[English] 日本語
 Yorodumi
Yorodumi- EMDB-4315: Subtomogram average of OST-containing ribosome-translocon complex... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-4315 | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Subtomogram average of OST-containing ribosome-translocon complexes from canine rough microsomal membranes | ||||||||||||
 Map data Map data | |||||||||||||
 Sample Sample |
| ||||||||||||
 Keywords Keywords | Protein translocon of the endoplasmic reticulum / PROTEIN TRANSPORT | ||||||||||||
| Function / homology |  Function and homology information Function and homology informationRegulation of CDH1 posttranslational processing and trafficking to plasma membrane / oligosaccharyltransferase complex A / oligosaccharyltransferase complex B / Insertion of tail-anchored proteins into the endoplasmic reticulum membrane / membrane docking / endoplasmic reticulum Sec complex / pronephric nephron development / dolichyl-diphosphooligosaccharide-protein glycotransferase / dolichyl-diphosphooligosaccharide-protein glycotransferase activity / cotranslational protein targeting to membrane ...Regulation of CDH1 posttranslational processing and trafficking to plasma membrane / oligosaccharyltransferase complex A / oligosaccharyltransferase complex B / Insertion of tail-anchored proteins into the endoplasmic reticulum membrane / membrane docking / endoplasmic reticulum Sec complex / pronephric nephron development / dolichyl-diphosphooligosaccharide-protein glycotransferase / dolichyl-diphosphooligosaccharide-protein glycotransferase activity / cotranslational protein targeting to membrane / Ssh1 translocon complex / Sec61 translocon complex / protein insertion into ER membrane / protein-transporting ATPase activity / oligosaccharyltransferase complex / protein targeting to ER / : / SRP-dependent cotranslational protein targeting to membrane, translocation / glycoprotein biosynthetic process / signal sequence binding / post-translational protein targeting to membrane, translocation / transmembrane protein transporter activity / ubiquitin ligase inhibitor activity / positive regulation of signal transduction by p53 class mediator / rough endoplasmic reticulum / MDM2/MDM4 family protein binding / post-translational protein modification / guanyl-nucleotide exchange factor activity / ribosomal large subunit biogenesis / maturation of LSU-rRNA from tricistronic rRNA transcript (SSU-rRNA, 5.8S rRNA, LSU-rRNA) / phospholipid binding / large ribosomal subunit / ribosome biogenesis / ribosome binding / 5S rRNA binding / ribosomal large subunit assembly / large ribosomal subunit rRNA binding / cytosolic large ribosomal subunit / cytoplasmic translation / tRNA binding / rRNA binding / structural constituent of ribosome / ribosome / translation / ribonucleoprotein complex / mRNA binding / synapse / endoplasmic reticulum membrane / nucleolus / endoplasmic reticulum / RNA binding / zinc ion binding / membrane / metal ion binding / nucleus / cytoplasm / cytosol Similarity search - Function | ||||||||||||
| Biological species |   | ||||||||||||
| Method | subtomogram averaging / cryo EM / Resolution: 9.1 Å | ||||||||||||
 Authors Authors | Pfeffer S / Foerster F | ||||||||||||
| Funding support |  Germany, 3 items Germany, 3 items
| ||||||||||||
 Citation Citation |  Journal: Science / Year: 2018 Journal: Science / Year: 2018Title: Structural basis for coupling protein transport and N-glycosylation at the mammalian endoplasmic reticulum. Authors: Katharina Braunger / Stefan Pfeffer / Shiteshu Shrimal / Reid Gilmore / Otto Berninghausen / Elisabet C Mandon / Thomas Becker / Friedrich Förster / Roland Beckmann /    Abstract: Protein synthesis, transport, and N-glycosylation are coupled at the mammalian endoplasmic reticulum by complex formation of a ribosome, the Sec61 protein-conducting channel, and ...Protein synthesis, transport, and N-glycosylation are coupled at the mammalian endoplasmic reticulum by complex formation of a ribosome, the Sec61 protein-conducting channel, and oligosaccharyltransferase (OST). Here we used different cryo-electron microscopy approaches to determine structures of native and solubilized ribosome-Sec61-OST complexes. A molecular model for the catalytic OST subunit STT3A (staurosporine and temperature sensitive 3A) revealed how it is integrated into the OST and how STT3-paralog specificity for translocon-associated OST is achieved. The OST subunit DC2 was placed at the interface between Sec61 and STT3A, where it acts as a versatile module for recruitment of STT3A-containing OST to the ribosome-Sec61 complex. This detailed structural view on the molecular architecture of the cotranslational machinery for N-glycosylation provides the basis for a mechanistic understanding of glycoprotein biogenesis at the endoplasmic reticulum. | ||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_4315.map.gz emd_4315.map.gz | 13.1 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-4315-v30.xml emd-4315-v30.xml emd-4315.xml emd-4315.xml | 101.8 KB 101.8 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_4315.png emd_4315.png | 97.5 KB | ||
| Filedesc metadata |  emd-4315.cif.gz emd-4315.cif.gz | 17.7 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-4315 http://ftp.pdbj.org/pub/emdb/structures/EMD-4315 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-4315 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-4315 | HTTPS FTP |
-Related structure data
| Related structure data |  6ftgMC  4306C  4307C  4308C  4309C  4310C  4311C  4312C  4313C 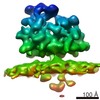 4314C  4316C  4317C  6ftiC  6ftjC C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_4315.map.gz / Format: CCP4 / Size: 45.2 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_4315.map.gz / Format: CCP4 / Size: 45.2 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 2.62 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
+Entire : 80S ribosome from WT canine rough microsomal membranes bound to t...
+Supramolecule #1: 80S ribosome from WT canine rough microsomal membranes bound to t...
+Supramolecule #2: Sec61 protein conducting channel
+Supramolecule #3: 60S ribosomal subunit
+Supramolecule #4: Mammalian Oligosaccharyltransferase
+Macromolecule #1: uL2
+Macromolecule #2: uL3
+Macromolecule #3: Ribosomal protein L4
+Macromolecule #4: 60S ribosomal protein L5
+Macromolecule #5: 60S ribosomal protein L6
+Macromolecule #6: uL30
+Macromolecule #7: eL8
+Macromolecule #8: uL6
+Macromolecule #9: Ribosomal protein L10 (Predicted)
+Macromolecule #10: Ribosomal protein L11
+Macromolecule #11: eL13
+Macromolecule #12: Ribosomal protein L14
+Macromolecule #13: Ribosomal protein L15
+Macromolecule #14: uL13
+Macromolecule #15: uL22
+Macromolecule #16: uL14
+Macromolecule #17: eL19
+Macromolecule #18: eL20
+Macromolecule #19: eL21
+Macromolecule #20: eL22
+Macromolecule #21: uL14
+Macromolecule #22: Ribosomal protein L24
+Macromolecule #23: uL23
+Macromolecule #24: Ribosomal protein L26
+Macromolecule #25: 60S ribosomal protein L27
+Macromolecule #26: uL15
+Macromolecule #27: 60S ribosomal protein L29
+Macromolecule #28: eL30
+Macromolecule #29: eL31
+Macromolecule #30: eL32
+Macromolecule #31: eL33
+Macromolecule #32: eL34
+Macromolecule #33: eL29
+Macromolecule #34: 60S ribosomal protein L36
+Macromolecule #35: Ribosomal protein L37
+Macromolecule #36: eL38
+Macromolecule #37: eL39
+Macromolecule #38: eL40
+Macromolecule #39: 60s ribosomal protein l41
+Macromolecule #40: eL42
+Macromolecule #41: Ribosomal protein L37a
+Macromolecule #42: eL28
+Macromolecule #43: 60S acidic ribosomal protein P0
+Macromolecule #44: Ribosomal protein L12
+Macromolecule #48: Protein transport protein Sec61 subunit alpha isoform 1
+Macromolecule #49: Protein transport protein Sec61 subunit gamma
+Macromolecule #50: Protein transport protein Sec61 subunit beta
+Macromolecule #51: Dolichyl-diphosphooligosaccharide--protein glycosyltransferase su...
+Macromolecule #52: TMEM258
+Macromolecule #53: Oligosaccharyltransferase complex subunit OSTC
+Macromolecule #54: OST4
+Macromolecule #55: Dolichyl-diphosphooligosaccharide--protein glycosyltransferase su...
+Macromolecule #56: DAD1
+Macromolecule #57: OST48
+Macromolecule #58: RPN2
+Macromolecule #45: 28S ribosomal RNA
+Macromolecule #46: 5S ribosomal RNA
+Macromolecule #47: 5.8S ribosomal RNA
+Macromolecule #60: MAGNESIUM ION
+Macromolecule #61: ZINC ION
+Macromolecule #62: [(2~{S},3~{R},4~{R},5~{S},6~{R})-3-acetamido-6-(hydroxymethyl)-4,...
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | subtomogram averaging |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 2.0 mg/mL |
|---|---|
| Buffer | pH: 7.6 |
| Grid | Material: MOLYBDENUM / Support film - Material: CARBON / Support film - topology: LACEY |
| Vitrification | Cryogen name: ETHANE-PROPANE / Chamber humidity: 70 % / Instrument: FEI VITROBOT MARK IV / Details: Blot 3 seconds before plunging.. |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Detector mode: COUNTING / Average electron dose: 1.5 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 4.0 µm / Nominal defocus min: 3.0 µm |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Final reconstruction | Applied symmetry - Point group: C1 (asymmetric) / Algorithm: BACK PROJECTION / Resolution.type: BY AUTHOR / Resolution: 9.1 Å / Resolution method: FSC 0.5 CUT-OFF / Number subtomograms used: 17600 |
|---|---|
| Extraction | Number tomograms: 211 / Number images used: 27500 / Method: Template matching / Software: (Name: PyTom, TOM Toolbox, AV3) |
| CTF correction | Software - Name: PyTom / Details: On each individual tilt image / Type: PHASE FLIPPING ONLY |
| Final angle assignment | Type: NOT APPLICABLE |
-Atomic model buiding 1
| Refinement | Space: REAL / Protocol: RIGID BODY FIT / Overall B value: 500 / Target criteria: Cross-correlation coefficient |
|---|---|
| Output model |  PDB-6ftg: |
 Movie
Movie Controller
Controller


















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)






















