+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-8069 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Prefusion structure of a human coronavirus spike protein | |||||||||
 Map data Map data | None | |||||||||
 Sample Sample |
| |||||||||
| Function / homology |  Function and homology information Function and homology informationendocytosis involved in viral entry into host cell / virion component / host cell endoplasmic reticulum-Golgi intermediate compartment membrane / receptor-mediated virion attachment to host cell / fusion of virus membrane with host plasma membrane / fusion of virus membrane with host endosome membrane / viral envelope / host cell plasma membrane / virion membrane / identical protein binding / membrane Similarity search - Function | |||||||||
| Biological species |  Human coronavirus HKU1 (isolate N5) / Human coronavirus HKU1 (isolate N5) /  Enterobacteria phage T4 (virus) Enterobacteria phage T4 (virus) | |||||||||
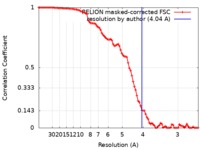
| Method | single particle reconstruction / cryo EM / negative staining / Resolution: 4.04 Å | |||||||||
 Authors Authors | Kirchdoerfer RN / Cottrell CA / Wang N / Pallesen J / Yassine HM / Turner HL / Corbett KS / Graham BS / McLellan JS / Ward AB | |||||||||
 Citation Citation |  Journal: Nature / Year: 2016 Journal: Nature / Year: 2016Title: Pre-fusion structure of a human coronavirus spike protein. Authors: Robert N Kirchdoerfer / Christopher A Cottrell / Nianshuang Wang / Jesper Pallesen / Hadi M Yassine / Hannah L Turner / Kizzmekia S Corbett / Barney S Graham / Jason S McLellan / Andrew B Ward /  Abstract: HKU1 is a human betacoronavirus that causes mild yet prevalent respiratory disease, and is related to the zoonotic SARS and MERS betacoronaviruses, which have high fatality rates and pandemic ...HKU1 is a human betacoronavirus that causes mild yet prevalent respiratory disease, and is related to the zoonotic SARS and MERS betacoronaviruses, which have high fatality rates and pandemic potential. Cell tropism and host range is determined in part by the coronavirus spike (S) protein, which binds cellular receptors and mediates membrane fusion. As the largest known class I fusion protein, its size and extensive glycosylation have hindered structural studies of the full ectodomain, thus preventing a molecular understanding of its function and limiting development of effective interventions. Here we present the 4.0 Å resolution structure of the trimeric HKU1 S protein determined using single-particle cryo-electron microscopy. In the pre-fusion conformation, the receptor-binding subunits, S1, rest above the fusion-mediating subunits, S2, preventing their conformational rearrangement. Surprisingly, the S1 C-terminal domains are interdigitated and form extensive quaternary interactions that occlude surfaces known in other coronaviruses to bind protein receptors. These features, along with the location of the two protease sites known to be important for coronavirus entry, provide a structural basis to support a model of membrane fusion mediated by progressive S protein destabilization through receptor binding and proteolytic cleavage. These studies should also serve as a foundation for the structure-based design of betacoronavirus vaccine immunogens. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_8069.map.gz emd_8069.map.gz | 59.8 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-8069-v30.xml emd-8069-v30.xml emd-8069.xml emd-8069.xml | 22.6 KB 22.6 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_8069_fsc.xml emd_8069_fsc.xml | 9 KB | Display |  FSC data file FSC data file |
| Images |  emd_8069.png emd_8069.png | 81.1 KB | ||
| Others |  emd_8069_additional.map.gz emd_8069_additional.map.gz emd_8069_half_map_1.map.gz emd_8069_half_map_1.map.gz emd_8069_half_map_2.map.gz emd_8069_half_map_2.map.gz | 59 MB 49.7 MB 49.7 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-8069 http://ftp.pdbj.org/pub/emdb/structures/EMD-8069 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-8069 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-8069 | HTTPS FTP |
-Validation report
| Summary document |  emd_8069_validation.pdf.gz emd_8069_validation.pdf.gz | 829.6 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_8069_full_validation.pdf.gz emd_8069_full_validation.pdf.gz | 829.2 KB | Display | |
| Data in XML |  emd_8069_validation.xml.gz emd_8069_validation.xml.gz | 15.7 KB | Display | |
| Data in CIF |  emd_8069_validation.cif.gz emd_8069_validation.cif.gz | 20.7 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-8069 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-8069 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-8069 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-8069 | HTTPS FTP |
-Related structure data
| Related structure data |  5i08MC  8066C  8067C 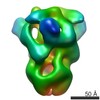 8068C M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_8069.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_8069.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | None | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.31 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Additional map: None
| File | emd_8069_additional.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | None | ||||||||||||
| Projections & Slices |
| ||||||||||||
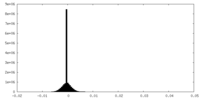
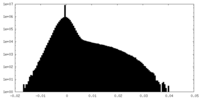
| Density Histograms |
-Half map: None
| File | emd_8069_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | None | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: None
| File | emd_8069_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | None | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : HKU1 spike with attached foldon domain and mutated furin-cleavage site
| Entire | Name: HKU1 spike with attached foldon domain and mutated furin-cleavage site |
|---|---|
| Components |
|
-Supramolecule #1: HKU1 spike with attached foldon domain and mutated furin-cleavage site
| Supramolecule | Name: HKU1 spike with attached foldon domain and mutated furin-cleavage site type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  Human coronavirus HKU1 (isolate N5) Human coronavirus HKU1 (isolate N5) |
| Recombinant expression | Organism:  Homo sapiens (human) / Recombinant cell: FreeStyle 293F / Recombinant plasmid: pVRC8400 Homo sapiens (human) / Recombinant cell: FreeStyle 293F / Recombinant plasmid: pVRC8400 |
| Molecular weight | Theoretical: 420 KDa |
-Macromolecule #1: Spike glycoprotein,Foldon chimera
| Macromolecule | Name: Spike glycoprotein,Foldon chimera / type: protein_or_peptide / ID: 1 / Number of copies: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Enterobacteria phage T4 (virus) Enterobacteria phage T4 (virus) |
| Molecular weight | Theoretical: 144.295891 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: VIGDFNCTNS FINDYNKTIP RISEDVVDVS LGLGTYYVLN RVYLNTTLLF TGYFPKSGAN FRDLALKGSI YLSTLWYKPP FLSDFNNGI FSKVKNTKLY VNNTLYSEFS TIVIGSVFVN TSYTIVVQPH NGILEITACQ YTMCEYPHTV CKSKGSIRNE S WHIDSSEP ...String: VIGDFNCTNS FINDYNKTIP RISEDVVDVS LGLGTYYVLN RVYLNTTLLF TGYFPKSGAN FRDLALKGSI YLSTLWYKPP FLSDFNNGI FSKVKNTKLY VNNTLYSEFS TIVIGSVFVN TSYTIVVQPH NGILEITACQ YTMCEYPHTV CKSKGSIRNE S WHIDSSEP LCLFKKNFTY NVSADWLYFH FYQERGVFYA YYADVGMPTT FLFSLYLGTI LSHYYVMPLT CNAISSNTDN ET LEYWVTP LSRRQYLLNF DEHGVITNAV DCSSSFLSEI QCKTQSFAPN TGVYDLSGFT VKPVATVYRR IPNLPDCDID NWL NNVSVP SPLNWERRIF SNCNFNLSTL LRLVHVDSFS CNNLDKSKIF GSCFNSITVD KFAIPNRRRD DLQLGSSGFL QSSN YKIDI SSSSCQLYYS LPLVNVTINN FNPSSWNRRY GFGSFNLSSY DVVYSDHCFS VNSDFCPCAD PSVVNSCAKS KPPSA ICPA GTKYRHCDLD TTLYVKNWCR CSCLPDPIST YSPNTCPQKK VVVGIGEHCP GLGINEEKCG TQLNHSSCFC SPDAFL GWS FDSCISNNRC NIFSNFIFNG INSGTTCSND LLYSNTEIST GVCVNYDLYG ITGQGIFKEV SAAYYNNWQN LLYDSNG NI IGFKDFLTNK TYTILPCYSG RVSAAFYQNS SSPALLYRNL KCSYVLNNIS FISQPFYFDS YLGCVLNAVN LTSYSVSS C DLRMGSGFCI DYALPSSGGS GSGISSPYRF VTFEPFNVSF VNDSVETVGG LFEIQIPTNF TIAGHEEFIQ TSSPKVTID CSAFVCSNYA ACHDLLSEYG TFCDNINSIL NEVNDLLDIT QLQVANALMQ GVTLSSNLNT NLHSDVDNID FKSLLGCLGS QCGSSSRSL LEDLLFNKVK LSDVGFVEAY NNCTGGSEIR DLLCVQSFNG IKVLPPILSE TQISGYTTAA TVAAMFPPWS A AAGVPFSL NVQYRINGLG VTMDVLNKNQ KLIANAFNKA LLSIQNGFTA TNSALAKIQS VVNANAQALN SLLQQLFNKF GA ISSSLQE ILSRLDNLEA QVQIDRLING RLTALNAYVS QQLSDITLIK AGASRAIEKV NECVKSQSPR INFCGNGNHI LSL VQNAPY GLLFIHFSYK PTSFKTVLVS PGLCLSGDRG IAPKQGYFIK QNDSWMFTGS SYYYPEPISD KNVVFMNSCS VNFT KAPFI YLNNSIPNLS DFEAELSLWF KNHTSIAPNL TFNSHINATF LDLYYEMNVI QESIKSLNGS GYIPEAPRDG QAYVR KDGE WVLLSTFLGL EVLFQ |
-Experimental details
-Structure determination
| Method | negative staining, cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.27 mg/mL | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 8 Component:
| |||||||||
| Staining | Type: NEGATIVE / Material: uranyl formate Details: 3 uL sample was applied to grid for 30 seconds and then blotted. Grids were stained with 3 uL 1% uranyl formate for 60 seconds followed by blotting. | |||||||||
| Grid | Model: EMS CF-2/2-4C C-Flat / Material: COPPER / Mesh: 400 / Support film - Material: CARBON / Support film - topology: HOLEY / Pretreatment - Type: PLASMA CLEANING / Pretreatment - Atmosphere: OTHER | |||||||||
| Vitrification | Cryogen name: ETHANE / Instrument: HOMEMADE PLUNGER Details: 3 uL sample was applied to grid, blotted, and plunged into liquid ethane.. |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Detector mode: COUNTING / Number grids imaged: 1 / Number real images: 1049 / Average exposure time: 10.0 sec. / Average electron dose: 57.0 e/Å2 Details: Images were collected using Legionon and processed using Appion. |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Calibrated magnification: 22500 / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 3.5 µm / Nominal defocus min: 1.0 µm / Nominal magnification: 22500 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Details | Model building and refinement were conducted using a combination of software programs. |
|---|---|
| Refinement | Space: REAL / Protocol: AB INITIO MODEL / Overall B value: 117 / Target criteria: EMRinger |
| Output model |  PDB-5i08: |
 Movie
Movie Controller
Controller
















 Z
Z Y
Y X
X