMDB-39040: 
Local structure of HCoV-HKU1C spike in complex with TMPRSS2 and glycan

EMDB-39041: 
Structure of HCoV-HKU1C spike in the inactive-closed conformation

EMDB-39042: 
Structure of HCoV-HKU1C spike in the inactive-1up conformation

EMDB-39043: 
Structure of HCoV-HKU1C spike in the inactive-2up conformation

EMDB-39044: 
Structure of HCoV-HKU1C spike in the glycan-activated-closed conformation

EMDB-39045: 
Structure of HCoV-HKU1C spike in the glycan-activated-1up conformation

EMDB-39046: 
Structure of HCoV-HKU1C spike in the glycan-activated-2up conformation

EMDB-39047: 
Structure of HCoV-HKU1C spike in the glycan-activated-3up conformation

EMDB-39048: 
Local structure of HCoV-HKU1C spike in complex with glycan

PDB-8y7x: 
Structure of HCoV-HKU1A spike in the functionally anchored-3up conformation with 3TMPRSS2

PDB-8y7y: 
Local structure of HCoV-HKU1A spike in complex with TMPRSS2 and glycan

PDB-8y87: 
Structure of HCoV-HKU1C spike in the functionally anchored-1up conformation with 1TMPRSS2

PDB-8y88: 
Structure of HCoV-HKU1C spike in the functionally anchored-2up conformation with 2TMPRSS2

PDB-8y89: 
Structure of HCoV-HKU1C spike in the functionally anchored-3up conformation with 2TMPRSS2

PDB-8y8a: 
Structure of HCoV-HKU1C spike in the functionally anchored-3up conformation with 3TMPRSS2

PDB-8y8b: 
Local structure of HCoV-HKU1C spike in complex with TMPRSS2 and glycan

PDB-8y8c: 
Structure of HCoV-HKU1C spike in the inactive-closed conformation
ページ:
 ムービー
ムービー コントローラー
コントローラー 構造ビューア
構造ビューア EMN検索について
EMN検索について



 wwPDBはEMDBデータモデルのバージョン3へ移行します
wwPDBはEMDBデータモデルのバージョン3へ移行します
