+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-9567 | |||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
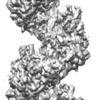
| Title | Human RAD51 post-synaptic complexes | |||||||||||||||||||||
 Map data Map data | ||||||||||||||||||||||
 Sample Sample |
| |||||||||||||||||||||
 Keywords Keywords | DNA repair / ATPase / homologous recombination / DNA BINDING PROTEIN-DNA complex | |||||||||||||||||||||
| Function / homology |  Function and homology information Function and homology informationpresynaptic intermediate filament cytoskeleton / response to glucoside / mitotic recombination-dependent replication fork processing / DNA recombinase assembly / chromosome organization involved in meiotic cell cycle / telomere maintenance via telomere lengthening / double-strand break repair involved in meiotic recombination / nuclear ubiquitin ligase complex / cellular response to cisplatin / DNA strand invasion ...presynaptic intermediate filament cytoskeleton / response to glucoside / mitotic recombination-dependent replication fork processing / DNA recombinase assembly / chromosome organization involved in meiotic cell cycle / telomere maintenance via telomere lengthening / double-strand break repair involved in meiotic recombination / nuclear ubiquitin ligase complex / cellular response to cisplatin / DNA strand invasion / mitotic recombination / cellular response to hydroxyurea / cellular response to camptothecin / replication-born double-strand break repair via sister chromatid exchange / lateral element / DNA strand exchange activity / regulation of DNA damage checkpoint / Impaired BRCA2 binding to PALB2 / telomere maintenance via recombination / reciprocal meiotic recombination / single-stranded DNA helicase activity / HDR through Single Strand Annealing (SSA) / ATP-dependent DNA damage sensor activity / regulation of double-strand break repair via homologous recombination / Homologous DNA Pairing and Strand Exchange / Defective homologous recombination repair (HRR) due to BRCA1 loss of function / Defective HDR through Homologous Recombination Repair (HRR) due to PALB2 loss of BRCA1 binding function / Defective HDR through Homologous Recombination Repair (HRR) due to PALB2 loss of BRCA2/RAD51/RAD51C binding function / Resolution of D-loop Structures through Synthesis-Dependent Strand Annealing (SDSA) / nuclear chromosome / Resolution of D-loop Structures through Holliday Junction Intermediates / Transcriptional Regulation by E2F6 / Impaired BRCA2 binding to RAD51 / replication fork processing / Presynaptic phase of homologous DNA pairing and strand exchange / response to X-ray / ATP-dependent activity, acting on DNA / interstrand cross-link repair / condensed chromosome / DNA polymerase binding / condensed nuclear chromosome / cellular response to ionizing radiation / male germ cell nucleus / meiotic cell cycle / cellular response to gamma radiation / protein-DNA complex / PML body / double-strand break repair via homologous recombination / Meiotic recombination / HDR through Homologous Recombination (HRR) / Hydrolases; Acting on acid anhydrides; Acting on acid anhydrides to facilitate cellular and subcellular movement / response to toxic substance / single-stranded DNA binding / site of double-strand break / double-stranded DNA binding / DNA recombination / chromosome, telomeric region / response to xenobiotic stimulus / mitochondrial matrix / DNA repair / DNA damage response / chromatin binding / centrosome / chromatin / nucleolus / perinuclear region of cytoplasm / enzyme binding / protein-containing complex / mitochondrion / nucleoplasm / ATP binding / identical protein binding / nucleus / cytoplasm / cytosol Similarity search - Function | |||||||||||||||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||||||||||||||
| Method | helical reconstruction / cryo EM / Resolution: 4.5 Å | |||||||||||||||||||||
 Authors Authors | Xu J / Zhao L | |||||||||||||||||||||
| Funding support |  China, China,  United States, 6 items United States, 6 items
| |||||||||||||||||||||
 Citation Citation |  Journal: Nat Struct Mol Biol / Year: 2017 Journal: Nat Struct Mol Biol / Year: 2017Title: Cryo-EM structures of human RAD51 recombinase filaments during catalysis of DNA-strand exchange. Authors: Jingfei Xu / Lingyun Zhao / Yuanyuan Xu / Weixing Zhao / Patrick Sung / Hong-Wei Wang /   Abstract: The central step in eukaryotic homologous recombination (HR) is ATP-dependent DNA-strand exchange mediated by the Rad51 recombinase. In this process, Rad51 assembles on single-stranded DNA (ssDNA) ...The central step in eukaryotic homologous recombination (HR) is ATP-dependent DNA-strand exchange mediated by the Rad51 recombinase. In this process, Rad51 assembles on single-stranded DNA (ssDNA) and generates a helical filament that is able to search for and invade homologous double-stranded DNA (dsDNA), thus leading to strand separation and formation of new base pairs between the initiating ssDNA and the complementary strand within the duplex. Here, we used cryo-EM to solve the structures of human RAD51 in complex with DNA molecules, in presynaptic and postsynaptic states, at near-atomic resolution. Our structures reveal both conserved and distinct structural features of the human RAD51-DNA complexes compared with their prokaryotic counterpart. Notably, we also captured the structure of an arrested synaptic complex. Our results provide new insight into the molecular mechanisms of the DNA homology search and strand-exchange processes. | |||||||||||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_9567.map.gz emd_9567.map.gz | 56.7 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-9567-v30.xml emd-9567-v30.xml emd-9567.xml emd-9567.xml | 18.4 KB 18.4 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_9567.png emd_9567.png | 96.8 KB | ||
| Filedesc metadata |  emd-9567.cif.gz emd-9567.cif.gz | 6.5 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-9567 http://ftp.pdbj.org/pub/emdb/structures/EMD-9567 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-9567 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-9567 | HTTPS FTP |
-Related structure data
| Related structure data |  5h1cMC  9566C  9568C  5h1bC C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_9567.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_9567.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.306 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : Human RAD51 post-synaptic complex
| Entire | Name: Human RAD51 post-synaptic complex |
|---|---|
| Components |
|
-Supramolecule #1: Human RAD51 post-synaptic complex
| Supramolecule | Name: Human RAD51 post-synaptic complex / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#3 |
|---|---|
| Molecular weight | Theoretical: 24.18 kDa/nm |
-Supramolecule #2: Human RAD51
| Supramolecule | Name: Human RAD51 / type: complex / ID: 2 / Parent: 1 / Macromolecule list: #1 |
|---|
-Supramolecule #3: DNA
| Supramolecule | Name: DNA / type: complex / ID: 3 / Parent: 1 / Macromolecule list: #2 |
|---|
-Supramolecule #4: DNA
| Supramolecule | Name: DNA / type: complex / ID: 4 / Parent: 1 / Macromolecule list: #3 |
|---|
-Macromolecule #1: DNA repair protein RAD51 homolog 1
| Macromolecule | Name: DNA repair protein RAD51 homolog 1 / type: protein_or_peptide / ID: 1 / Number of copies: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 37.008074 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MAMQMQLEAN ADTSVEEESF GPQPISRLEQ CGINANDVKK LEEAGFHTVE AVAYAPKKEL INIKGISEAK ADKILAEAAK LVPMGFTTA TEFHQRRSEI IQITTGSKEL DKLLQGGIET GSITEMFGEF RTGKTQICHT LAVTCQLPID RGGGEGKAMY I DTEGTFRP ...String: MAMQMQLEAN ADTSVEEESF GPQPISRLEQ CGINANDVKK LEEAGFHTVE AVAYAPKKEL INIKGISEAK ADKILAEAAK LVPMGFTTA TEFHQRRSEI IQITTGSKEL DKLLQGGIET GSITEMFGEF RTGKTQICHT LAVTCQLPID RGGGEGKAMY I DTEGTFRP ERLLAVAERY GLSGSDVLDN VAYARAFNTD HQTQLLYQAS AMMVESRYAL LIVDSATALY RTDYSGRGEL SA RQMHLAR FLRMLLRLAD EFGVAVVITN QVVAQVDGAA MFAADPKKPI GGNIIAHAST TRLYLRKGRG ETRICQIYDS PCL PEAEAM FAINADGVGD AKD UniProtKB: DNA repair protein RAD51 homolog 1 |
-Macromolecule #2: DNA (5'-D(P*TP*TP*TP*TP*TP*TP*TP*TP*T)-3')
| Macromolecule | Name: DNA (5'-D(P*TP*TP*TP*TP*TP*TP*TP*TP*T)-3') / type: dna / ID: 2 / Number of copies: 1 / Classification: DNA |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 2.692778 KDa |
| Sequence | String: (DT)(DT)(DT)(DT)(DT)(DT)(DT)(DT)(DT) |
-Macromolecule #3: DNA (5'-D(P*AP*AP*AP*AP*AP*AP*AP*AP*A)-3')
| Macromolecule | Name: DNA (5'-D(P*AP*AP*AP*AP*AP*AP*AP*AP*A)-3') / type: dna / ID: 3 / Number of copies: 1 / Classification: DNA |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 2.773904 KDa |
| Sequence | String: (DA)(DA)(DA)(DA)(DA)(DA)(DA)(DA)(DA) |
-Macromolecule #4: MAGNESIUM ION
| Macromolecule | Name: MAGNESIUM ION / type: ligand / ID: 4 / Number of copies: 3 / Formula: MG |
|---|---|
| Molecular weight | Theoretical: 24.305 Da |
-Macromolecule #5: PHOSPHOAMINOPHOSPHONIC ACID-ADENYLATE ESTER
| Macromolecule | Name: PHOSPHOAMINOPHOSPHONIC ACID-ADENYLATE ESTER / type: ligand / ID: 5 / Number of copies: 3 / Formula: ANP |
|---|---|
| Molecular weight | Theoretical: 506.196 Da |
| Chemical component information |  ChemComp-ANP: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | helical reconstruction |
| Aggregation state | filament |
- Sample preparation
Sample preparation
| Concentration | 0.075 mg/mL |
|---|---|
| Buffer | pH: 7.5 Details: 25mM Tris-HCl, pH 7.5, 50mM KCl, 1mM dithiothreitol, 1mM AMP-PNP and 2mM MgCl2 |
| Grid | Model: Quantifoil R1.2/1.3 / Material: COPPER / Mesh: 300 / Support film - Material: CARBON / Support film - topology: HOLEY / Support film - Film thickness: 20 / Pretreatment - Type: PLASMA CLEANING / Pretreatment - Time: 30 sec. / Pretreatment - Atmosphere: AIR / Pretreatment - Pressure: 101.325 kPa |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 289 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Detector mode: SUPER-RESOLUTION / Digitization - Dimensions - Width: 7676 pixel / Digitization - Dimensions - Height: 7420 pixel / Digitization - Frames/image: 3-14 / Number grids imaged: 2 / Number real images: 528 / Average exposure time: 8.0 sec. / Average electron dose: 50.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 70.0 µm / Calibrated defocus max: 2.5 µm / Calibrated defocus min: 1.5 µm / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 2.5 µm / Nominal defocus min: 1.5 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Final reconstruction | Number classes used: 78 Applied symmetry - Helical parameters - Δz: 15.8 Å Applied symmetry - Helical parameters - Δ&Phi: 56.18 ° Applied symmetry - Helical parameters - Axial symmetry: C1 (asymmetric) Algorithm: BACK PROJECTION / Resolution.type: BY AUTHOR / Resolution: 4.5 Å / Resolution method: FSC 0.143 CUT-OFF Software:
Number images used: 30194 | ||||||
|---|---|---|---|---|---|---|---|
| Segment selection | Number selected: 41146 | ||||||
| Startup model | Type of model: OTHER Details: Randomly assigned euler angle for each segment and using back projection to generate initial model. | ||||||
| Final angle assignment | Type: NOT APPLICABLE Software:
|
 Movie
Movie Controller
Controller



















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)






















