[English] 日本語
 Yorodumi
Yorodumi- EMDB-4480: CI Peripheral Arm focused refinement from Ovine respiratory SC I+III2 -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-4480 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | CI Peripheral Arm focused refinement from Ovine respiratory SC I+III2 | |||||||||
 Map data Map data | Focused refinement around complex I peripheral arm | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | complex I / peripheral arm / cellular respiration / mitochondria / ELECTRON TRANSPORT | |||||||||
| Function / homology |  Function and homology information Function and homology information: / : / NADH:ubiquinone reductase (H+-translocating) / apoptotic mitochondrial changes / mitochondrial electron transport, NADH to ubiquinone / acyl binding / mitochondrial respiratory chain complex I assembly / NADH dehydrogenase activity / respiratory chain complex I / NADH dehydrogenase (ubiquinone) activity ...: / : / NADH:ubiquinone reductase (H+-translocating) / apoptotic mitochondrial changes / mitochondrial electron transport, NADH to ubiquinone / acyl binding / mitochondrial respiratory chain complex I assembly / NADH dehydrogenase activity / respiratory chain complex I / NADH dehydrogenase (ubiquinone) activity / acyl carrier activity / ATP metabolic process / reactive oxygen species metabolic process / regulation of mitochondrial membrane potential / respiratory electron transport chain / electron transport chain / circadian rhythm / mitochondrial intermembrane space / 2 iron, 2 sulfur cluster binding / mitochondrial membrane / NAD binding / FMN binding / 4 iron, 4 sulfur cluster binding / response to oxidative stress / mitochondrial inner membrane / mitochondrial matrix / protein-containing complex binding / mitochondrion / membrane / metal ion binding Similarity search - Function | |||||||||
| Biological species |  | |||||||||
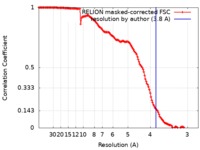
| Method | single particle reconstruction / cryo EM / Resolution: 3.8 Å | |||||||||
 Authors Authors | Letts JA / Sazanov LA | |||||||||
| Funding support |  Austria, 1 items Austria, 1 items
| |||||||||
 Citation Citation |  Journal: Mol Cell / Year: 2019 Journal: Mol Cell / Year: 2019Title: Structures of Respiratory Supercomplex I+III Reveal Functional and Conformational Crosstalk. Authors: James A Letts / Karol Fiedorczuk / Gianluca Degliesposti / Mark Skehel / Leonid A Sazanov /    Abstract: The mitochondrial electron transport chain complexes are organized into supercomplexes (SCs) of defined stoichiometry, which have been proposed to regulate electron flux via substrate channeling. We ...The mitochondrial electron transport chain complexes are organized into supercomplexes (SCs) of defined stoichiometry, which have been proposed to regulate electron flux via substrate channeling. We demonstrate that CoQ trapping in the isolated SC I+III limits complex (C)I turnover, arguing against channeling. The SC structure, resolved at up to 3.8 Å in four distinct states, suggests that CoQ oxidation may be rate limiting because of unequal access of CoQ to the active sites of CIII. CI shows a transition between "closed" and "open" conformations, accompanied by the striking rotation of a key transmembrane helix. Furthermore, the state of CI affects the conformational flexibility within CIII, demonstrating crosstalk between the enzymes. CoQ was identified at only three of the four binding sites in CIII, suggesting that interaction with CI disrupts CIII symmetry in a functionally relevant manner. Together, these observations indicate a more nuanced functional role for the SCs. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_4480.map.gz emd_4480.map.gz | 228.8 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-4480-v30.xml emd-4480-v30.xml emd-4480.xml emd-4480.xml | 45.5 KB 45.5 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_4480_fsc.xml emd_4480_fsc.xml | 14.1 KB | Display |  FSC data file FSC data file |
| Images |  emd_4480.png emd_4480.png | 63.9 KB | ||
| Masks |  emd_4480_msk_1.map emd_4480_msk_1.map | 244.1 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-4480.cif.gz emd-4480.cif.gz | 11 KB | ||
| Others |  emd_4480_half_map_1.map.gz emd_4480_half_map_1.map.gz emd_4480_half_map_2.map.gz emd_4480_half_map_2.map.gz | 194.7 MB 194.7 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-4480 http://ftp.pdbj.org/pub/emdb/structures/EMD-4480 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-4480 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-4480 | HTTPS FTP |
-Related structure data
| Related structure data |  6q9dMC  4479C  4481C  4482C  4493C  4494C  4495C  4496C  4497C  4498C  4499C  4500C  4501C  4502C  4505C  4506C  4507C  6q9bC  6q9eC  6qa9C  6qbxC  6qc2C  6qc3C  6qc4C  6qc5C  6qc6C  6qc7C  6qc8C  6qc9C  6qcaC  6qcfC C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_4480.map.gz / Format: CCP4 / Size: 244.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_4480.map.gz / Format: CCP4 / Size: 244.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
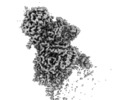
| Annotation | Focused refinement around complex I peripheral arm | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.4 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
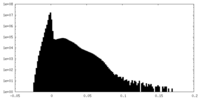
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Mask #1
| File |  emd_4480_msk_1.map emd_4480_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
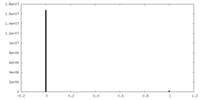
| Density Histograms |
-Half map: Focused refinement around complex I peripheral arm. Half map 1
| File | emd_4480_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Focused refinement around complex I peripheral arm. Half map 1 | ||||||||||||
| Projections & Slices |
| ||||||||||||
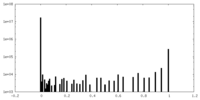
| Density Histograms |
-Half map: Focused refinement around complex I peripheral arm. Half map 2
| File | emd_4480_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Focused refinement around complex I peripheral arm. Half map 2 | ||||||||||||
| Projections & Slices |
| ||||||||||||
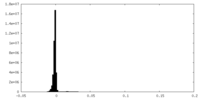
| Density Histograms |
- Sample components
Sample components
+Entire : Ovine mitochondrial SC I+III2
+Supramolecule #1: Ovine mitochondrial SC I+III2
+Macromolecule #1: NADH dehydrogenase [ubiquinone] flavoprotein 1, mitochondrial
+Macromolecule #2: NADH dehydrogenase [ubiquinone] flavoprotein 2, mitochondrial
+Macromolecule #3: NADH:ubiquinone oxidoreductase core subunit S1
+Macromolecule #4: NADH:ubiquinone oxidoreductase core subunit S2,NADH:ubiquinone ox...
+Macromolecule #5: NADH:ubiquinone oxidoreductase core subunit S3
+Macromolecule #6: NADH:ubiquinone oxidoreductase core subunit S7
+Macromolecule #7: NADH:ubiquinone oxidoreductase core subunit S8
+Macromolecule #8: NADH:ubiquinone oxidoreductase subunit V3
+Macromolecule #9: NADH dehydrogenase [ubiquinone] iron-sulfur protein 6, mitochondrial
+Macromolecule #10: NADH:ubiquinone oxidoreductase subunit S4
+Macromolecule #11: NADH:ubiquinone oxidoreductase subunit A9
+Macromolecule #12: NADH dehydrogenase [ubiquinone] 1 alpha subcomplex subunit 2
+Macromolecule #13: NADH dehydrogenase [ubiquinone] 1 alpha subcomplex subunit 5
+Macromolecule #14: NADH:ubiquinone oxidoreductase subunit A6
+Macromolecule #15: NADH dehydrogenase [ubiquinone] 1 alpha subcomplex subunit 7
+Macromolecule #16: NADH dehydrogenase [ubiquinone] 1 alpha subcomplex subunit 12
+Macromolecule #17: Acyl carrier protein
+Macromolecule #18: NADH dehydrogenase [ubiquinone] 1 alpha subcomplex subunit 13
+Macromolecule #19: IRON/SULFUR CLUSTER
+Macromolecule #20: FLAVIN MONONUCLEOTIDE
+Macromolecule #21: FE2/S2 (INORGANIC) CLUSTER
+Macromolecule #22: ZINC ION
+Macromolecule #23: NADPH DIHYDRO-NICOTINAMIDE-ADENINE-DINUCLEOTIDE PHOSPHATE
+Macromolecule #24: S-[2-({N-[(2S)-2-hydroxy-3,3-dimethyl-4-(phosphonooxy)butanoyl]-b...
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 2 mg/mL | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 7.4 Component:
Details: 250 mM NaCl, 20 mM HEPES, pH 7.7, 0.02% Brij-35 | ||||||||||||
| Vitrification | Cryogen name: PROPANE / Chamber humidity: 95 % / Chamber temperature: 277 K / Instrument: FEI VITROBOT MARK IV Details: blotting for 30 seconds at 4 degrees Celsius, 95% humidity and flash freezing. |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: FEI FALCON II (4k x 4k) / Number grids imaged: 1 / Number real images: 1854 / Average exposure time: 2.0 sec. / Average electron dose: 51.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: OTHER / Imaging mode: BRIGHT FIELD |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller

















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)