-Search query
-Search result
Showing all 39 items for (author: liedl & a)

EMDB-28135: 
Cryo-EM structure of L9 Fab in complex with rsCSP

PDB-8eh5: 
Cryo-EM structure of L9 Fab in complex with rsCSP

EMDB-13883: 
Broad-spectrum virus-trapping with heparan sulfate-modified DNA origami shells: T1 shell trapping a chikungunya VLP

EMDB-13884: 
Broad-spectrum virus-trapping with heparan sulfate-modified DNA origami shells: Octahedron shell trapping a HPV

EMDB-25634: 
Negative stain map of monoclonal Fab 047-09 4F04 binding the anchor epitope of H1 HA

EMDB-25635: 
Negative stain map of monoclonal Fab 241 IgA 2F04 binding the anchor epitope of H1 HA

EMDB-25636: 
Negative stain map of polyclonal Fab 236.7 binding the anchor and esterase epitopes of H1 HA

EMDB-25637: 
Negative stain map of polyclonal Fab 236.7 binding the RBS epitope of H1 HA

EMDB-25638: 
Negative stain map of polyclonal Fab 236.14 binding an epitope on the top of the head of H1 HA
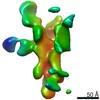
EMDB-25639: 
Negative stain map of polyclonal Fab 236.14 binding the esterase epitope of H1 HA

EMDB-25640: 
Negative stain map of polycolonal Fab 236.14 binding the RBS epitope of H1 HA

EMDB-25641: 
Negative stain map of polyclonal Fab 236.14 binding the anchor epitope of H1 HA
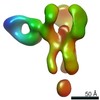
EMDB-25642: 
Negative stain map of polyclonal Fab 241.7 binding the esterase epitope of H1 HA
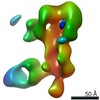
EMDB-25643: 
Negative stain map of polyclonal Fab 241.14 binding the anchor epitope of H1 HA
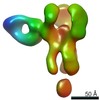
EMDB-25644: 
Negative stain map of polyclonal Fab 241.14 binding the esterase epitope of H1 HA

EMDB-25645: 
Negative stain map of polyclonal Fab 241.14 binding an epitope on the top of the head of H1 HA

EMDB-25646: 
Negative stain map of polyclonal Fab 241.14 binding the RBS epitope of H1 HA

EMDB-25655: 
CryoEM map of anchor 222-1C06 Fab and lateral patch 2B05 Fab binding H1 HA

PDB-7t3d: 
CryoEM map of anchor 222-1C06 Fab and lateral patch 2B05 Fab binding H1 HA

EMDB-12007: 
Programmable icosahedral shell system for virus trapping: DNA-origami half octahedron

EMDB-12008: 
Programmable icosahedral shell system for virus trapping: DNA-origami T=9 triangle (pentamer)

EMDB-12009: 
Programmable icosahedral shell system for virus trapping: DNA-origami octahedron triangle

EMDB-12010: 
Programmable icosahedral shell system for virus trapping: DNA-origami T=1 triangle

EMDB-12011: 
Programmable icosahedral shell system for virus trapping: DNA-origami T=3 triangle

EMDB-12012: 
Programmable icosahedral shell system for virus trapping: DNA-origami T=4 triangle (isosceles)

EMDB-12013: 
Programmable icosahedral shell system for virus trapping: DNA-origami T=4 triangle (equilateral)

EMDB-12014: 
Programmable icosahedral shell system for virus trapping: DNA-origami T=9 triangle (hexamer 1)

EMDB-12015: 
Programmable icosahedral shell system for virus trapping: DNA-origami T=9 triangle (hexamer 2)

EMDB-12016: 
Programmable icosahedral shell system for virus trapping: DNA-origami octahedron shell

EMDB-12019: 
Programmable icosahedral shell system for virus trapping: DNA-origami T=3 shell

EMDB-12020: 
Programmable icosahedral shell system for virus trapping: DNA-origami T=4 shell

EMDB-12021: 
Programmable icosahedral shell system for virus trapping: DNA-origami T=1 shell

EMDB-12022: 
Programmable icosahedral shell system for virus trapping: T=1 shell trapping a HBV core particle

EMDB-12023: 
Programmable icosahedral shell system for virus trapping: DNA-origami T=1 shell with pentagonal aperture

EMDB-12024: 
Programmable icosahedral shell system for virus trapping: DNA-origami T=1 shell (at 25mM MgCl2)

EMDB-12044: 
Programmable icosahedral shell system for virus trapping: DNA-origami half octahedron shell trapping a Hepatitis B core particle

EMDB-12045: 
Programmable icosahedral shell system for virus trapping: DNA-origami half T=1 shell

EMDB-12046: 
Programmable icosahedral shell system for virus trapping: DNA-origami triangular brick

EMDB-12049: 
Programmable icosahedral shell system for virus trapping: spiky DNA-origami T=1 shell
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
