+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-3877 | |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Inner membrane Bcs complex (E. coli) | |||||||||||||||
 Map data Map data | An inner membrane subcomplex of the E. coli cellulose secretion system (Bcs). The complex was purified from solubilised membranes of Bl21(DE3) cells over-expressing the BcsRQAB subunits. | |||||||||||||||
 Sample Sample |
| |||||||||||||||
| Biological species |  | |||||||||||||||
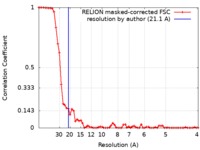
| Method | single particle reconstruction / negative staining / Resolution: 21.1 Å | |||||||||||||||
 Authors Authors | Krasteva PV / Fronzes R | |||||||||||||||
| Funding support |  France, 4 items France, 4 items
| |||||||||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2017 Journal: Nat Commun / Year: 2017Title: Insights into the structure and assembly of a bacterial cellulose secretion system. Authors: Petya Violinova Krasteva / Joaquin Bernal-Bayard / Laetitia Travier / Fernando Ariel Martin / Pierre-Alexandre Kaminski / Gouzel Karimova / Rémi Fronzes / Jean-Marc Ghigo /   Abstract: Secreted exopolysaccharides present important determinants for bacterial biofilm formation, survival, and virulence. Cellulose secretion typically requires the concerted action of a c-di-GMP- ...Secreted exopolysaccharides present important determinants for bacterial biofilm formation, survival, and virulence. Cellulose secretion typically requires the concerted action of a c-di-GMP-responsive inner membrane synthase (BcsA), an accessory membrane-anchored protein (BcsB), and several additional Bcs components. Although the BcsAB catalytic duo has been studied in great detail, its interplay with co-expressed subunits remains enigmatic. Here we show that E. coli Bcs proteins partake in a complex protein interaction network. Electron microscopy reveals a stable, megadalton-sized macromolecular assembly, which encompasses most of the inner membrane and cytosolic Bcs components and features a previously unobserved asymmetric architecture. Heterologous reconstitution and mutational analyses point toward a structure-function model, where accessory proteins regulate secretion by affecting both the assembly and stability of the system. Altogether, these results lay the foundation for more comprehensive models of synthase-dependent exopolysaccharide secretion in biofilms and add a sophisticated secretory nanomachine to the diverse bacterial arsenal for virulence and adaptation. | |||||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_3877.map.gz emd_3877.map.gz | 8.9 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-3877-v30.xml emd-3877-v30.xml emd-3877.xml emd-3877.xml | 13.3 KB 13.3 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_3877_fsc.xml emd_3877_fsc.xml | 6.3 KB | Display |  FSC data file FSC data file |
| Images |  emd_3877.png emd_3877.png | 31.2 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-3877 http://ftp.pdbj.org/pub/emdb/structures/EMD-3877 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-3877 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-3877 | HTTPS FTP |
-Validation report
| Summary document |  emd_3877_validation.pdf.gz emd_3877_validation.pdf.gz | 232.3 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_3877_full_validation.pdf.gz emd_3877_full_validation.pdf.gz | 231.4 KB | Display | |
| Data in XML |  emd_3877_validation.xml.gz emd_3877_validation.xml.gz | 9 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-3877 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-3877 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-3877 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-3877 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_3877.map.gz / Format: CCP4 / Size: 22.2 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_3877.map.gz / Format: CCP4 / Size: 22.2 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | An inner membrane subcomplex of the E. coli cellulose secretion system (Bcs). The complex was purified from solubilised membranes of Bl21(DE3) cells over-expressing the BcsRQAB subunits. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.9 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
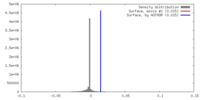
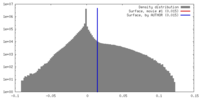
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : Bcs inner membrane sub-complex.
| Entire | Name: Bcs inner membrane sub-complex. |
|---|---|
| Components |
|
-Supramolecule #1: Bcs inner membrane sub-complex.
| Supramolecule | Name: Bcs inner membrane sub-complex. / type: complex / ID: 1 / Parent: 0 Details: Sample purified using BcsA as bait from solubilised E. coli Bl21(DE3) cells over-expressing the BcsRQAB sub-units of cellulose-secreting E. coli 1094. |
|---|---|
| Source (natural) | Organism:  |
| Recombinant expression | Organism:  |
| Molecular weight | Theoretical: 1 MDa |
-Experimental details
-Structure determination
| Method | negative staining |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.1 mg/mL |
|---|---|
| Buffer | pH: 8 Details: 20 mM HEPES 8.0 120 mM NaCl 10 glycerol 5 mM MgCl2 10 uM AppCp 2 uM c-di-GMP 0.008% LM-NPG cOmplete protease inhibitors |
| Staining | Type: NEGATIVE / Material: uranyl acetate Details: 5 ul of sample were spotted on glow-discharged carbon-coated copper grids and incubated for 1 minute. Liquid was blotted off and the sample was passed through 3 drops of 10 ul 2% uranyl ...Details: 5 ul of sample were spotted on glow-discharged carbon-coated copper grids and incubated for 1 minute. Liquid was blotted off and the sample was passed through 3 drops of 10 ul 2% uranyl acetate with a 30-second incubation over the third. After that the stain was blotted off and the grid was air-dried. |
| Grid | Material: COPPER / Support film - Material: CARBON / Support film - topology: CONTINUOUS / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Atmosphere: OTHER |
| Details | Samples were purified using anti-FLAG affinity gel from detergent-solubilised membranes of E. coli(DE3) cells over-expressing the BcsRQAB subunits of an E. coli 1094-derived strain (pCDF vector, IPTG-inducible overnight expression in TB, affinity tag on BcsA). Eluted samples were further purified by centrifugation over a 10-40% glycerol density gradient. |
- Electron microscopy
Electron microscopy
| Microscope | FEI TECNAI F20 |
|---|---|
| Image recording | Film or detector model: FEI FALCON II (4k x 4k) / Number grids imaged: 2 / Average electron dose: 12.0 e/Å2 |
| Electron beam | Acceleration voltage: 200 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Calibrated defocus max: 3.5 µm / Illumination mode: OTHER / Imaging mode: BRIGHT FIELD / Nominal defocus max: 0.5 µm / Nominal magnification: 50000 |
| Sample stage | Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Tecnai F20 / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller











 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)