-Search query
-Search result
Showing 1 - 50 of 142 items for (author: bianchi & s)

EMDB-27177: 
sd1.040 Fab in complex with SARS-CoV-2 Spike 2P glycoprotein
Method: single particle / : Abernathy ME, Barnes CO

PDB-8d48: 
sd1.040 Fab in complex with SARS-CoV-2 Spike 2P glycoprotein
Method: single particle / : Abernathy ME, Barnes CO

EMDB-23523: 
SARS-2 CoV 6P Mut7 in complex with Fab J08
Method: single particle / : Torres JL, Ward AB

EMDB-23524: 
SARS-2 CoV 6P Mut7 in complex with Fab F05
Method: single particle / : Torres JL, Ward AB

EMDB-23525: 
SARS-2 CoV 6P Mut 7 in complex with fragment antigen binding (fab) I14
Method: single particle / : Torres JL, Ward AB

EMDB-24607: 
SARS-CoV-2 spike in complex with the S2X58 neutralizing antibody Fab fragment (two receptor-binding domains open)
Method: single particle / : Park YJ, Seattle Structural Genomics Center for Infectious Disease (SSGCID), Veesler D

EMDB-24608: 
SARS-CoV-2 spike in complex with the S2X58 neutralizing antibody Fab fragment (three receptor-binding domains open)
Method: single particle / : Park YJ, Seattle Structural Genomics Center for Infectious Disease (SSGCID), Veesler D

EMDB-24533: 
SARS-CoV-2 spike protein bound to the S2P6 and S2M11 Fab fragments
Method: single particle / : Sauer MM, Seattle Structural Genomics Center for Infectious Disease (SSGCID), Veesler D

EMDB-24236: 
SARS-CoV-2 S (B.1.429 / epsilon variant) + S2M11 + S2L20 Global Refinement
Method: single particle / : McCallum M, Veesler D, Seattle Structural Genomics Center for Infectious Disease (SSGCID)

EMDB-24237: 
SARS-CoV-2 S (B.1.429 / epsilon variant) + S2M11 + S2L20 (Local Refinement of the NTD/S2L20)
Method: single particle / : McCallum M, Seattle Structural Genomics Center for Infectious Disease (SSGCID), Veesler D

PDB-7n8h: 
SARS-CoV-2 S (B.1.429 / epsilon variant) + S2M11 + S2L20 Global Refinement
Method: single particle / : McCallum M, Veesler D, Seattle Structural Genomics Center for Infectious Disease (SSGCID)

PDB-7n8i: 
SARS-CoV-2 S (B.1.429 / epsilon variant) + S2M11 + S2L20 (Local Refinement of the NTD/S2L20)
Method: single particle / : McCallum M, Seattle Structural Genomics Center for Infectious Disease (SSGCID), Veesler D

EMDB-23577: 
SARS-CoV-2 S/S2M11/S2X333 Local Refinement
Method: single particle / : McCallum M, Veesler D, Seattle Structural Genomics Center for Infectious Disease (SSGCID)

EMDB-23578: 
SARS-CoV-2 S/S2M11/S2L28 Local Refinement
Method: single particle / : McCallum M, Veesler D, Seattle Structural Genomics Center for Infectious Disease (SSGCID)

EMDB-23579: 
SARS-CoV-2 S/S2M11/S2X333 Global Refinement
Method: single particle / : McCallum M, Veesler D, Seattle Structural Genomics Center for Infectious Disease (SSGCID)

EMDB-23580: 
SARS-CoV-2 S/S2M11/S2L28 Global Refinement
Method: single particle / : McCallum M, Veesler D, Seattle Structural Genomics Center for Infectious Disease (SSGCID)

EMDB-23581: 
SARS-CoV-2 S/S2M11/S2M28 Local Refinement
Method: single particle / : McCallum M, Veesler D, Seattle Structural Genomics Center for Infectious Disease (SSGCID)

EMDB-23582: 
SARS-CoV-2 S/S2M11/S2M28 Global Refinement
Method: single particle / : McCallum M, Veesler D, Seattle Structural Genomics Center for Infectious Disease (SSGCID)

EMDB-23583: 
SARS-CoV-2 S/S2M11/S2M24 map
Method: single particle / : McCallum M, Veesler D

EMDB-23584: 
SARS-CoV-2 S/S2X28 map
Method: single particle / : McCallum M, Veesler D

EMDB-23585: 
SARS-CoV-2 S/S2M11/S2X316 map
Method: single particle / : McCallum M, Veesler D

EMDB-23586: 
SARS-CoV-2 S/S2M11/S2L20 map
Method: single particle / : McCallum M, Veesler D

PDB-7lxw: 
SARS-CoV-2 S/S2M11/S2X333 Local Refinement
Method: single particle / : McCallum M, Veesler D, Seattle Structural Genomics Center for Infectious Disease (SSGCID)

PDB-7lxx: 
SARS-CoV-2 S/S2M11/S2L28 Local Refinement
Method: single particle / : McCallum M, Veesler D, Seattle Structural Genomics Center for Infectious Disease (SSGCID)

PDB-7lxy: 
SARS-CoV-2 S/S2M11/S2X333 Global Refinement
Method: single particle / : McCallum M, Veesler D, Seattle Structural Genomics Center for Infectious Disease (SSGCID)

PDB-7lxz: 
SARS-CoV-2 S/S2M11/S2L28 Global Refinement
Method: single particle / : McCallum M, Veesler D, Seattle Structural Genomics Center for Infectious Disease (SSGCID)
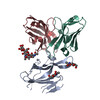
PDB-7ly0: 
SARS-CoV-2 S/S2M11/S2M28 Local Refinement
Method: single particle / : McCallum M, Veesler D, Seattle Structural Genomics Center for Infectious Disease (SSGCID)

PDB-7ly2: 
SARS-CoV-2 S/S2M11/S2M28 Global Refinement
Method: single particle / : McCallum M, Veesler D, Seattle Structural Genomics Center for Infectious Disease (SSGCID)

EMDB-22925: 
cryo-EM structure of SARS-CoV-2 spike in complex with Fab 15033-7, two RBDs bound
Method: single particle / : Li Z, Rini JM

EMDB-22926: 
cryo-EM structure of SARS-CoV-2 spike in complex with Fab 15033-7, three RBDs bound
Method: single particle / : Li Z, Rini JM

PDB-7kmk: 
cryo-EM structure of SARS-CoV-2 spike in complex with Fab 15033-7, two RBDs bound
Method: single particle / : Li Z, Rini JM

PDB-7kml: 
cryo-EM structure of SARS-CoV-2 spike in complex with Fab 15033-7, three RBDs bound
Method: single particle / : Li Z, Rini JM

EMDB-23064: 
SARS-CoV-2 spike protein in complex with Fab 15033-7, 3-"up", asymmetric
Method: single particle / : Li Z, Rini J

EMDB-23065: 
SARS-CoV-2 spike protein in complex with Fab 15033-7, 2-"up"-1-"down" conformation
Method: single particle / : Li Z, Rini J

PDB-7kxj: 
SARS-CoV-2 spike protein in complex with Fab 15033-7, 3-"up", asymmetric
Method: single particle / : Li Z, Rini J

PDB-7kxk: 
SARS-CoV-2 spike protein in complex with Fab 15033-7, 2-"up"-1-"down" conformation
Method: single particle / : Li Z, Rini J

EMDB-22659: 
SARS-CoV-2 spike in complex with the S2M11 neutralizing antibody Fab fragment
Method: single particle / : Tortorici MA, Veesler D, Seattle Structural Genomics Center for Infectious Disease (SSGCID)

EMDB-22660: 
SARS-CoV-2 spike in complex with the S2E12 neutralizing antibody Fab fragment (local refinement of the RBD and Fab variable domains)
Method: single particle / : Tortorici MA, Veesler D, Seattle Structural Genomics Center for Infectious Disease (SSGCID)

EMDB-22668: 
SARS-CoV-2 spike in complex with the S2E12 neutralizing antibody Fab fragment
Method: single particle / : Tortorici M, Veesler D, Seattle Structural Genomics Center for Infectious Disease (SSGCID)

PDB-7k43: 
SARS-CoV-2 spike in complex with the S2M11 neutralizing antibody Fab fragment
Method: single particle / : Tortorici MA, Veesler D, Seattle Structural Genomics Center for Infectious Disease (SSGCID)

PDB-7k45: 
SARS-CoV-2 spike in complex with the S2E12 neutralizing antibody Fab fragment (local refinement of the RBD and Fab variable domains)
Method: single particle / : Tortorici MA, Veesler D, Seattle Structural Genomics Center for Infectious Disease (SSGCID)

PDB-7k4n: 
SARS-CoV-2 spike in complex with the S2E12 neutralizing antibody Fab fragment
Method: single particle / : Tortorici M, Veesler D, Seattle Structural Genomics Center for Infectious Disease (SSGCID)

EMDB-21864: 
Structure of the SARS-CoV-2 spike glycoprotein in complex with the S309 neutralizing antibody Fab fragment
Method: single particle / : Pinto D, Park YJ, Beltramello M, Walls AC, Tortorici MA, Bianchi S, Jaconi S, Culap K, Zatta F, De Marco A, Peter A, Guarino B, Spreafico R, Cameroni E, Case JB, Chen RE, Havenar-Daughton C, Snell G, Virgin HW, Lanzavecchia A, Diamond MS, Fink K, Veesler D, Corti D, Seattle Structural Genomics Center for Infectious Disease (SSGCID)

EMDB-21865: 
Structure of the SARS-CoV-2 spike glycoprotein in complex with the S309 neutralizing antibody Fab fragment (open state)
Method: single particle / : Pinto D, Park YJ, Beltramello M, Walls AC, Tortorici MA, Bianchi S, Jaconi S, Culap K, Zatta F, De Marco A, Peter A, Guarino B, Spreafico R, Cameroni E, Case JB, Chen RE, Havenar-Daughton C, Snell G, Virgin HW, Lanzavecchia A, Diamond MS, Fink K, Veesler D, Corti D, Seattle Structural Genomics Center for Infectious Disease (SSGCID)

PDB-6wps: 
Structure of the SARS-CoV-2 spike glycoprotein in complex with the S309 neutralizing antibody Fab fragment
Method: single particle / : Pinto D, Park YJ, Beltramello M, Walls AC, Tortorici MA, Bianchi S, Jaconi S, Culap K, Zatta F, De Marco A, Peter A, Guarino B, Spreafico R, Cameroni E, Case JB, Chen RE, Havenar-Daughton C, Snell G, Virgin HW, Lanzavecchia A, Diamond MS, Fink K, Veesler D, Corti D, Seattle Structural Genomics Center for Infectious Disease (SSGCID)

PDB-6wpt: 
Structure of the SARS-CoV-2 spike glycoprotein in complex with the S309 neutralizing antibody Fab fragment (open state)
Method: single particle / : Pinto D, Park YJ, Beltramello M, Walls AC, Tortorici MA, Bianchi S, Jaconi S, Culap K, Zatta F, De Marco A, Peter A, Guarino B, Spreafico R, Cameroni E, Case JB, Chen RE, Havenar-Daughton C, Snell G, Virgin HW, Lanzavecchia A, Diamond MS, Fink K, Veesler D, Corti D, Seattle Structural Genomics Center for Infectious Disease (SSGCID)

EMDB-20396: 
Cryo-EM Reconstruction of Polyclonal Serum Fab From a Protected Rhesus Macaque Complexed with BG505 SOSIP.664
Method: single particle / : Nogal B, Ward AB

EMDB-20399: 
Unimmunized control rhesus macaque BZ05 (challenged with SHIV) serum fab complexed with BG505 SOSIP.664 trimer
Method: single particle / : Nogal B, Ward AB
EMDB-20400: 
Unimmunized control rhesus macaque BZ11 (challenged with SHIV) serum fab complexed with BG505 SOSIP.664 trimer
Method: single particle / : Nogal B, Ward AB

EMDB-20401: 
Unimmunized control rhesus macaque BZ13 (challenged with SHIV) serum fab complexed with BG505 SOSIP.664 trimer
Method: single particle / : Nogal B, Ward AB
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
