[English] 日本語
 Yorodumi
Yorodumi- EMDB-44517: Pseudomonas phage DEV neck and tail (portal, head-to-tail and tai... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Pseudomonas phage DEV neck and tail (portal, head-to-tail and tail tube proteins) | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | portal / tail hub / tail tube / complex / STRUCTURAL PROTEIN / gp75 / gp83 / gp80 / VIRAL PROTEIN | |||||||||
| Function / homology | : / Phage SU10 portal protein / Uncharacterized protein / N4 gp54-like protein / Portal protein Function and homology information Function and homology information | |||||||||
| Biological species |  Pseudomonas phage vB_PaeP_DEV (virus) Pseudomonas phage vB_PaeP_DEV (virus) | |||||||||
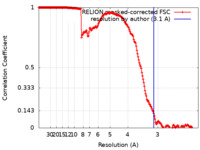
| Method | single particle reconstruction / cryo EM / Resolution: 3.1 Å | |||||||||
 Authors Authors | Iglesias SM / Hou CFD / Li F / Cingolani G | |||||||||
| Funding support |  United States, 2 items United States, 2 items
| |||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2024 Journal: Nat Commun / Year: 2024Title: Integrative structural analysis of Pseudomonas phage DEV reveals a genome ejection motor. Authors: Ravi K Lokareddy / Chun-Feng David Hou / Francesca Forti / Stephano M Iglesias / Fenglin Li / Mikhail Pavlenok / David S Horner / Michael Niederweis / Federica Briani / Gino Cingolani /   Abstract: DEV is an obligatory lytic Pseudomonas phage of the N4-like genus, recently reclassified as Schitoviridae. The DEV genome encodes 91 ORFs, including a 3398 amino acid virion-associated RNA polymerase ...DEV is an obligatory lytic Pseudomonas phage of the N4-like genus, recently reclassified as Schitoviridae. The DEV genome encodes 91 ORFs, including a 3398 amino acid virion-associated RNA polymerase (vRNAP). Here, we describe the complete architecture of DEV, determined using a combination of cryo-electron microscopy localized reconstruction, biochemical methods, and genetic knockouts. We built de novo structures of all capsid factors and tail components involved in host attachment. We demonstrate that DEV long tail fibers are essential for infection of Pseudomonas aeruginosa but dispensable for infecting mutants with a truncated lipopolysaccharide devoid of the O-antigen. We determine that DEV vRNAP is part of a three-gene operon conserved in 191 Schitoviridae genomes. We propose these three proteins are ejected into the host to form a genome ejection motor spanning the cell envelope. We posit that the design principles of the DEV ejection apparatus are conserved in all Schitoviridae. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_44517.map.gz emd_44517.map.gz | 400.9 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-44517-v30.xml emd-44517-v30.xml emd-44517.xml emd-44517.xml | 16.6 KB 16.6 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_44517_fsc.xml emd_44517_fsc.xml | 18 KB | Display |  FSC data file FSC data file |
| Images |  emd_44517.png emd_44517.png | 33.4 KB | ||
| Masks |  emd_44517_msk_1.map emd_44517_msk_1.map | 512 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-44517.cif.gz emd-44517.cif.gz | 6 KB | ||
| Others |  emd_44517_half_map_1.map.gz emd_44517_half_map_1.map.gz emd_44517_half_map_2.map.gz emd_44517_half_map_2.map.gz | 405.7 MB 405.7 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-44517 http://ftp.pdbj.org/pub/emdb/structures/EMD-44517 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-44517 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-44517 | HTTPS FTP |
-Related structure data
| Related structure data |  9bgmMC  8vxqC  9bgnC  9bgoC  9codC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_44517.map.gz / Format: CCP4 / Size: 512 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_44517.map.gz / Format: CCP4 / Size: 512 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.12 Å | ||||||||||||||||||||||||||||||||||||
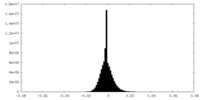
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_44517_msk_1.map emd_44517_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
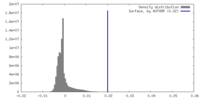
| Density Histograms |
-Half map: #1
| File | emd_44517_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
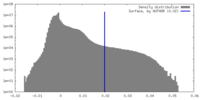
| Density Histograms |
-Half map: #2
| File | emd_44517_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
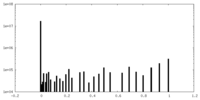
| Density Histograms |
- Sample components
Sample components
-Entire : Portal to tail complex of Pseudomonas phage DEV
| Entire | Name: Portal to tail complex of Pseudomonas phage DEV |
|---|---|
| Components |
|
-Supramolecule #1: Portal to tail complex of Pseudomonas phage DEV
| Supramolecule | Name: Portal to tail complex of Pseudomonas phage DEV / type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  Pseudomonas phage vB_PaeP_DEV (virus) Pseudomonas phage vB_PaeP_DEV (virus) |
-Macromolecule #1: gp75 tail tube
| Macromolecule | Name: gp75 tail tube / type: protein_or_peptide / ID: 1 / Number of copies: 12 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Pseudomonas phage vB_PaeP_DEV (virus) Pseudomonas phage vB_PaeP_DEV (virus) |
| Molecular weight | Theoretical: 35.267215 KDa |
| Sequence | String: MAVEPITIAD LTEVKLDGKG ALDQLLQVTR LHLAKEHDAG RLKGQEYAAV LTGGITAVLQ NAVMFLLQKD EAANKAALVE AQIKLTEKQ GELLDKQIAQ ADKDAELIAA KVKLTLEQAK LPDSQIRSAG FQDLLVQEQT KVQTAQTRRI DQEILSAGFQ D LLVKEQTA ...String: MAVEPITIAD LTEVKLDGKG ALDQLLQVTR LHLAKEHDAG RLKGQEYAAV LTGGITAVLQ NAVMFLLQKD EAANKAALVE AQIKLTEKQ GELLDKQIAQ ADKDAELIAA KVKLTLEQAK LPDSQIRSAG FQDLLVQEQT KVQTAQTRRI DQEILSAGFQ D LLVKEQTA KTKQDVLTAV QQTKVMEQQV LESTQKVLNM KQELLNLVAQ ECLLKAQFDL TKDQGLNTQE QTILVRQKVA SE RAQTIGA GVDADSVIGR QKELYKAQAD GFKRDAEQKA AKILIDTWNV RRTTDTGTQA NTTNRLDDAN VGRVVNMLMT GVG A UniProtKB: N4 gp54-like protein |
-Macromolecule #2: gp83 head-to-tail
| Macromolecule | Name: gp83 head-to-tail / type: protein_or_peptide / ID: 2 / Number of copies: 12 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Pseudomonas phage vB_PaeP_DEV (virus) Pseudomonas phage vB_PaeP_DEV (virus) |
| Molecular weight | Theoretical: 27.868934 KDa |
| Sequence | String: MTIQLKQVID LLAEGELSNI KYVNIDTGAL VLERVPSLIR AINLGVLDLH KRFLLKEGML KIQLEEGRRL YPLRPAYQVG QKPKPGVPQ FITEGNKLGR QSILKIEKII GDNGVEYYLN DTWQPLNITT PEFDVLEISD EFYCHSSSKT LEVRYRRAPT P MKICVDNL ...String: MTIQLKQVID LLAEGELSNI KYVNIDTGAL VLERVPSLIR AINLGVLDLH KRFLLKEGML KIQLEEGRRL YPLRPAYQVG QKPKPGVPQ FITEGNKLGR QSILKIEKII GDNGVEYYLN DTWQPLNITT PEFDVLEISD EFYCHSSSKT LEVRYRRAPT P MKICVDNL DSWGCIDIDL PYTHLQALLY FVASRCQTPI GFMENTAQEG FNFSQKYEAE CANLDAQNLR IDPVGNQDRF TR GGWV UniProtKB: Uncharacterized protein |
-Macromolecule #3: gp80 portal protein
| Macromolecule | Name: gp80 portal protein / type: protein_or_peptide / ID: 3 / Number of copies: 12 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Pseudomonas phage vB_PaeP_DEV (virus) Pseudomonas phage vB_PaeP_DEV (virus) |
| Molecular weight | Theoretical: 81.739453 KDa |
| Sequence | String: MADVDEDYLT LPNEDGDPSK RLQPEWSNAP SLAQLKQDYQ EAKQVTDEKI TQINRWLDYM HVRGEGKPKT EKGKSAVQPP TIRKQAEWR YSSLSEPFLS SPNIFEVNPV TWEDAESARQ NGLVLNQQFN TKLNKQRFID EYVRAGVDEG TIIVKVGWNY Q SRTVKEQV ...String: MADVDEDYLT LPNEDGDPSK RLQPEWSNAP SLAQLKQDYQ EAKQVTDEKI TQINRWLDYM HVRGEGKPKT EKGKSAVQPP TIRKQAEWR YSSLSEPFLS SPNIFEVNPV TWEDAESARQ NGLVLNQQFN TKLNKQRFID EYVRAGVDEG TIIVKVGWNY Q SRTVKEQV VTYEMMPDSS EELAQIYQTA AQIREESPSE YPEIPEDVRL GLEETEANGI QVRAVPVGSE EEEREETVEN HP TVQVCDY NNIVIDPSCG SDFSKAKFLI ETFESSYAEL KADGRYKNLD KIQVEGQNLL SEPDYTGPSE GVRNFDFQDK SRK RLVVHE YWGYYDIHGD GVLHPIVATW VGAVMIRMEE NPFPDKKIPY VVVSYIPRKR DLYGESDGAL LIDNQRIIGA VTRG MIDTM ARSANGQVGV MKGALDVTNR RRFDRGENYE FNPGADPRAA VHMHTFPEIP QSAQYMINLQ QAEAESMTGV KAFNA GISG AALGDTATAV RGALDAASKR ELGILRRLSA GIIEIGRKII AMNAEFLDDV EVVRITNEHF VDIRRDDLAG NFDLKL DIS TAEEDNAKVN DLTFMLQTMG PNMDPMMAQQ IMGQIMELKK MPDFAKRIRE FQPQPDPIAQ QKAQLELMLL QAQIEAE RA RAAHYMSGAG LQDSKVGTEQ AKARALASQA DMTDLNFLEQ ESGVQQARKR ELQQAQSEAQ GKLAMLNSQL KRLDEATS A RTSQK UniProtKB: Portal protein |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Average electron dose: 50.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 1.6 µm / Nominal defocus min: 0.8 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller







 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)