-Search query
-Search result
Showing all 31 items for (author: sakamoto & j)

EMDB-42676: 
5-HT2AR bound to Lisuride in complex with a mini-Gq protein and an active-state stabilizing single-chain variable fragment (scFv16) obtained by cryo-electron microscopy (cryoEM)

EMDB-42999: 
5HT2AR-miniGq heterotrimer in complex with a novel agonist obtained from large scale docking

PDB-8uwl: 
5-HT2AR bound to Lisuride in complex with a mini-Gq protein and an active-state stabilizing single-chain variable fragment (scFv16) obtained by cryo-electron microscopy (cryoEM)

PDB-8v6u: 
5HT2AR-miniGq heterotrimer in complex with a novel agonist obtained from large scale docking

EMDB-15851: 
Cryo-EM structure of cytochrome bd oxidase from C. glutamicum

PDB-8b4o: 
Cryo-EM structure of cytochrome bd oxidase from C. glutamicum

EMDB-14587: 
Cryo-EM structure of the human GS-GN complex in the inhibited state

PDB-7zbn: 
Cryo-EM structure of the human GS-GN complex in the inhibited state

EMDB-12324: 
In situ cryo-electron tomogram of the Synechocystis wild-type VIPP1 strain grown in high light

EMDB-12325: 
In situ cryo-electron tomogram of the Synechocystis F4E VIPP1 mutant grown in high light

EMDB-12326: 
In situ cryo-electron tomogram of the Synechocystis V11E VIPP1 mutant grown in high light

EMDB-12327: 
In situ cryo-electron tomogram of a cluster of VIPP1-mCherry structures inside the Chlamydomonas chloroplast

EMDB-12328: 
In situ cryo-electron tomogram of a cluster of VIPP1-mCherry structures inside the Chlamydomonas chloroplast

EMDB-12329: 
In situ cryo-electron tomogram of a cluster of VIPP1-mCherry structures inside the Chlamydomonas chloroplast

EMDB-12330: 
In situ cryo-electron tomogram of a cluster of VIPP1-mCherry structures inside the Chlamydomonas chloroplast

EMDB-12710: 
Structural basis for VIPP1 oligomerization and maintenance of thylakoid membrane integrity

EMDB-12711: 
Structural basis for VIPP1 oligomerization and maintenance of thylakoid membrane integrity

EMDB-12712: 
Structural basis for VIPP1 oligomerization and maintenance of thylakoid membrane integrity

EMDB-12713: 
Structural basis for VIPP1 oligomerization and maintenance of thylakoid membrane integrity

EMDB-12714: 
Structural basis for VIPP1 oligomerization and maintenance of thylakoid membrane integrity

PDB-7o3w: 
Structural basis for VIPP1 oligomerization and maintenance of thylakoid membrane integrity

PDB-7o3x: 
Structural basis for VIPP1 oligomerization and maintenance of thylakoid membrane integrity

PDB-7o3y: 
Structural basis for VIPP1 oligomerization and maintenance of thylakoid membrane integrity

PDB-7o3z: 
Structural basis for VIPP1 oligomerization and maintenance of thylakoid membrane integrity

PDB-7o40: 
Structural basis for VIPP1 oligomerization and maintenance of thylakoid membrane integrity
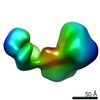
EMDB-11054: 
Linear Ubiquitin Chain Assembly Complex

EMDB-4908: 
Cryo-EM structure of the E. coli cytochrome bd-I oxidase at 2.68 A resolution

PDB-6rko: 
Cryo-EM structure of the E. coli cytochrome bd-I oxidase at 2.68 A resolution

EMDB-6122: 
RNA-targeting by the Type III-A CRISPR-Cas Csm complex of Thermus thermophilus

EMDB-5719: 
Electron microscopy of the negatively-stained Cmr complex from Thermus thermophilus HB8.

EMDB-2418: 
Structure and activity of an RNA-targeting Type III-B CRISPER-Cas complex
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
