+ データを開く
データを開く
- 基本情報
基本情報
| 登録情報 |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| タイトル | Bovine adult muscle nAChR bound to ACh | |||||||||
 マップデータ マップデータ | adult_desen_sharp_map | |||||||||
 試料 試料 |
| |||||||||
 キーワード キーワード | Bovine muscle nicotinic acetylcholine receptor / MEMBRANE PROTEIN | |||||||||
| 機能・相同性 |  機能・相同性情報 機能・相同性情報Highly sodium permeable postsynaptic acetylcholine nicotinic receptors / Highly calcium permeable postsynaptic nicotinic acetylcholine receptors / Highly calcium permeable nicotinic acetylcholine receptors / postsynaptic membrane organization / skeletal muscle tissue growth / acetylcholine receptor activity / acetylcholine-gated channel complex / behavioral response to nicotine / neuromuscular synaptic transmission / acetylcholine-gated monoatomic cation-selective channel activity ...Highly sodium permeable postsynaptic acetylcholine nicotinic receptors / Highly calcium permeable postsynaptic nicotinic acetylcholine receptors / Highly calcium permeable nicotinic acetylcholine receptors / postsynaptic membrane organization / skeletal muscle tissue growth / acetylcholine receptor activity / acetylcholine-gated channel complex / behavioral response to nicotine / neuromuscular synaptic transmission / acetylcholine-gated monoatomic cation-selective channel activity / muscle cell development / acetylcholine binding / synaptic transmission, cholinergic / excitatory extracellular ligand-gated monoatomic ion channel activity / nervous system process / acetylcholine receptor signaling pathway / postsynaptic specialization membrane / membrane depolarization / monoatomic cation transport / skeletal muscle contraction / presynaptic modulation of chemical synaptic transmission / muscle contraction / response to nicotine / neuromuscular junction / transmitter-gated monoatomic ion channel activity involved in regulation of postsynaptic membrane potential / transmembrane signaling receptor activity / presynapse / channel activity / monoatomic ion transmembrane transport / chemical synaptic transmission / postsynaptic membrane / neuron projection / synapse / signal transduction / plasma membrane 類似検索 - 分子機能 | |||||||||
| 生物種 |  | |||||||||
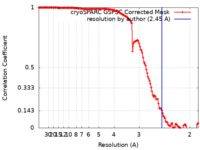
| 手法 | 単粒子再構成法 / クライオ電子顕微鏡法 / 解像度: 2.45 Å | |||||||||
 データ登録者 データ登録者 | Li H / Hibbs RE | |||||||||
| 資金援助 |  米国, 1件 米国, 1件
| |||||||||
 引用 引用 |  ジャーナル: Nature / 年: 2024 ジャーナル: Nature / 年: 2024タイトル: Structural switch in acetylcholine receptors in developing muscle. 著者: Huanhuan Li / Jinfeng Teng / Ryan E Hibbs /  要旨: During development, motor neurons originating in the brainstem and spinal cord form elaborate synapses with skeletal muscle fibres. These neurons release acetylcholine (ACh), which binds to nicotinic ...During development, motor neurons originating in the brainstem and spinal cord form elaborate synapses with skeletal muscle fibres. These neurons release acetylcholine (ACh), which binds to nicotinic ACh receptors (AChRs) on the muscle, initiating contraction. Two types of AChR are present in developing muscle cells, and their differential expression serves as a hallmark of neuromuscular synapse maturation. The structural principles underlying the switch from fetal to adult muscle receptors are unknown. Here, we present high-resolution structures of both fetal and adult muscle nicotinic AChRs, isolated from bovine skeletal muscle in developmental transition. These structures, obtained in the absence and presence of ACh, provide a structural context for understanding how fetal versus adult receptor isoforms are tuned for synapse development versus the all-or-none signalling required for high-fidelity skeletal muscle contraction. We find that ACh affinity differences are driven by binding site access, channel conductance is tuned by widespread surface electrostatics and open duration changes result from intrasubunit interactions and structural flexibility. The structures further reveal pathogenic mechanisms underlying congenital myasthenic syndromes. | |||||||||
| 履歴 |
|
- 構造の表示
構造の表示
| 添付画像 |
|---|
- ダウンロードとリンク
ダウンロードとリンク
-EMDBアーカイブ
| マップデータ |  emd_43925.map.gz emd_43925.map.gz | 226.9 MB |  EMDBマップデータ形式 EMDBマップデータ形式 | |
|---|---|---|---|---|
| ヘッダ (付随情報) |  emd-43925-v30.xml emd-43925-v30.xml emd-43925.xml emd-43925.xml | 28.1 KB 28.1 KB | 表示 表示 |  EMDBヘッダ EMDBヘッダ |
| FSC (解像度算出) |  emd_43925_fsc.xml emd_43925_fsc.xml | 13.1 KB | 表示 |  FSCデータファイル FSCデータファイル |
| 画像 |  emd_43925.png emd_43925.png | 45.7 KB | ||
| Filedesc metadata |  emd-43925.cif.gz emd-43925.cif.gz | 8 KB | ||
| その他 |  emd_43925_additional_1.map.gz emd_43925_additional_1.map.gz emd_43925_additional_2.map.gz emd_43925_additional_2.map.gz emd_43925_half_map_1.map.gz emd_43925_half_map_1.map.gz emd_43925_half_map_2.map.gz emd_43925_half_map_2.map.gz | 205.6 MB 226.9 MB 222.8 MB 222.8 MB | ||
| アーカイブディレクトリ |  http://ftp.pdbj.org/pub/emdb/structures/EMD-43925 http://ftp.pdbj.org/pub/emdb/structures/EMD-43925 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-43925 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-43925 | HTTPS FTP |
-関連構造データ
- リンク
リンク
| EMDBのページ |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| 「今月の分子」の関連する項目 |
- マップ
マップ
| ファイル |  ダウンロード / ファイル: emd_43925.map.gz / 形式: CCP4 / 大きさ: 512 MB / タイプ: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) ダウンロード / ファイル: emd_43925.map.gz / 形式: CCP4 / 大きさ: 512 MB / タイプ: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 注釈 | adult_desen_sharp_map | ||||||||||||||||||||||||||||||||||||
| 投影像・断面図 | 画像のコントロール
画像は Spider により作成 | ||||||||||||||||||||||||||||||||||||
| ボクセルのサイズ | X=Y=Z: 0.935 Å | ||||||||||||||||||||||||||||||||||||
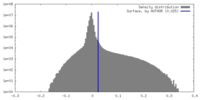
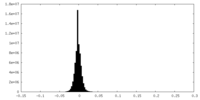
| 密度 |
| ||||||||||||||||||||||||||||||||||||
| 対称性 | 空間群: 1 | ||||||||||||||||||||||||||||||||||||
| 詳細 | EMDB XML:
|
-添付データ
-追加マップ: adult desen EMhancermap
| ファイル | emd_43925_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 注釈 | adult_desen_EMhancermap | ||||||||||||
| 投影像・断面図 |
| ||||||||||||
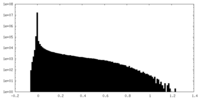
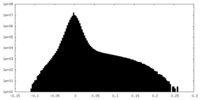
| 密度ヒストグラム |
-追加マップ: adult desen map
| ファイル | emd_43925_additional_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 注釈 | adult_desen_map | ||||||||||||
| 投影像・断面図 |
| ||||||||||||
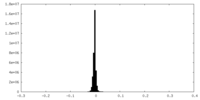
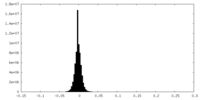
| 密度ヒストグラム |
-ハーフマップ: adult desen halfmapA
| ファイル | emd_43925_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 注釈 | adult_desen_halfmapA | ||||||||||||
| 投影像・断面図 |
| ||||||||||||
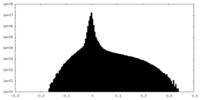
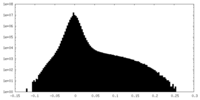
| 密度ヒストグラム |
-ハーフマップ: adult desen halfmapB
| ファイル | emd_43925_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 注釈 | adult_desen_halfmapB | ||||||||||||
| 投影像・断面図 |
| ||||||||||||
| 密度ヒストグラム |
- 試料の構成要素
試料の構成要素
-全体 : Bovine muscle nicotinic acetylcholine receptor
| 全体 | 名称: Bovine muscle nicotinic acetylcholine receptor |
|---|---|
| 要素 |
|
-超分子 #1: Bovine muscle nicotinic acetylcholine receptor
| 超分子 | 名称: Bovine muscle nicotinic acetylcholine receptor / タイプ: complex / ID: 1 / 親要素: 0 / 含まれる分子: #1-#4 |
|---|---|
| 由来(天然) | 生物種:  |
-分子 #1: Acetylcholine receptor subunit alpha
| 分子 | 名称: Acetylcholine receptor subunit alpha / タイプ: protein_or_peptide / ID: 1 / コピー数: 2 / 光学異性体: LEVO |
|---|---|
| 由来(天然) | 生物種:  |
| 分子量 | 理論値: 49.949828 KDa |
| 配列 | 文字列: SEHETRLVAK LFEDYNSVVR PVEDHRQAVE VTVGLQLIQL INVDEVNQIV TTNVRLKQQW VDYNLKWNPD DYGGVKKIHI PSEKIWRPD LVLYNNADGD FAIVKFTKVL LDYTGHITWT PPAIFKSYCE IIVTHFPFDE QNCSMKLGTW TYDGSVVVIN P ESDQPDLS ...文字列: SEHETRLVAK LFEDYNSVVR PVEDHRQAVE VTVGLQLIQL INVDEVNQIV TTNVRLKQQW VDYNLKWNPD DYGGVKKIHI PSEKIWRPD LVLYNNADGD FAIVKFTKVL LDYTGHITWT PPAIFKSYCE IIVTHFPFDE QNCSMKLGTW TYDGSVVVIN P ESDQPDLS NFMESGEWVI KESRGWKHWV FYACCPSTPY LDITYHFVMQ RLPLYFIVNV IIPCLLFSFL TGLVFYLPTD SG EKMTLSI SVLLSLTVFL LVIVELIPST SSAVPLIGKY MLFTMVFVIA SIIITVIVIN THHRSPSTHV MPEWVRKVFI DTI PNIMFF STMKRPSREK QDKKIFTEDI DISDISGKPG PPPMGFHSPL IKHPEVKSAI EGIKYIAETM KSDQESNNAA EEWK YVAMV MDHILLAVFM LVCIIGTLAV FAGRLIELNQ QG UniProtKB: Acetylcholine receptor subunit alpha |
-分子 #2: Acetylcholine receptor subunit beta
| 分子 | 名称: Acetylcholine receptor subunit beta / タイプ: protein_or_peptide / ID: 2 / コピー数: 1 / 光学異性体: LEVO |
|---|---|
| 由来(天然) | 生物種:  |
| 分子量 | 理論値: 55.111832 KDa |
| 配列 | 文字列: SEAEGRLREK LFSGYDSTVR PAREVGDRVW VSIGLTLAQL ISLNEKDEEM STKVYLDLEW TDYRLSWDPE EHEGIDSLRI SAESVWLPD VVLLNNNDGN FDVALDINVV VSSDGSMRWQ PPGIYRSSCS IQVTYFPFDW QNCTMVFSSY SYDSSEVSLQ T GLSPEGQE ...文字列: SEAEGRLREK LFSGYDSTVR PAREVGDRVW VSIGLTLAQL ISLNEKDEEM STKVYLDLEW TDYRLSWDPE EHEGIDSLRI SAESVWLPD VVLLNNNDGN FDVALDINVV VSSDGSMRWQ PPGIYRSSCS IQVTYFPFDW QNCTMVFSSY SYDSSEVSLQ T GLSPEGQE RQEVYIHEGT FIENGQWEII HKPSRLIQPS VDPRGGGEGR REEVTFYLII RRKPLFYLVN VIAPCILITL LA IFVFYLP PDAGEKMGLS IFALLTLTVF LLLLADKVPE TSLSVPIIIK YLMFTMVLVT FSVILSVVVL NLHHRSPHTH QMP LWVRQI FIHKLPLYLG LKRPKPERDQ MQEPPSIAPR DSPGSGWGRG TDEYFIRKPP NDFLFPKPNR FQPELSAPDL RRFI DGPNR AVGLPPELRE VVSSISYIAR QLQEQEDHDV LKEDWQFVAM VVDRLFLWTF IIFTSVGTLV IFLDATYHLP PADPF P UniProtKB: Acetylcholine receptor subunit beta |
-分子 #3: Acetylcholine receptor subunit delta
| 分子 | 名称: Acetylcholine receptor subunit delta / タイプ: protein_or_peptide / ID: 3 / コピー数: 1 / 光学異性体: LEVO |
|---|---|
| 由来(天然) | 生物種:  |
| 分子量 | 理論値: 56.543926 KDa |
| 配列 | 文字列: LNEEERLIRH LFEEKAYNKE LRPAAHKESV EISLALTLSN LISLKEVEET LTTNVWIEQG WTDSRLQWDA EDFGNISVLR LPADMVWLP EIVLENNNDG SFQISYSCNV LIYPSGSVYW LPPAIFRSSC PISVTYFPFD WQNCSLKFSS LKYTTKEITL S LKQAEEDG ...文字列: LNEEERLIRH LFEEKAYNKE LRPAAHKESV EISLALTLSN LISLKEVEET LTTNVWIEQG WTDSRLQWDA EDFGNISVLR LPADMVWLP EIVLENNNDG SFQISYSCNV LIYPSGSVYW LPPAIFRSSC PISVTYFPFD WQNCSLKFSS LKYTTKEITL S LKQAEEDG RSYPVEWIII DPEGFTENGE WEIVHRPARV NVDPSVPLDS PNRQDVTFYL IIRRKPLFYV INILVPCVLI SF MINLVFY LPADCGEKTS MAISVLLAQS VFLLLISKRL PATSMAIPLI GKFLLFGMVL VTMVVVICVI VLNIHFRTPS THV LSEPVK KLFLETLPEI LHMSRPAEDG PSPGTLIRRS SSLGYISKAE EYFSLKSRSD LMFEKQSERH GLARRLTTAR RPPA GSEQA QQELFSELKP AVDGANFIVN HMKDQNNYNE EKDCWNRVAR TVDRLCLFVV TPIMVVGTAW IFLQGAYNQP PPQPF PGDP FSYLEKDKRF I UniProtKB: Acetylcholine receptor subunit delta |
-分子 #4: Acetylcholine receptor subunit epsilon
| 分子 | 名称: Acetylcholine receptor subunit epsilon / タイプ: protein_or_peptide / ID: 4 / コピー数: 1 / 光学異性体: LEVO |
|---|---|
| 由来(天然) | 生物種:  |
| 分子量 | 理論値: 52.613262 KDa |
| 配列 | 文字列: KNEELRLYHY LFDTYDPGRR PVQEPEDTVT ISLKVTLTNL ISLNEKEETL TTSVWIGIDW QDYRLNYSKG DFGGVETLRV PSELVWLPE IVLENNIDGQ FGVAYEANVL VSEGGYLSWL PPAIYRSTCA VEVTYFPFDW QNCSLVFRSQ TYNAEEVEFV F AVDDEGKT ...文字列: KNEELRLYHY LFDTYDPGRR PVQEPEDTVT ISLKVTLTNL ISLNEKEETL TTSVWIGIDW QDYRLNYSKG DFGGVETLRV PSELVWLPE IVLENNIDGQ FGVAYEANVL VSEGGYLSWL PPAIYRSTCA VEVTYFPFDW QNCSLVFRSQ TYNAEEVEFV F AVDDEGKT ISKIDIDTEA YTENGEWAID FCPGVIRRHD GDSAGGPGET DVIYSLIIRR KPLFYVINII VPCVLISGLV LL AYFLPAQ AGGQKCTVSI NVLLAQTVFL FLIAQKTPET SLSVPLLGRY LIFVMVVATL IVMNCVIVLN VSLRTPTTHA MSP RLRYVL LELLPQLLGS GAPPEIPRAA SPPRRASSLG LLLRAEELIL KKPRSELVFE QQRHRHGTWT ATLCQNLGAA APEI RCCVD AVNFVASSTR DQEATGEEVS DWVRMGKALD SICFWAALVL FLVGSSLIFL GAYFNRVPQL PYPPCM UniProtKB: Acetylcholine receptor subunit epsilon |
-分子 #11: ACETYLCHOLINE
| 分子 | 名称: ACETYLCHOLINE / タイプ: ligand / ID: 11 / コピー数: 2 / 式: ACH |
|---|---|
| 分子量 | 理論値: 146.207 Da |
| Chemical component information |  ChemComp-ACH: |
-分子 #12: (2S)-3-(hexadecanoyloxy)-2-[(9Z)-octadec-9-enoyloxy]propyl 2-(tri...
| 分子 | 名称: (2S)-3-(hexadecanoyloxy)-2-[(9Z)-octadec-9-enoyloxy]propyl 2-(trimethylammonio)ethyl phosphate タイプ: ligand / ID: 12 / コピー数: 2 / 式: POV |
|---|---|
| 分子量 | 理論値: 760.076 Da |
| Chemical component information |  ChemComp-POV: |
-分子 #13: water
| 分子 | 名称: water / タイプ: ligand / ID: 13 / コピー数: 5 / 式: HOH |
|---|---|
| 分子量 | 理論値: 18.015 Da |
| Chemical component information |  ChemComp-HOH: |
-実験情報
-構造解析
| 手法 | クライオ電子顕微鏡法 |
|---|---|
 解析 解析 | 単粒子再構成法 |
| 試料の集合状態 | particle |
- 試料調製
試料調製
| 緩衝液 | pH: 7.4 |
|---|---|
| 凍結 | 凍結剤: ETHANE |
- 電子顕微鏡法
電子顕微鏡法
| 顕微鏡 | FEI TITAN KRIOS |
|---|---|
| 撮影 | フィルム・検出器のモデル: FEI FALCON III (4k x 4k) 平均電子線量: 50.0 e/Å2 |
| 電子線 | 加速電圧: 300 kV / 電子線源:  FIELD EMISSION GUN FIELD EMISSION GUN |
| 電子光学系 | 照射モード: OTHER / 撮影モード: BRIGHT FIELD / 最大 デフォーカス(公称値): 2.0 µm / 最小 デフォーカス(公称値): 1.6 µm |
| 実験機器 |  モデル: Titan Krios / 画像提供: FEI Company |
+ 画像解析
画像解析
-原子モデル構築 1
| 初期モデル | Chain - Source name: Other / Chain - Initial model type: experimental model / 詳細: cryosparc ab initial model |
|---|---|
| 精密化 | 空間: REAL / プロトコル: AB INITIO MODEL |
| 得られたモデル |  PDB-9awj: |
 ムービー
ムービー コントローラー
コントローラー











 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)