[English] 日本語
 Yorodumi
Yorodumi- EMDB-5765: A new topology of the HK97-like fold revealed in Bordetella bacte... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-5765 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | A new topology of the HK97-like fold revealed in Bordetella bacteriophage by cryoEM at 3.5A resolution | |||||||||
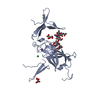
 Map data Map data | Averaged MCP density of Bordetella bacteriophage (EMD-5764) | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Protein topology / HK97 like / cryoEM / bacteriophage | |||||||||
| Biological species |  Bordetella phage BPP-1 (virus) Bordetella phage BPP-1 (virus) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.5 Å | |||||||||
 Authors Authors | Zhang X / Guo H / Jin L / Czornyj E / Hodes A / Hui WH / Nieh AW / Miller JF / Zhou ZH | |||||||||
 Citation Citation |  Journal: Elife / Year: 2013 Journal: Elife / Year: 2013Title: A new topology of the HK97-like fold revealed in Bordetella bacteriophage by cryoEM at 3.5 A resolution. Authors: Xing Zhang / Huatao Guo / Lei Jin / Elizabeth Czornyj / Asher Hodes / Wong H Hui / Angela W Nieh / Jeff F Miller / Z Hong Zhou /  Abstract: Bacteriophage BPP-1 infects and kills Bordetella species that cause whooping cough. Its diversity-generating retroelement (DGR) provides a naturally occurring phage-display system, but engineering ...Bacteriophage BPP-1 infects and kills Bordetella species that cause whooping cough. Its diversity-generating retroelement (DGR) provides a naturally occurring phage-display system, but engineering efforts are hampered without atomic structures. Here, we report a cryo electron microscopy structure of the BPP-1 head at 3.5 Å resolution. Our atomic model shows two of the three protein folds representing major viral lineages: jellyroll for its cement protein (CP) and HK97-like ('Johnson') for its major capsid protein (MCP). Strikingly, the fold topology of MCP is permuted non-circularly from the Johnson fold topology previously seen in viral and cellular proteins. We illustrate that the new topology is likely the only feasible alternative of the old topology. β-sheet augmentation and electrostatic interactions contribute to the formation of non-covalent chainmail in BPP-1, unlike covalent inter-protein linkages of the HK97 chainmail. Despite these complex interactions, the termini of both CP and MCP are ideally positioned for DGR-based phage-display engineering. DOI: http://dx.doi.org/10.7554/eLife.01299.001. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_5765.map.gz emd_5765.map.gz | 218.5 KB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-5765-v30.xml emd-5765-v30.xml emd-5765.xml emd-5765.xml | 9.9 KB 9.9 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_5765.jpg emd_5765.jpg | 145 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-5765 http://ftp.pdbj.org/pub/emdb/structures/EMD-5765 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-5765 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-5765 | HTTPS FTP |
-Validation report
| Summary document |  emd_5765_validation.pdf.gz emd_5765_validation.pdf.gz | 79.2 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_5765_full_validation.pdf.gz emd_5765_full_validation.pdf.gz | 78.3 KB | Display | |
| Data in XML |  emd_5765_validation.xml.gz emd_5765_validation.xml.gz | 492 B | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-5765 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-5765 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-5765 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-5765 | HTTPS FTP |
-Related structure data
| Related structure data |  5764C  5766C  3j4uFC F: fitted*YM C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_5765.map.gz / Format: CCP4 / Size: 29.8 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_5765.map.gz / Format: CCP4 / Size: 29.8 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Averaged MCP density of Bordetella bacteriophage (EMD-5764) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.1 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : Bordetella bacteriophage major capsid protein (MCP)
| Entire | Name: Bordetella bacteriophage major capsid protein (MCP) |
|---|---|
| Components |
|
-Supramolecule #1000: Bordetella bacteriophage major capsid protein (MCP)
| Supramolecule | Name: Bordetella bacteriophage major capsid protein (MCP) / type: sample / ID: 1000 / Details: This is the averaged MCP density of EMD-5764. / Number unique components: 1 |
|---|
-Macromolecule #1: major capsid protein
| Macromolecule | Name: major capsid protein / type: protein_or_peptide / ID: 1 / Name.synonym: MCP / Details: This is the averaged MCP density of EMD-5764. / Recombinant expression: No / Database: NCBI |
|---|---|
| Source (natural) | Organism:  Bordetella phage BPP-1 (virus) / synonym: Bordetella bacteriophage Bordetella phage BPP-1 (virus) / synonym: Bordetella bacteriophage |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 / Details: 50 mM Tris-HCl, 250 mM NaCl |
|---|---|
| Grid | Details: 400 mesh holey carbon film |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Temperature | Average: 80 K |
| Alignment procedure | Legacy - Astigmatism: manual correction / Legacy - Electron beam tilt params: 0 |
| Date | Jul 12, 2009 |
| Image recording | Category: FILM / Film or detector model: KODAK SO-163 FILM / Digitization - Scanner: NIKON SUPER COOLSCAN 9000 / Digitization - Sampling interval: 6.35 µm / Number real images: 895 / Average electron dose: 25 e/Å2 / Bits/pixel: 2 |
| Tilt angle min | 0 |
| Tilt angle max | 0 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Calibrated magnification: 57660 / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 2.3 µm / Nominal defocus min: 0.8 µm / Nominal magnification: 59000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| CTF correction | Details: Each particle |
|---|---|
| Final reconstruction | Algorithm: OTHER / Resolution.type: BY AUTHOR / Resolution: 3.5 Å / Resolution method: OTHER / Software - Name: eLite3D, Frealign / Details: This is the averaged MCP density of EMD-5764. / Number images used: 39549 |
| Final two d classification | Number classes: 1 |
 Movie
Movie Controller
Controller










 Z (Sec.)
Z (Sec.) X (Row.)
X (Row.) Y (Col.)
Y (Col.)





















