[English] 日本語
 Yorodumi
Yorodumi- EMDB-3680: Full-length complex of S.cerevisiae Cdt1-MCM in ATPgS-bound state. -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-3680 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Full-length complex of S.cerevisiae Cdt1-MCM in ATPgS-bound state. | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
| Biological species |  | |||||||||
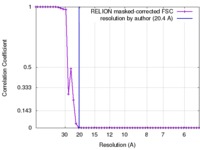
| Method | single particle reconstruction / negative staining / Resolution: 20.4 Å | |||||||||
 Authors Authors | He J / Frigola J / Kinkelin K / Pye VE / Renault L / Douglas M / Remus D / Cherepanov P / Costa A / Diffley JFX | |||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2017 Journal: Nat Commun / Year: 2017Title: Cdt1 stabilizes an open MCM ring for helicase loading. Authors: Jordi Frigola / Jun He / Kerstin Kinkelin / Valerie E Pye / Ludovic Renault / Max E Douglas / Dirk Remus / Peter Cherepanov / Alessandro Costa / John F X Diffley /   Abstract: ORC, Cdc6 and Cdt1 act together to load hexameric MCM, the motor of the eukaryotic replicative helicase, into double hexamers at replication origins. Here we show that Cdt1 interacts with MCM ...ORC, Cdc6 and Cdt1 act together to load hexameric MCM, the motor of the eukaryotic replicative helicase, into double hexamers at replication origins. Here we show that Cdt1 interacts with MCM subunits Mcm2, 4 and 6, which both destabilizes the Mcm2-5 interface and inhibits MCM ATPase activity. Using X-ray crystallography, we show that Cdt1 contains two winged-helix domains in the C-terminal half of the protein and a catalytically inactive dioxygenase-related N-terminal domain, which is important for MCM loading, but not for subsequent replication. We used these structures together with single-particle electron microscopy to generate three-dimensional models of MCM complexes. These show that Cdt1 stabilizes MCM in a left-handed spiral open at the Mcm2-5 gate. We propose that Cdt1 acts as a brace, holding MCM open for DNA entry and bound to ATP until ORC-Cdc6 triggers ATP hydrolysis by MCM, promoting both Cdt1 ejection and MCM ring closure. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_3680.map.gz emd_3680.map.gz | 7.3 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-3680-v30.xml emd-3680-v30.xml emd-3680.xml emd-3680.xml | 7.6 KB 7.6 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_3680_fsc.xml emd_3680_fsc.xml | 4.3 KB | Display |  FSC data file FSC data file |
| Images |  emd_3680.png emd_3680.png | 441.1 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-3680 http://ftp.pdbj.org/pub/emdb/structures/EMD-3680 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-3680 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-3680 | HTTPS FTP |
-Validation report
| Summary document |  emd_3680_validation.pdf.gz emd_3680_validation.pdf.gz | 235.4 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_3680_full_validation.pdf.gz emd_3680_full_validation.pdf.gz | 234.5 KB | Display | |
| Data in XML |  emd_3680_validation.xml.gz emd_3680_validation.xml.gz | 7.8 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-3680 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-3680 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-3680 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-3680 | HTTPS FTP |
-Related structure data
| Related structure data |  3679C 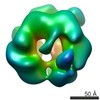 3681C  5me9C  5meaC  5mebC  5mecC C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_3680.map.gz / Format: CCP4 / Size: 8 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_3680.map.gz / Format: CCP4 / Size: 8 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 2.7 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : Full-length yeast Cdt1-MCM complex in ATPgS-bound state.
| Entire | Name: Full-length yeast Cdt1-MCM complex in ATPgS-bound state. |
|---|---|
| Components |
|
-Supramolecule #1: Full-length yeast Cdt1-MCM complex in ATPgS-bound state.
| Supramolecule | Name: Full-length yeast Cdt1-MCM complex in ATPgS-bound state. type: complex / ID: 1 / Parent: 0 |
|---|---|
| Source (natural) | Organism:  |
| Recombinant expression | Organism:  |
-Experimental details
-Structure determination
| Method | negative staining |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.6 |
|---|---|
| Staining | Type: NEGATIVE / Material: Uranyl formate |
- Electron microscopy
Electron microscopy
| Microscope | JEOL 2100 |
|---|---|
| Image recording | Film or detector model: GATAN ULTRASCAN 4000 (4k x 4k) / Average electron dose: 20.0 e/Å2 |
| Electron beam | Acceleration voltage: 120 kV / Electron source: LAB6 |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
 Movie
Movie Controller
Controller


 UCSF Chimera
UCSF Chimera







 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)