-Search query
-Search result
Showing all 44 items for (author: hamel & v)

EMDB-56682: 
In situ ribosome structure from environmental sample of Pseudo-nitzschia
Method: subtomogram averaging / : Leisch N, Pyle E

EMDB-55390: 
In situ cryo-electron tomogram of a mouse rod photoreceptor cell containing the centriolar luminal distal ring
Method: electron tomography / : Mukherjee S, Daraspe J, Genoud C, Hamel V, Guichard P

EMDB-55391: 
In situ cryo-electron tomogram of a mouse rod photoreceptor cell containing the centriolar luminal distal ring
Method: electron tomography / : Mukherjee S, Daraspe J, Genoud C, Hamel V, Guichard P

EMDB-55393: 
Subtomogram average of the hook density between microtubule doublets/triplets
Method: subtomogram averaging / : McCafferty C, van den Hoek HG, Righetto RD, Van der Stappen P, Mueller A, Stearns T, Engel BD

EMDB-55394: 
Subtomogram average of the luminal distal ring from MTEC centrioles
Method: subtomogram averaging / : McCafferty C, van den Hoek HG, Righetto RD, Mueller A, Van der Stappen P, Stearns T, Engel BD

EMDB-55386: 
Tomogram of a mouse tracheal epithelial cell containing the C2CD3 luminal ring protein
Method: electron tomography / : van den Hoek HG, McCafferty C, Righetto RD, Stearns T, Engel BD

EMDB-19790: 
Restriction on Ku Inward Translocation Caps Telomere Ends
Method: single particle / : Mattarocci S, Baconnais S, Roisne-Hamelin F, Pobiega S, Alibert O, Morin V, Deshayes A, Veaute X, Ropars V, Mazon G, Busso D, Fernandez Varela P, Le Cam E, Charbonnier J, Cuniasse P, Marcand S

EMDB-19811: 
Restriction on Ku Inward Translocation Caps Telomere Ends
Method: single particle / : Mattarocci S, Baconnais S, Roisne-Hamelin F, Pobiega S, Alibert O, Morin V, Deshayes A, Veaute X, Ropars V, Mazon G, Busso D, Fernandez Varela P, Le Cam E, Charbonnier J, Cuniasse P, Marcand S

PDB-8s82: 
Restriction on Ku Inward Translocation Caps Telomere Ends
Method: single particle / : Mattarocci S, Baconnais S, Roisne-Hamelin F, Pobiega S, Alibert O, Morin V, Deshayes A, Veaute X, Ropars V, Mazon G, Busso D, Fernandez Varela P, Le Cam E, Charbonnier J, Cuniasse P, Marcand S

PDB-8s8p: 
Restriction on Ku Inward Translocation Caps Telomere Ends
Method: single particle / : Mattarocci S, Baconnais S, Roisne-Hamelin F, Pobiega S, Alibert O, Morin V, Deshayes A, Veaute X, Ropars V, Mazon G, Busso D, Fernandez Varela P, Le Cam E, Charbonnier J, Cuniasse P, Marcand S

EMDB-42247: 
Degrader-induced complex between PTPN2 and CRBN-DDB1
Method: single particle / : Catalano C, Bratkowski M, Scapin G, Hao Q

PDB-8uh6: 
Degrader-induced complex between PTPN2 and CRBN-DDB1
Method: single particle / : Catalano C, Bratkowski M, Scapin G, Hao Q

EMDB-15609: 
E. coli Wadjet JetABC monomer
Method: single particle / : Roisne-Hamelin F, Beckert B, Li Y, Myasnikov A, Gruber S

EMDB-16020: 
E. coli Wadjet JetABC dimer of dimers
Method: single particle / : Roisne-Hamelin F, Beckert B, Myasnikov A, Li Y, Gruber S

PDB-8as8: 
E. coli Wadjet JetABC monomer
Method: single particle / : Roisne-Hamelin F, Beckert B, Li Y, Myasnikov A, Gruber S

PDB-8bfn: 
E. coli Wadjet JetABC dimer of dimers
Method: single particle / : Roisne-Hamelin F, Beckert B, Myasnikov A, Li Y, Gruber S

EMDB-15252: 
In situ subtomogram average of the C. reinhardtii stellate at the ciliary transition zone
Method: subtomogram averaging / : van den Hoek HG, Righetto RD, Schaffer M, Erdmann PS, Wan WN, Plitzko JM, Baumeister W, Engel BD

EMDB-15253: 
In situ subtomogram average of the C. reinhardtii Y-link at the ciliary transition zone
Method: subtomogram averaging / : van den Hoek HG, Righetto RD, Schaffer M, Erdmann PS, Wan WN, Plitzko JM, Baumeister W, Engel BD

EMDB-15254: 
In situ subtomogram average of the C. reinhardtii MTD sleeve at the ciliary transition zone
Method: subtomogram averaging / : van den Hoek HG, Righetto RD, Schaffer M, Erdmann PS, Wan WN, Plitzko JM, Baumeister W, Engel BD

EMDB-15255: 
In situ subtomogram average of the C. reinhardtii MTD at the ciliary transition zone
Method: subtomogram averaging / : van den Hoek HG, Righetto RD, Schaffer M, Erdmann PS, Wan WN, Plitzko JM, Baumeister W, Engel BD

EMDB-15256: 
Composite map of the C. reinhardtii ciliary transition zone (structures attached to a single MTD) from in situ subtomogram averaging
Method: subtomogram averaging / : van den Hoek HG, Righetto RD, Schaffer M, Erdmann PS, Wan WN, Plitzko JM, Baumeister W, Engel BD

EMDB-15257: 
Composite map of the C. reinhardtii ciliary transition zone (full 9-fold assembly) from in situ subtomogram averaging
Method: subtomogram averaging / : van den Hoek HG, Righetto RD, Schaffer M, Erdmann PS, Wan WN, Plitzko JM, Baumeister W, Engel BD

EMDB-15258: 
In situ subtomogram average of C. reinhardtii IFT-B at the ciliary transition zone
Method: subtomogram averaging / : van den Hoek HG, Jordan MA, Alvarez Viar G, Righetto RD, Schaffer M, Erdmann PS, Wan WN, Plitzko JM, Baumeister W, Pigino G, Engel BD

EMDB-15259: 
In situ subtomogram average of C. reinhardtii IFT-A at the ciliary transition zone
Method: subtomogram averaging / : van den Hoek HG, Jordan MA, Alvarez Viar G, Righetto RD, Schaffer M, Erdmann PS, Wan WN, Plitzko JM, Baumeister W, Pigino G, Engel BD

EMDB-15260: 
In situ subtomogram average of C. reinhardtii dynein-1b at the ciliary transition zone
Method: subtomogram averaging / : van den Hoek HG, Jordan MA, Alvarez Viar G, Righetto RD, Schaffer M, Erdmann PS, Wan WN, Plitzko JM, Baumeister W, Pigino G, Engel BD

EMDB-15261: 
Composite map of the C. reinhardtii IFT (segment of a fully assembled train) at the ciliary transition zone
Method: subtomogram averaging / : van den Hoek HG, Jordan MA, Alvarez Viar G, Righetto RD, Schaffer M, Erdmann PS, Wan WN, Plitzko JM, Baumeister W, Pigino G, Engel BD

EMDB-15262: 
Tomogram #3 of the C. reinhardtii ciliary transition zone
Method: electron tomography / : van den Hoek HG, Schaffer M, Erdmann PS, Wan WN, Plitzko JM, Baumeister W, Engel BD

EMDB-15263: 
Tomogram #12 of the C. reinhardtii ciliary transition zone
Method: electron tomography / : van den Hoek HG, Schaffer M, Erdmann PS, Wan WN, Plitzko JM, Baumeister W, Engel BD

EMDB-23792: 
CryoEM structure of monoclonal Fab 045-09 2B05 binding the lateral patch of influenza virus H1 HA
Method: single particle / : Han J, Ward A
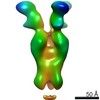
EMDB-23793: 
Negative stain map of monoclonal Fab SFV009 2G01 binding the RBS of H1 HA
Method: single particle / : Han J, Ward AB

EMDB-23794: 
Negative stain map of monoclonal Fab 045-09 2B05 binding the lateral patch of H1 HA
Method: single particle / : Han J, Ward AB

EMDB-23795: 
Negative stain map of monoclonal Fab SFV019 2A06 binding the lateral patch of H1 HA
Method: single particle / : Han J, Ward AB
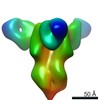
EMDB-23796: 
Negative stain map of monoclonal Fab SFV015 2F02 binding the lateral patch of H1 HA
Method: single particle / : Han J, Ward AB

EMDB-23797: 
Negative stain map of monoclonal Fab 047-09 4G02 binding the lateral patch of H1 HA
Method: single particle / : Han J, Ward AB
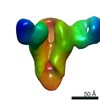
EMDB-23798: 
Negative stain map of monoclonal Fab 047-09 4B06 binding the lateral patch of H1 HA
Method: single particle / : Han J, Ward AB

PDB-7mem: 
CryoEM structure of monoclonal Fab 045-09 2B05 binding the lateral patch of influenza virus H1 HA
Method: single particle / : Han J, Ward A

EMDB-10726: 
Electron cryo-tomography reconstruction and subvolume averaging of the A-C linker from the end of the proximal region of Paramecium tetraurelia basal body
Method: subtomogram averaging / : Guichard P, Hamel V, Le Guennec M

EMDB-10727: 
Electron cryo-tomography reconstruction and subvolume averaging of the A-C linker from the start of the proximal region of Paramecium tetraurelia basal body
Method: subtomogram averaging / : Guichard P, Hamel V, Le Guennec M

EMDB-10728: 
Electron cryo-tomography reconstruction and subvolume averaging of the microtubule triplet from the end of the proximal region of Paramecium tetraurelia basal body
Method: subtomogram averaging / : Guichard P, Hamel V, Le Guennec M

EMDB-10729: 
Electron cryo-tomography reconstruction and subvolume averaging of the microtubule triplet from the start of the proximal region of Paramecium tetraurelia basal body
Method: subtomogram averaging / : Guichard P, Hamel V, Le Guennec M

EMDB-4926: 
Microtubule triplet of isolated Paramecium tetraurelia centriole - inner core
Method: subtomogram averaging / : Le Guennec M, Klena N, Tassin AM, Van der Hoek H, Erdmann PS, Schaffer M, Kovacik L, Goldie KN, Stahlberg H, Engel BD, Hamel V, Guichard P

EMDB-4927: 
Junction between microtubule triplet of isolated Paramecium tetraurelia centriole - inner core region
Method: subtomogram averaging / : Le Guennec M, Klena N, Tassin AM, Van der Hoek H, Erdmann PS, Schaffer M, Kovacik L, Goldie KN, Stahlberg H, Engel BD, Hamel V, Guichard P

EMDB-4929: 
Junction between microtubules triplets of in situ Chlamydomonas reinardtii centriole - inner core region
Method: subtomogram averaging / : Le Guennec M, Klena N, Tassin AM, Van der Hoek H, Erdmann PS, Schaffer M, Kovacik L, Goldie KN, Stahlberg H, Engel BD, Hamel V, Guichard P

EMDB-4930: 
Microtubules triplet of in situ Chlamydomonas reinhardtii centriole - inner core region
Method: subtomogram averaging / : Le Guennec M, Klena N, Tassin AM, Van der Hoek H, Erdmann PS, Schaffer M, Kovacik L, Goldie KN, Stahlberg H, Engel BD, Hamel V, Guichard P
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
