[English] 日本語
 Yorodumi
Yorodumi- EMDB-6538: Structure of Simian Immunodeficiency Virus Envelope Spikes bound ... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-6538 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Structure of Simian Immunodeficiency Virus Envelope Spikes bound with CD4 and Monoclonal Antibody 36D5 | |||||||||
 Map data Map data | Reconstruction of native SIV envelope spike | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Cryoelectron tomography / image processing / electron microscopy / immunology / AIDS / HIV | |||||||||
| Biological species |  Simian immunodeficiency virus Simian immunodeficiency virus | |||||||||
| Method | subtomogram averaging / cryo EM | |||||||||
 Authors Authors | Hu G / Liu J / Roux KH / Taylor KA | |||||||||
 Citation Citation |  Journal: J Virol / Year: 2017 Journal: J Virol / Year: 2017Title: Structure of Simian Immunodeficiency Virus Envelope Spikes Bound with CD4 and Monoclonal Antibody 36D5. Authors: Guiqing Hu / Jun Liu / Kenneth H Roux / Kenneth A Taylor /  Abstract: The human immunodeficiency virus type 1 (HIV-1)/simian immunodeficiency virus (SIV) envelope spike (Env) mediates viral entry into host cells. The V3 loop of the gp120 component of the Env trimer ...The human immunodeficiency virus type 1 (HIV-1)/simian immunodeficiency virus (SIV) envelope spike (Env) mediates viral entry into host cells. The V3 loop of the gp120 component of the Env trimer contributes to the coreceptor binding site and is a target for neutralizing antibodies. We used cryo-electron tomography to visualize the binding of CD4 and the V3 loop monoclonal antibody (MAb) 36D5 to gp120 of the SIV Env trimer. Our results show that 36D5 binds gp120 at the base of the V3 loop and suggest that the antibody exerts its neutralization effect by blocking the coreceptor binding site. The antibody does this without altering the dynamics of the spike motion between closed and open states when CD4 is bound. The interaction between 36D5 and SIV gp120 is similar to the interaction between some broadly neutralizing anti-V3 loop antibodies and HIV-1 gp120. Two conformations of gp120 bound with CD4 are revealed, suggesting an intrinsic dynamic nature of the liganded Env trimer. CD4 binding substantially increases the binding of 36D5 to gp120 in the intact Env trimer, consistent with CD4-induced changes in the conformation of gp120 and the antibody binding site. Binding by MAb 36D5 does not substantially alter the proportions of the two CD4-bound conformations. The position of MAb 36D5 at the V3 base changes little between conformations, indicating that the V3 base serves as a pivot point during the transition between these two states. Glycoprotein spikes on the surfaces of SIV and HIV are the sole targets available to the immune system for antibody neutralization. Spikes evade the immune system by a combination of a thick layer of polysaccharide on the surface (the glycan shield) and movement between spike domains that masks the epitope conformation. Using SIV virions whose spikes were "decorated" with the primary cellular receptor (CD4) and an antibody (36D5) at part of the coreceptor binding site, we visualized multiple conformations trapped by the rapid freezing step, which were separated using statistical analysis. Our results show that the CD4-induced conformational dynamics of the spike enhances binding of the antibody. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_6538.map.gz emd_6538.map.gz | 153.6 KB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-6538-v30.xml emd-6538-v30.xml emd-6538.xml emd-6538.xml | 8.2 KB 8.2 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_6538.tif emd_6538.tif | 67.8 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-6538 http://ftp.pdbj.org/pub/emdb/structures/EMD-6538 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-6538 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-6538 | HTTPS FTP |
-Related structure data
| Related structure data |  6539C  6540C  6541C  6542C  6543C  3jcbC  3jccC C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_6538.map.gz / Format: CCP4 / Size: 348.6 KB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_6538.map.gz / Format: CCP4 / Size: 348.6 KB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Reconstruction of native SIV envelope spike | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 5.7 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
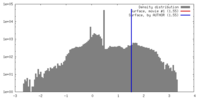
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : Structure of wild SIV envelope
| Entire | Name: Structure of wild SIV envelope |
|---|---|
| Components |
|
-Supramolecule #1000: Structure of wild SIV envelope
| Supramolecule | Name: Structure of wild SIV envelope / type: sample / ID: 1000 / Oligomeric state: native gp120 trimer / Number unique components: 1 |
|---|
-Supramolecule #1: Simian immunodeficiency virus
| Supramolecule | Name: Simian immunodeficiency virus / type: virus / ID: 1 / Name.synonym: SIV / NCBI-ID: 11723 / Sci species name: Simian immunodeficiency virus / Sci species strain: SIV239/251tail/Supt-CCR5 CL.30 / Database: NCBI / Virus type: VIRION / Virus isolate: STRAIN / Virus enveloped: Yes / Virus empty: No / Syn species name: SIV |
|---|---|
| Host (natural) | Organism:  Cercopithecidae (Old World monkeys) / synonym: VERTEBRATES Cercopithecidae (Old World monkeys) / synonym: VERTEBRATES |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | subtomogram averaging |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Vitrification | Cryogen name: ETHANE / Instrument: HOMEMADE PLUNGER |
|---|
- Electron microscopy
Electron microscopy
| Microscope | FEI POLARA 300 |
|---|---|
| Date | Aug 16, 2008 |
| Image recording | Category: CCD / Film or detector model: TVIPS TEMCAM-F415 (4k x 4k) / Number real images: 130 / Average electron dose: 100 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 5.0 µm / Nominal defocus min: 4.0 µm / Nominal magnification: 31000 |
| Sample stage | Specimen holder: cryo holder / Specimen holder model: OTHER / Tilt series - Axis1 - Min angle: -65 ° / Tilt series - Axis1 - Max angle: 65 ° |
| Experimental equipment |  Model: Tecnai Polara / Image courtesy: FEI Company |
- Image processing
Image processing
| Final reconstruction | Software - Name: I3, protomo / Number subtomograms used: 3296 |
|---|
-Atomic model buiding 1
| Initial model | PDB ID: |
|---|---|
| Refinement | Space: REAL / Protocol: RIGID BODY FIT |
 Movie
Movie Controller
Controller




 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)