-Search query
-Search result
Showing 1 - 50 of 66 items for (author: palese & p)

EMDB-46833: 
Negative stain map of cH5/1 apo
Method: single particle / : Han J, Rodriguez AJ, Ward AB

EMDB-46834: 
Negative stain map of cH11/1 closed apo
Method: single particle / : Han J, Rodriguez AJ, Ward AB

EMDB-46835: 
Negative stain of cH5/1 HA in complex with 31.b.09 Fab
Method: single particle / : Han J, Rodriguez AJ, Ward AB

EMDB-46836: 
Negative stain of cH5/1 HA in complex with CR9114 Fab
Method: single particle / : Han J, Rodriguez AJ, Ward AB

EMDB-46837: 
Negative stain of cH11/1 HA in complex with 31.b.09 Fab
Method: single particle / : Han J, Rodriguez AJ, Ward AB

EMDB-46838: 
Negative stain of cH8/1 HA in complex with 31.b.09 Fab
Method: single particle / : Han J, Rodriguez AJ, Ward AB

EMDB-46839: 
Negative stain map of cH8/1 HA in complex with CR9114 Fab
Method: single particle / : Han J, Rodriguez AJ, Ward AB

EMDB-46840: 
Negative stain of cH5/1 HA in complex with FluA20 fab
Method: single particle / : Han J, Rodriguez AJ, Ward AB

EMDB-46841: 
Negative stain of cH8/1 HA in complex with FluA20 Fab
Method: single particle / : Han J, Rodriguez AJ, Ward AB

EMDB-46842: 
Negative stain maps of cH8/1 apo HA
Method: single particle / : Han J, Rodriguez AJ, Ward AB

EMDB-26928: 
Negative stain map of cH4/3
Method: single particle / : Han J, Ward AB

EMDB-26929: 
Negative stain half map of cH4/3 state 1
Method: single particle / : Han J, Ward AB

EMDB-26930: 
Negative stain map of cH4/3 state 3
Method: single particle / : Han J, Ward AB

EMDB-26931: 
Negative stain map of cH4/3 state 3
Method: single particle / : Han J, Ward AB

EMDB-26932: 
Negative stain map of cH4/3 state 4
Method: single particle / : Han J, Ward AB

EMDB-26933: 
Negative stain map of cH15/3
Method: single particle / : Han J, Ward AB

EMDB-26934: 
Negative stain map of cH15/3 state 1
Method: single particle / : Han J, Ward AB

EMDB-26935: 
Negative stain map of cH15/3 state 2
Method: single particle / : Han J, Ward AB

EMDB-26937: 
Negative stain map of cH15/3 state 3
Method: single particle / : Han J, Ward AB

EMDB-26938: 
Negative stain map of cH15/3 state 4
Method: single particle / : Han J, Ward AB

EMDB-26939: 
Negative stain map of cH15/3 in complex with CR9114 Fab
Method: single particle / : Han J, Ward AB

EMDB-25634: 
Negative stain map of monoclonal Fab 047-09 4F04 binding the anchor epitope of H1 HA
Method: single particle / : Han J, Richey ST, Ward AB

EMDB-25635: 
Negative stain map of monoclonal Fab 241 IgA 2F04 binding the anchor epitope of H1 HA
Method: single particle / : Han J, Richey ST, Ward AB

EMDB-25636: 
Negative stain map of polyclonal Fab 236.7 binding the anchor and esterase epitopes of H1 HA
Method: single particle / : Han J, Richey ST, Ward AB

EMDB-25637: 
Negative stain map of polyclonal Fab 236.7 binding the RBS epitope of H1 HA
Method: single particle / : Han J, Richey ST, Ward AB

EMDB-25638: 
Negative stain map of polyclonal Fab 236.14 binding an epitope on the top of the head of H1 HA
Method: single particle / : Han J, Richey ST, Ward AB
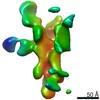
EMDB-25639: 
Negative stain map of polyclonal Fab 236.14 binding the esterase epitope of H1 HA
Method: single particle / : Han J, Richey ST, Ward AB

EMDB-25640: 
Negative stain map of polycolonal Fab 236.14 binding the RBS epitope of H1 HA
Method: single particle / : Han J, Richey ST, Ward AB

EMDB-25641: 
Negative stain map of polyclonal Fab 236.14 binding the anchor epitope of H1 HA
Method: single particle / : Han J, Richey ST, Ward AB
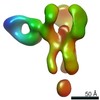
EMDB-25642: 
Negative stain map of polyclonal Fab 241.7 binding the esterase epitope of H1 HA
Method: single particle / : Han J, Richey ST, Ward AB
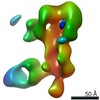
EMDB-25643: 
Negative stain map of polyclonal Fab 241.14 binding the anchor epitope of H1 HA
Method: single particle / : Han J, Richey ST, Ward AB
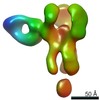
EMDB-25644: 
Negative stain map of polyclonal Fab 241.14 binding the esterase epitope of H1 HA
Method: single particle / : Han J, Richey ST, Ward AB

EMDB-25645: 
Negative stain map of polyclonal Fab 241.14 binding an epitope on the top of the head of H1 HA
Method: single particle / : Han J, Richey ST, Ward AB

EMDB-25646: 
Negative stain map of polyclonal Fab 241.14 binding the RBS epitope of H1 HA
Method: single particle / : Han J, Richey ST, Ward AB

EMDB-25655: 
CryoEM map of anchor 222-1C06 Fab and lateral patch 2B05 Fab binding H1 HA
Method: single particle / : Han J, Ward AB

PDB-7t3d: 
CryoEM map of anchor 222-1C06 Fab and lateral patch 2B05 Fab binding H1 HA
Method: single particle / : Han J, Ward AB

EMDB-23792: 
CryoEM structure of monoclonal Fab 045-09 2B05 binding the lateral patch of influenza virus H1 HA
Method: single particle / : Han J, Ward A
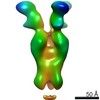
EMDB-23793: 
Negative stain map of monoclonal Fab SFV009 2G01 binding the RBS of H1 HA
Method: single particle / : Han J, Ward AB

EMDB-23794: 
Negative stain map of monoclonal Fab 045-09 2B05 binding the lateral patch of H1 HA
Method: single particle / : Han J, Ward AB

EMDB-23795: 
Negative stain map of monoclonal Fab SFV019 2A06 binding the lateral patch of H1 HA
Method: single particle / : Han J, Ward AB
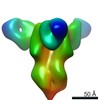
EMDB-23796: 
Negative stain map of monoclonal Fab SFV015 2F02 binding the lateral patch of H1 HA
Method: single particle / : Han J, Ward AB

EMDB-23797: 
Negative stain map of monoclonal Fab 047-09 4G02 binding the lateral patch of H1 HA
Method: single particle / : Han J, Ward AB
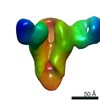
EMDB-23798: 
Negative stain map of monoclonal Fab 047-09 4B06 binding the lateral patch of H1 HA
Method: single particle / : Han J, Ward AB

PDB-7mem: 
CryoEM structure of monoclonal Fab 045-09 2B05 binding the lateral patch of influenza virus H1 HA
Method: single particle / : Han J, Ward A
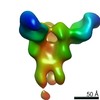
EMDB-23313: 
Negative stain map of monoclonal Fab 23 binding the lateral patch of H1 HA
Method: single particle / : Han J, Freyn AW, Ward AB

EMDB-23314: 
Negative stain map of monoclonal Fab 45 binding the esterase domain of H1 HA
Method: single particle / : Han J, Freyn AW, Ward AB
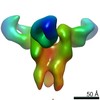
EMDB-23315: 
Negative stain map of monoclonal Fab 50 binding the lateral patch of H1 HA
Method: single particle / : Han J, Freyn AW, Ward AB
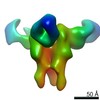
EMDB-23316: 
Negative stain map of monoclonal Fab 56 binding the lateral patch of H1 HA
Method: single particle / : Han J, Freyn AW, Ward AB

EMDB-23317: 
Negative stain map of monoclonal Fab 68 binding the Sa antigenic site of H1 HA
Method: single particle / : Han J, Freyn AW, Ward AB
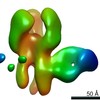
EMDB-23318: 
Negative stain map of monoclonal Fab 76 binding the stem of H1 HA
Method: single particle / : Han J, Freyn AW, Ward AB
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
