-検索条件
-検索結果
検索 (著者・登録者: mueller & dm)の結果79件中、1から50件目までを表示しています

EMDB-18953: 
Cryo-electron tomogram of Yersinia entomophaga chi2-sfGFP cells

EMDB-18954: 
Cryo-electron tomogram of Yersinia entomophaga chi2-sfGFP cells
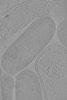
EMDB-18955: 
Cryo-electron tomogram of Yersinia entomophaga chi2-sfGFP cells

EMDB-18957: 
Cryo-electron tomogram of Yersinia entomophaga MH96 cells

EMDB-18958: 
Cryo-electron tomogram of Yersinia entomophaga MH96 cells

EMDB-18960: 
Cryo-electron tomogram of Yersinia entomophaga delta LC cells

EMDB-18961: 
Cryo-electron tomogram of mechanically cryo-milled Yersinia entomophaga delta LC cells

EMDB-18962: 
Cryo-electron tomogram of a lysate preparation of Yersinia entomophaga delta LC cells
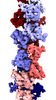
EMDB-18970: 
Subtomogram average of M66 filaments in Yersinia entomophaga cells
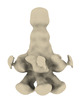
EMDB-18971: 
Subtomogram average of YenTc-Chi2-sfGFP from Yersinia entomophaga chi2-sfGFP
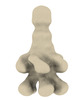
EMDB-18972: 
Subtomogram average of YenTc from Yersinia entomophaga MH96

EMDB-19370: 
Cryo-electron tomogram of Yersinia entomophaga delta YenTc cells
EMDB-19371: 
Cryo-electron tomogram of Yersinia entomophaga delta YenTc cells

EMDB-19372: 
Cryo-electron tomogram of Yersinia entomophaga delta YenTc cells

EMDB-19373: 
Cryo-electron tomogram of Yersinia entomophaga delta YenTc cells

EMDB-19374: 
Cryo-electron tomogram of Yersinia entomophaga chi2-sfGFP cells

EMDB-19375: 
Cryo-electron tomogram of Yersinia entomophaga chi2-sfGFP cells

EMDB-19376: 
Cryo-electron tomogram of Yersinia entomophaga delta LC delta YenTc cells
EMDB-19377: 
Cryo-electron tomogram of Yersinia entomophaga MH96 cells grown at 37 degrees
EMDB-19378: 
Cryo-electron tomogram of Yersinia entomophaga delta LC cells grown at 37 degrees

EMDB-19379: 
Cryo-electron tomogram of Yersinia entomophaga delta LC delta M66 cells

EMDB-19380: 
Cryo-electron tomogram of Yersinia entomophaga delta LC delta M66 cells

EMDB-19381: 
Cryo-electron tomogram of a lysate preparation of Yersinia entomophaga delta LC delta M66 cells

EMDB-28809: 
Yeast ATP synthase map in conformation-1 at pH 6

EMDB-28835: 
Yeast ATP synthase map in conformation-2, at pH 6

EMDB-29250: 
Yeast ATP Synthase in conformation-3, at pH 6

EMDB-28685: 
Yeast ATP synthase consensus map in conformation-0 at pH 6

EMDB-29270: 
Yeast ATP Synthase map in presence of MgATP

EMDB-29278: 
F1-focused map of yeast ATP Synthase in presence of MgATP

EMDB-29251: 
Fo and central rotor focused map of yeast ATP Synthase in conformation-3, at pH 6

EMDB-28811: 
F1-focused map of yeast ATP synthase in conformation-1, at pH 6

EMDB-28814: 
Fo and central rotor focused map of yeast ATP synthase in conformation-1, at pH 6

EMDB-28836: 
F1-focused map of yeast ATP synthase in conformation-2, at pH 6

EMDB-28837: 
Fo and central rotor focused map of yeast ATP synthase in conformation-2, at pH 6

EMDB-28813: 
Fo-focused map of yeast ATP synthase in conformation-1, at pH 6

EMDB-28687: 
Yeast ATP synthase F1-focused map in conformation-0 at pH 6

EMDB-28689: 
Yeast ATP synthase Fo-focused map in conformation-0 at pH 6

EMDB-16284: 
Cryotomogram of L-form-like cytoplasmic membrane vesicles of Listeria monocytogenes Rev2 cell

EMDB-16285: 
Cryotomogram of endolysin Ply006 treated Listeria monocytogenes Rev2 cell (stage 1)

EMDB-16286: 
Cryotomogram of endolysin Ply006 treated Listeria monocytogenes Rev2 cell (stage 2)

EMDB-16287: 
Cryotomogram of a A006 virion attaching to a Listeria monocytogenes Rev2 cell

EMDB-16288: 
Cryotomogram of endolysin Ply006 treated Listeria monocytogenes Rev2 cell (stage 3)

EMDB-16289: 
Cryotomogram of endolysin Ply007 treated Enterococcus faecalis cells

EMDB-16290: 
Cryotomogram of endolysin Ply007 treated Enterococcus faecalis cells

EMDB-16291: 
Cryotomogram of individual Efs7 virions and phage attachment to a Enterococcus faecalis cell

EMDB-16305: 
Cryotomogram of endolysin Ply007 treated Enterococcus faecalis cells

EMDB-16306: 
Cryotomogram of Listeria monocytogenes Rev2 cells

EMDB-11745: 
Cryo-EM structure of an extracellular contractile injection system in marine bacterium Algoriphagus machipongonensis, the baseplate complex in extended state applied 3-fold symmetry.

PDB-7aef: 
Cryo-EM structure of an extracellular contractile injection system in marine bacterium Algoriphagus machipongonensis, the baseplate complex in extended state applied 3-fold symmetry.

EMDB-11734: 
Cryo-EM structure of an extracellular contractile injection system in marine bacterium Algoriphagus machipongonensis, the cap portion in extended state.
ページ:
 ムービー
ムービー コントローラー
コントローラー 構造ビューア
構造ビューア EMN検索について
EMN検索について



 wwPDBはEMDBデータモデルのバージョン3へ移行します
wwPDBはEMDBデータモデルのバージョン3へ移行します
