+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-9535 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM Structure of the PaeCas3-AcrF3 complex | |||||||||
 Map data Map data | Cryo-EM structure of PaeCas3-AcrF3 complex | |||||||||
 Sample Sample |
| |||||||||
| Function / homology |  Function and homology information Function and homology informationhelicase activity / Hydrolases; Acting on acid anhydrides; Acting on acid anhydrides to facilitate cellular and subcellular movement / defense response to virus / Hydrolases; Acting on ester bonds / hydrolase activity / ATP binding / metal ion binding Similarity search - Function | |||||||||
| Biological species |   Pseudomonas phage JBD5 (virus) Pseudomonas phage JBD5 (virus) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 4.5 Å | |||||||||
 Authors Authors | Zhang X / Ma J / Wang Y / Wang J | |||||||||
 Citation Citation |  Journal: Cell Res / Year: 2016 Journal: Cell Res / Year: 2016Title: A CRISPR evolutionary arms race: structural insights into viral anti-CRISPR/Cas responses. Authors: Jiuyu Wang / Jun Ma / Zhi Cheng / Xu Meng / Lilan You / Min Wang / Xinzheng Zhang / Yanli Wang /  | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_9535.map.gz emd_9535.map.gz | 9.7 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-9535-v30.xml emd-9535-v30.xml emd-9535.xml emd-9535.xml | 13.8 KB 13.8 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_9535.png emd_9535.png | 56.5 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-9535 http://ftp.pdbj.org/pub/emdb/structures/EMD-9535 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-9535 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-9535 | HTTPS FTP |
-Related structure data
| Related structure data |  5gqhMC  5gnfC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_9535.map.gz / Format: CCP4 / Size: 10.5 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_9535.map.gz / Format: CCP4 / Size: 10.5 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Cryo-EM structure of PaeCas3-AcrF3 complex | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.82 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
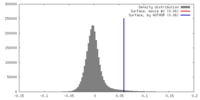
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : PaeCas3-AcrF3 complex
| Entire | Name: PaeCas3-AcrF3 complex |
|---|---|
| Components |
|
-Supramolecule #1: PaeCas3-AcrF3 complex
| Supramolecule | Name: PaeCas3-AcrF3 complex / type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Molecular weight | Theoretical: 158 kDa/nm |
-Supramolecule #2: type I-F CRISPR-associated nuclease/helicase Cas3
| Supramolecule | Name: type I-F CRISPR-associated nuclease/helicase Cas3 / type: complex / ID: 2 / Parent: 1 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:  |
| Recombinant expression | Organism:  |
-Supramolecule #3: anti-CRISPR protein 3 (AcrF3)
| Supramolecule | Name: anti-CRISPR protein 3 (AcrF3) / type: complex / ID: 3 / Parent: 1 / Macromolecule list: #2 |
|---|---|
| Source (natural) | Organism:  Pseudomonas phage JBD5 (virus) Pseudomonas phage JBD5 (virus) |
| Recombinant expression | Organism:  |
-Macromolecule #1: type I-F CRISPR-associated nuclease/helicase Cas3
| Macromolecule | Name: type I-F CRISPR-associated nuclease/helicase Cas3 / type: protein_or_peptide / ID: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Recombinant expression | Organism:  |
| Sequence | String: MGSSHHHHHH SQDPMNILLV SQCEKRALSE TRRILDQFAE RRGERTWQTP ITQAGLDTLR RLLKKSARRN TAVACHWIRG RDHSELLWIV GDASRFNAQG AVPTNRTCRD ILRKEDENDW HSAEDIRLLT VMAALFHDIG KASQAFQAKL RNRGKPMADA YRHEWVSLRL ...String: MGSSHHHHHH SQDPMNILLV SQCEKRALSE TRRILDQFAE RRGERTWQTP ITQAGLDTLR RLLKKSARRN TAVACHWIRG RDHSELLWIV GDASRFNAQG AVPTNRTCRD ILRKEDENDW HSAEDIRLLT VMAALFHDIG KASQAFQAKL RNRGKPMADA YRHEWVSLRL FEAFVGPGSS DEDWLRRLAD KRETGDAWLS QLARDDRQSA PPGPFQKSRL PPLAQAVGWL IVSHHRLPNG DHRGSASLAR LPAPIQSQWC GARDADAKEK AACWQFPHGL PFASAHWRAR TALCAQSMLE RPGLLARGPA LLHDSYVMHV SRLILMLADH HYSSLPADSR LGDPNFPLHA NTDRDSGKLK QRLDEHLLGV ALHSRKLAGT LPRLERQLPR LARHKGFTRR VEQPRFRWQD KAYDCAMACR EQAMEHGFFG LNLASTGCGK TLANGRILYA LADPQRGARF SIALGLRSLT LQTGQAYRER LGLGDDDLAI LVGGSAAREL FEKQQERLER SGSESAQELL AENSHVHFAG TLEDGPLREW LGRNSAGNRL LQAPILACTI DHLMPASESL RGGHQIAPLL RLMTSDLVLD EVDDFDIDDL PALSRLVHWA GLFGSRVLLS SATLPPALVQ GLFEAYRSGR EIFQRHRGAP GRATEIRCAW FDEFSSQSSA HGAVTSFSEA HATFVAQRLA KLEQLPPRRQ AQLCTVHAAG EARPALCREL AGQMNTWMAD LHRCHHTEHQ GRRISFGLLR LANIEPLIEL AQAILAQGAP EGLHVHLCVY HSRHPLLVRS AIERQLDELL KRSDDDAAAL FARPTLAKAL QASTERDHLF VVLASPVAEV GRDHDYDWAI VEPSSMRSII QLAGRIRRHR SGFSGEANLY LLSRNIRSLE GQNPAFQRPG FETPDFPLDS HDLHDLLDPA LLARIDASPR IVEPFPLFPR SRLVDLEHRR LRALMLADDP PSSLLGVPLW WQTPASLSGA LQTSQPFRAG AKERCYALLP DEDDEERLHF SRYEEGTWSN QDNLLRNLDL TYGPRIQTWG TVNYREELVA MAGREDLDLR QCAMRYGEVR LRENTQGWSY HPYLGFKKYN |
-Macromolecule #2: anti-CRISPR protein 3 (AcrF3)
| Macromolecule | Name: anti-CRISPR protein 3 (AcrF3) / type: protein_or_peptide / ID: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Pseudomonas phage JBD5 (virus) Pseudomonas phage JBD5 (virus) |
| Recombinant expression | Organism:  |
| Sequence | String: MSNTISDRIV ARSVIEAARF IQSWEDADPD SLTEDQVLAA AGFAARLHEG LQATVLQRLV DESNHEEYRE FKAWEEALLN ADGRVASSPF ADWGWWYRIA NVMLATASQN VGVTWGSRVH GRLMAIFQDK FKQRYEEQA |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 1 mg/mL |
|---|---|
| Buffer | pH: 7.5 |
| Grid | Model: GiG-322 / Material: COPPER / Mesh: 300 / Support film - Material: CARBON / Support film - topology: HOLEY / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Atmosphere: OTHER |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 16 K / Instrument: LEICA EM GP |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Detector mode: SUPER-RESOLUTION / Average electron dose: 8.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Refinement | Protocol: BACKBONE TRACE |
|---|---|
| Output model |  PDB-5gqh: |
 Movie
Movie Controller
Controller















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)