-検索条件
-検索結果
検索 (著者・登録者: meyerson & jr)の結果全39件を表示しています

EMDB-26719: 
FMC63 scFv in complex with soluble CD19

EMDB-26720: 
SJ25C1 Fab in complex with soluble CD19

EMDB-25414: 
Structure of human Kv1.3 with Fab-ShK fusion

EMDB-25416: 
Structure of human Kv1.3

EMDB-25417: 
Structure of human Kv1.3 with A0194009G09 nanobodies

EMDB-24444: 
Mouse GITR (mGITR) with DTA-1 Fab fragment

EMDB-24789: 
Isoproterenol bound beta1 adrenergic receptor in complex with heterotrimeric Gi protein

EMDB-24790: 
Isoproterenol bound beta1 adrenergic receptor in complex with heterotrimeric Gi/s chimera protein

EMDB-23542: 
Structure of full-length GluK1 with L-Glu

EMDB-23494: 
Cryo-EM of the SLFN12-PDE3A complex: PDE3A body refinement

EMDB-23495: 
Cryo-EM of the SLFN12-PDE3A complex: Consensus subset model

EMDB-23496: 
Cryo-EM of the SLFN12-PDE3A complex: SLFN12 body refinement

EMDB-23014: 
GluK2/K5 with 6-Cyano-7-nitroquinoxaline-2,3-dione (CNQX)

EMDB-23015: 
GluK2/K5 with L-Glu

EMDB-22357: 
Structural Basis of the Activation of Heterotrimeric Gs-protein by Isoproterenol-bound Beta1-Adrenergic Receptor

EMDB-8289: 
GluK2EM with 2S,4R-4-methylglutamate

EMDB-8290: 
GluK2EM with LY466195

EMDB-2680: 
Density map of GluA2em in complex with ZK200775

EMDB-2684: 
Density map of GluA2em in complex with LY451646 and glutamate

EMDB-2685: 
Density map of GluK2 desensitized state in complex with 2S,4R-4-methylglutamate

EMDB-2686: 
Density map of GluA2em desensitized state in complex with quisqualate (class 1)

EMDB-2687: 
Density map of GluA2em desensitized state in complex with quisqualate (class 2)

EMDB-2688: 
Density map of GluA2em desensitized state in complex with quisqualate (class 3)

EMDB-2689: 
Density map of GluA2em in complex with quisqualate and LY451646

EMDB-5682: 
Molecular structure of the native influenza hemagglutinin H1 on virions

EMDB-5683: 
Molecular structure of the native influenza hemagglutinin H1 on virions

EMDB-5684: 
Molecular structure of the native influenza hemagglutinin H1 on virions complexed with broadly neutralizing stem antibody C179

EMDB-5685: 
Molecular structure of the native influenza hemagglutinin H1 on virions complexed with broadly neutralizing stem antibody C179

EMDB-5686: 
Molecular structure of the native influenza hemagglutinin H3 on virions

EMDB-5687: 
Molecular structure of the native influenza hemagglutinin H3 on virions
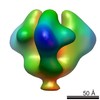
EMDB-5544: 
Molecular structure of the native HIV-1 Env trimer bound to VHH A12: Spike region

EMDB-5551: 
Molecular structure of the native HIV-1 Env trimer bound to VHH A12: Membrane region
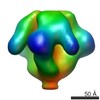
EMDB-5552: 
Molecular structure of the native HIV-1 Env trimer bound to m36: Spike region

EMDB-5553: 
Molecular structure of the native HIV-1 Env trimer bound to m36: Membrane region

EMDB-5554: 
Molecular structure of the native HIV-1 Env trimer bound to m36 and sCD4: Spike region

EMDB-5555: 
Molecular structure of the native HIV-1 Env trimer bound to m36 and sCD4: Membrane region

EMDB-5272: 
Molecular Structure of Unliganded Native SIVmneE11S gp120 trimer: Spike region
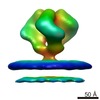
EMDB-5273: 
Molecular Structure of Unliganded Native SIVmac239 gp120 trimer: Spike region

EMDB-5274: 
Molecular Structure of Unliganded Native CP-MAC gp120 trimer: Spike region
 ムービー
ムービー コントローラー
コントローラー 構造ビューア
構造ビューア EMN検索について
EMN検索について



 wwPDBはEMDBデータモデルのバージョン3へ移行します
wwPDBはEMDBデータモデルのバージョン3へ移行します
