[English] 日本語
 Yorodumi
Yorodumi- EMDB-38357: Human Cx36/GJD2 (Ala14 deletion mutant) gap junction channel prep... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Human Cx36/GJD2 (Ala14 deletion mutant) gap junction channel prepared with mefloquine, showing no bound mefloquine (C1 symmetry) | |||||||||
 Map data Map data | Cryo-EM density map, C1 symmetry | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | connexin 36 / Cx36 / Gap Junction Channel / MEMBRANE PROTEIN | |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.1 Å | |||||||||
 Authors Authors | Cho HJ / Lee HH | |||||||||
| Funding support |  Korea, Republic Of, 1 items Korea, Republic Of, 1 items
| |||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2024 Journal: Nat Commun / Year: 2024Title: Mefloquine-induced conformational shift in Cx36 N-terminal helix leading to channel closure mediated by lipid bilayer. Authors: Hwa-Jin Cho / Dong Kyu Chung / Hyung Ho Lee /  Abstract: Connexin 36 (Cx36) forms interneuronal gap junctions, establishing electrical synapses for rapid synaptic transmission. In disease conditions, inhibiting Cx36 gap junction channels (GJCs) is ...Connexin 36 (Cx36) forms interneuronal gap junctions, establishing electrical synapses for rapid synaptic transmission. In disease conditions, inhibiting Cx36 gap junction channels (GJCs) is beneficial, as it prevents abnormal synchronous neuronal firing and apoptotic signal propagation, mitigating seizures and progressive cell death. Here, we present cryo-electron microscopy structures of human Cx36 GJC in complex with known channel inhibitors, such as mefloquine, arachidonic acid, and 1-hexanol. Notably, these inhibitors competitively bind to the binding pocket of the N-terminal helices (NTH), inducing a conformational shift from the pore-lining NTH (PLN) state to the flexible NTH (FN) state. This leads to the obstruction of the channel pore by flat double-layer densities of lipids. These studies elucidate the molecular mechanisms of how Cx36 GJC can be modulated by inhibitors, providing valuable insights into potential therapeutic applications. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_38357.map.gz emd_38357.map.gz | 168 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-38357-v30.xml emd-38357-v30.xml emd-38357.xml emd-38357.xml | 12.2 KB 12.2 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_38357_fsc.xml emd_38357_fsc.xml | 12 KB | Display |  FSC data file FSC data file |
| Images |  emd_38357.png emd_38357.png | 108.8 KB | ||
| Filedesc metadata |  emd-38357.cif.gz emd-38357.cif.gz | 3.8 KB | ||
| Others |  emd_38357_half_map_1.map.gz emd_38357_half_map_1.map.gz emd_38357_half_map_2.map.gz emd_38357_half_map_2.map.gz | 165.1 MB 165.1 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-38357 http://ftp.pdbj.org/pub/emdb/structures/EMD-38357 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-38357 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-38357 | HTTPS FTP |
-Validation report
| Summary document |  emd_38357_validation.pdf.gz emd_38357_validation.pdf.gz | 1.2 MB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_38357_full_validation.pdf.gz emd_38357_full_validation.pdf.gz | 1.2 MB | Display | |
| Data in XML |  emd_38357_validation.xml.gz emd_38357_validation.xml.gz | 20.7 KB | Display | |
| Data in CIF |  emd_38357_validation.cif.gz emd_38357_validation.cif.gz | 26.7 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-38357 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-38357 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-38357 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-38357 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_38357.map.gz / Format: CCP4 / Size: 178 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_38357.map.gz / Format: CCP4 / Size: 178 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Cryo-EM density map, C1 symmetry | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.6 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: Half map A
| File | emd_38357_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half map A | ||||||||||||
| Projections & Slices |
| ||||||||||||
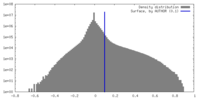
| Density Histograms |
-Half map: Half map B
| File | emd_38357_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half map B | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Human Cx36/GJD2 (Ala14 deletion mutant) gap junction channel in b...
| Entire | Name: Human Cx36/GJD2 (Ala14 deletion mutant) gap junction channel in brain lipids |
|---|---|
| Components |
|
-Supramolecule #1: Human Cx36/GJD2 (Ala14 deletion mutant) gap junction channel in b...
| Supramolecule | Name: Human Cx36/GJD2 (Ala14 deletion mutant) gap junction channel in brain lipids type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: FEI FALCON IV (4k x 4k) / Average electron dose: 50.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 1.6 µm / Nominal defocus min: 0.8 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller























 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)