-Search query
-Search result
Showing 1 - 50 of 324 items for (author: yoo & y)

EMDB entry, No image
EMDB-37139: 
Structure of SARS-CoV Spike protein complexed with antibody PW5-5

EMDB entry, No image
EMDB-37143: 
The local refined map of SARS-CoV-2 XBB Variant Spike protein complexed with antibody PW5-535

EMDB entry, No image
EMDB-37144: 
Trimer state of SARS-CoV Spike protein complexed with antibody PW5-535

EMDB entry, No image
EMDB-37145: 
The local refined map of SARS-CoV Spike protein complexed with antibody PW5-5

EMDB entry, No image
EMDB-37160: 
Monomer state of SARS-CoV-2 XBB Variant Spike protein trimer complexed with antibody PW5-5

EMDB entry, No image
EMDB-37161: 
Monomer state of SARS-CoV Spike protein complexed with antibody PW5-535

EMDB entry, No image
EMDB-37162: 
Structure of SARS-CoV-2 Omicron BA.1 Spike complexed with antibody PW5-570

EMDB entry, No image
EMDB-37163: 
The local refined map of SARS-CoV-2 Omicron BA.1 Spike complexed with antibody PW5-570

EMDB entry, No image
EMDB-37164: 
State 1 of SARS-CoV-2 XBB Variant Spike protein trimer complexed with antibody PW5-5

EMDB entry, No image
EMDB-37165: 
Structure of SARS-CoV-2 XBB Variant Spike protein complexed with broadly neutralizing antibody PW5-535

PDB-8kdm: 
Structure of SARS-CoV Spike protein complexed with antibody PW5-5

PDB-8kdr: 
The local refined map of SARS-CoV-2 XBB Variant Spike protein complexed with antibody PW5-535

PDB-8kds: 
Trimer state of SARS-CoV Spike protein complexed with antibody PW5-535

PDB-8kdt: 
The local refined map of SARS-CoV Spike protein complexed with antibody PW5-5

PDB-8kej: 
Monomer state of SARS-CoV-2 XBB Variant Spike protein trimer complexed with antibody PW5-5

PDB-8kek: 
Monomer state of SARS-CoV Spike protein complexed with antibody PW5-535

PDB-8keo: 
Structure of SARS-CoV-2 Omicron BA.1 Spike complexed with antibody PW5-570

PDB-8kep: 
The local refined map of SARS-CoV-2 Omicron BA.1 Spike complexed with antibody PW5-570

PDB-8keq: 
State 1 of SARS-CoV-2 XBB Variant Spike protein trimer complexed with antibody PW5-5

PDB-8ker: 
Structure of SARS-CoV-2 XBB Variant Spike protein complexed with broadly neutralizing antibody PW5-535
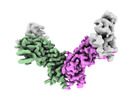
EMDB-44389: 
Cryo-EM structure of the ZBTB5 BTB domain filament
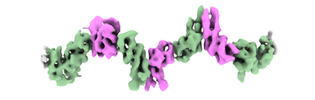
EMDB-44391: 
Cryo-EM structure of the ZBTB9 BTB domain filament

PDB-9b9r: 
Cryo-EM structure of the ZBTB5 BTB domain filament

PDB-9b9v: 
Cryo-EM structure of the ZBTB9 BTB domain filament

EMDB-42837: 
Cryo-EM structure of GeoCas9 in complex with sgRNA and target DNA

EMDB-42838: 
Cryo-EM structure of iGeoCas9 in complex with sgRNA and target DNA

PDB-8uza: 
Cryo-EM structure of GeoCas9 in complex with sgRNA and target DNA

PDB-8uzb: 
Cryo-EM structure of iGeoCas9 in complex with sgRNA and target DNA

EMDB-42676: 
5-HT2AR bound to Lisuride in complex with a mini-Gq protein and an active-state stabilizing single-chain variable fragment (scFv16) obtained by cryo-electron microscopy (cryoEM)

EMDB-42999: 
5HT2AR-miniGq heterotrimer in complex with a novel agonist obtained from large scale docking

PDB-8uwl: 
5-HT2AR bound to Lisuride in complex with a mini-Gq protein and an active-state stabilizing single-chain variable fragment (scFv16) obtained by cryo-electron microscopy (cryoEM)

PDB-8v6u: 
5HT2AR-miniGq heterotrimer in complex with a novel agonist obtained from large scale docking
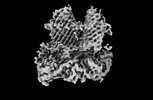
EMDB-40682: 
The cryo-EM structure of the EcBAM/EspP(beta1-12) complex
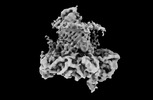
EMDB-40700: 
The cryo-EM structure of the EcBAM/EspP(beta8-12) complex
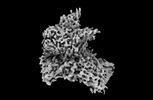
EMDB-40701: 
The cryo-EM structure of the EcBAM/EspP(beta7-12) complex

EMDB-38453: 
Structure of the SARS-CoV-2 EG.5.1 spike glycoprotein in complex with ACE2 (1-up state)

EMDB-38454: 
Structure of the SARS-CoV-2 EG.5.1 spike RBD in complex with ACE2

PDB-8xlm: 
Structure of the SARS-CoV-2 EG.5.1 spike glycoprotein in complex with ACE2 (1-up state)

PDB-8xln: 
Structure of the SARS-CoV-2 EG.5.1 spike RBD in complex with ACE2

EMDB-37648: 
SARS-CoV-2 EG.5.1 spike glycoprotein (1-up state)

EMDB-37650: 
SARS-CoV-2 EG.5.1 spike glycoprotein (closed-2 state)

EMDB-37651: 
SARS-CoV-2 EG.5.1 spike glycoprotein (closed-1 state)

PDB-8wmd: 
Structure of the SARS-CoV-2 EG.5.1 spike glycoprotein (closed-2 state)

PDB-8wmf: 
Structure of the SARS-CoV-2 EG.5.1 spike glycoprotein (closed-1 state)
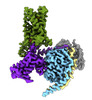
EMDB-43647: 
CryoEM structure of Gi-coupled TAS2R14 with cholesterol and an intracellular tastant
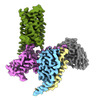
EMDB-43650: 
CryoEM structure of Ggust-coupled TAS2R14 with cholesterol and an intracellular tastant

EMDB-43656: 
CryoEM structure of Gi-coupled TAS2R14 with cholesterol and an intracellular tastant (Locally refined map)

EMDB-43657: 
CryoEM structure of Ggust-coupled TAS2R14 with cholesterol and an intracellular tastant (Locally refined map)

PDB-8vy7: 
CryoEM structure of Gi-coupled TAS2R14 with cholesterol and an intracellular tastant

PDB-8vy9: 
CryoEM structure of Ggust-coupled TAS2R14 with cholesterol and an intracellular tastant
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
