-Search query
-Search result
Showing all 20 items for (author: tran & pt)

EMDB-46908: 
Recombinant AD-fold Paired Helical Filament Polymorph
Method: helical / : Vaquer-Alicea J, Diamond MI, Kunach P

EMDB-46915: 
Recombinant AD-fold Quadruple Helical Filament Polymorph
Method: helical / : Vaquer-Alicea J, Diamond MI, Kunach P

EMDB-46870: 
Recombinant AD PHF
Method: helical / : Vaquer-Alicea J, Diamond MI, Kunach P

EMDB-46885: 
AD-fold Tri Helical Filament Polymorph
Method: helical / : Vaquer-Alicea J, Kunach P, Diamond MI

EMDB-46909: 
Recombinant AD-fold Paired Helical Filament II Polymorph
Method: helical / : Vaquer-Alicea J, Diamond MI, Kunach P

EMDB-46911: 
Recombinant AD-fold Triple Helical Filament Polymorph
Method: helical / : Vaquer-Alicea J, Diamond MI, Kunach P

EMDB-47092: 
Recombinant Truncated Tau 266-391 fibrillized in NaCl Second Polymorph
Method: helical / : Vaquer-Alicea J, Diamond MI, Kunach P

EMDB-46937: 
Recombinant Truncated Tau 266-391 fibrillized in NaCl
Method: helical / : Vaquer-Alicea J, Diamond MI, Kunach P

EMDB-15488: 
Cryo-EM structure of the proline-rich antimicrobial peptide drosocin bound to the terminating ribosome
Method: single particle / : Koller TO, Morici M, Wilson DN

EMDB-15523: 
Cryo-EM structure of the proline-rich antimicrobial peptide drosocin bound to the elongating ribosome
Method: single particle / : Koller TO, Morici M, Wilson DN

EMDB-15533: 
Cryo-EM structure of the proline-rich antimicrobial peptide drosocin bound to the 50S ribosomal subunit
Method: single particle / : Koller TO, Morici M, Wilson DN

PDB-8akn: 
Cryo-EM structure of the proline-rich antimicrobial peptide drosocin bound to the terminating ribosome
Method: single particle / : Koller TO, Morici M, Wilson DN

PDB-8am9: 
Cryo-EM structure of the proline-rich antimicrobial peptide drosocin bound to the elongating ribosome
Method: single particle / : Koller TO, Morici M, Wilson DN

PDB-8ana: 
Cryo-EM structure of the proline-rich antimicrobial peptide drosocin bound to the 50S ribosomal subunit
Method: single particle / : Koller TO, Morici M, Wilson DN

EMDB-25448: 
Negative-stain EM reconstruction of SpFN_1B-06-PL, a SARS-CoV-2 spike fused to H.pylori ferritin nanoparticle vaccine candidate
Method: single particle / : Thomas PV, Smith C, Chen WH, Sankhala RS, Hajduczki A, Choe M, Martinez E, Chang W, Peterson CE, Karch C, Gohain N, Kannadka CB, de Val N, Joyce MG, Modjarrad K
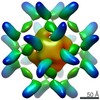
EMDB-25449: 
RFN_131, a Ferritin-based Nanoparticle Vaccine Candidate Displaying the SARS-CoV-2 Receptor-Binding Domain
Method: single particle / : Thomas PV, Smith C, Chen WH, Sankhala RS, Hajduczki A, Choe M, Martinez E, Chang W, Peterson CE, Karch C, Gohain N, Kannadka CB, de Val N, Joyce MG, Modjarrad K

EMDB-25450: 
pCoV146, a Ferritin-based Nanoparticle Vaccine Candidate Displaying the SARS-CoV-2 Spike Receptor-Binding and N-Terminal Domains
Method: single particle / : Thomas PV, Smith C, Chen WH, Sankhala RS, Hajduczki A, Choe M, Martinez E, Chang W, Peterson CE, Karch C, Gohain N, Kannadka CB, de Val N, Joyce MG, Modjarrad K

EMDB-25451: 
pCoV111, a Ferritin-based Nanoparticle Vaccine Candidate Displaying the SARS-CoV-2 Spike S1 Subunit
Method: single particle / : Thomas PV, Smith C, Chen WH, Sankhala RS, Hajduczki A, Choe M, Martinez E, Chang W, Peterson CE, Karch C, Gohain N, Kannadka CB, de Val N, Joyce MG, Modjarrad K

EMDB-8673: 
Cryo-EM structure of yeast cytoplasmic dynein-1 with Lis1 and ATP
Method: single particle / : Cianfrocco MA, DeSantis ME

EMDB-8706: 
Cryo-EM structure of yeast cytoplasmic dynein with Walker B mutation at AAA3 in presence of ATP-VO4
Method: single particle / : Cianfrocco MA, DeSantis ME
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
