[English] 日本語
 Yorodumi
Yorodumi- EMDB-36107: Cryo-EM composite map of Euglena gracilis respiratory complex I, ... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM composite map of Euglena gracilis respiratory complex I, deactive state | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Electron transport chain / complex / membrane protein / Euglena gracilis / ELECTRON TRANSPORT | |||||||||
| Function / homology |  Function and homology information Function and homology informationrespiratory chain complex I / NADH:ubiquinone reductase (H+-translocating) / ubiquinone binding / NADH dehydrogenase (ubiquinone) activity / electron transport coupled proton transport / ATP synthesis coupled electron transport / mitochondrial membrane Similarity search - Function | |||||||||
| Biological species |  Euglena gracilis (euglena) Euglena gracilis (euglena) | |||||||||
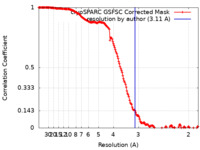
| Method | single particle reconstruction / cryo EM / Resolution: 3.11 Å | |||||||||
 Authors Authors | Wu MC / He ZX / Tian HT / Hu YQ / Han FZ / Zhou L | |||||||||
| Funding support |  China, 1 items China, 1 items
| |||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2024 Journal: Nat Commun / Year: 2024Title: Euglena's atypical respiratory chain adapts to the discoidal cristae and flexible metabolism. Authors: Zhaoxiang He / Mengchen Wu / Hongtao Tian / Liangdong Wang / Yiqi Hu / Fangzhu Han / Jiancang Zhou / Yong Wang / Long Zhou /  Abstract: Euglena gracilis, a model organism of the eukaryotic supergroup Discoba harbouring also clinically important parasitic species, possesses diverse metabolic strategies and an atypical electron ...Euglena gracilis, a model organism of the eukaryotic supergroup Discoba harbouring also clinically important parasitic species, possesses diverse metabolic strategies and an atypical electron transport chain. While structures of the electron transport chain complexes and supercomplexes of most other eukaryotic clades have been reported, no similar structure is currently available for Discoba, limiting the understandings of its core metabolism and leaving a gap in the evolutionary tree of eukaryotic bioenergetics. Here, we report high-resolution cryo-EM structures of Euglena's respirasome I + III + IV and supercomplex III + IV. A previously unreported fatty acid synthesis domain locates on the tip of complex I's peripheral arm, providing a clear picture of its atypical subunit composition identified previously. Individual complexes are re-arranged in the respirasome to adapt to the non-uniform membrane curvature of the discoidal cristae. Furthermore, Euglena's conformationally rigid complex I is deactivated by restricting ubiquinone's access to its substrate tunnel. Our findings provide structural insights for therapeutic developments against euglenozoan parasite infections. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_36107.map.gz emd_36107.map.gz | 372 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-36107-v30.xml emd-36107-v30.xml emd-36107.xml emd-36107.xml | 81.9 KB 81.9 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_36107_fsc.xml emd_36107_fsc.xml | 15.9 KB | Display |  FSC data file FSC data file |
| Images |  emd_36107.png emd_36107.png | 104.5 KB | ||
| Filedesc metadata |  emd-36107.cif.gz emd-36107.cif.gz | 18.6 KB | ||
| Others |  emd_36107_half_map_1.map.gz emd_36107_half_map_1.map.gz emd_36107_half_map_2.map.gz emd_36107_half_map_2.map.gz | 390.9 MB 390.9 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-36107 http://ftp.pdbj.org/pub/emdb/structures/EMD-36107 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-36107 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-36107 | HTTPS FTP |
-Validation report
| Summary document |  emd_36107_validation.pdf.gz emd_36107_validation.pdf.gz | 1.1 MB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_36107_full_validation.pdf.gz emd_36107_full_validation.pdf.gz | 1.1 MB | Display | |
| Data in XML |  emd_36107_validation.xml.gz emd_36107_validation.xml.gz | 25.5 KB | Display | |
| Data in CIF |  emd_36107_validation.cif.gz emd_36107_validation.cif.gz | 33.1 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-36107 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-36107 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-36107 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-36107 | HTTPS FTP |
-Related structure data
| Related structure data |  8j9hMC  8iufC  8iujC  8j9iC  8j9jC C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_36107.map.gz / Format: CCP4 / Size: 421.9 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_36107.map.gz / Format: CCP4 / Size: 421.9 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Voxel size | X=Y=Z: 0.93 Å | ||||||||||||||||||||
| Density |
| ||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: #2
| File | emd_36107_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #1
| File | emd_36107_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
+Entire : Euglena gracilis respiratory complex I, deactive state
+Supramolecule #1: Euglena gracilis respiratory complex I, deactive state
+Macromolecule #1: NDUS1A
+Macromolecule #2: NDUS1B
+Macromolecule #3: NADH dehydrogenase subunit 5
+Macromolecule #4: ND4L
+Macromolecule #5: NDUFA1
+Macromolecule #6: NDUFA2
+Macromolecule #7: NDUFA3
+Macromolecule #8: NDUFA5
+Macromolecule #9: NDUFA6
+Macromolecule #10: NDUFA7
+Macromolecule #11: NDUFA8
+Macromolecule #12: NDUFA9
+Macromolecule #13: NDUFAB1-alpha
+Macromolecule #14: NDUFA12
+Macromolecule #15: NDUFA13
+Macromolecule #16: NDUFA11
+Macromolecule #17: NDUFB2
+Macromolecule #18: NDUFB3
+Macromolecule #19: NDUFB4
+Macromolecule #20: NDUFB5
+Macromolecule #21: NDUFB6
+Macromolecule #22: NDUFB7
+Macromolecule #23: NDUFB8
+Macromolecule #24: NDUFB9
+Macromolecule #25: NDUFB10
+Macromolecule #26: NDUFB11
+Macromolecule #27: NDUFC2
+Macromolecule #28: NDUEG1
+Macromolecule #29: NDUEG2
+Macromolecule #30: NDUEG3
+Macromolecule #31: NDUEG4
+Macromolecule #32: NDUEG5
+Macromolecule #33: NDUEG6
+Macromolecule #34: NDUEG8
+Macromolecule #35: NDUEG10
+Macromolecule #36: NDUEG11
+Macromolecule #37: NDUEG12
+Macromolecule #38: NDUEG13
+Macromolecule #39: NDUFX
+Macromolecule #40: NDUCA1
+Macromolecule #41: NDUCA2
+Macromolecule #42: NDUCA3
+Macromolecule #43: ND1
+Macromolecule #44: ND2A
+Macromolecule #45: ND3
+Macromolecule #46: NADH-ubiquinone oxidoreductase chain 4
+Macromolecule #47: ND5
+Macromolecule #48: NDUFS2
+Macromolecule #49: NDUFS3
+Macromolecule #50: NDUFS4
+Macromolecule #51: NDUFS5
+Macromolecule #52: NDUFS6
+Macromolecule #53: NDUFS7
+Macromolecule #54: NDUFS8
+Macromolecule #55: UNK-UNK-UNK-UNK-UNK-UNK-UNK-UNK-UNK-UNK-UNK-UNK
+Macromolecule #56: NDUFV1
+Macromolecule #57: NDUFV2
+Macromolecule #58: NDUEG7
+Macromolecule #59: NDUFAB1-beta
+Macromolecule #60: FE2/S2 (INORGANIC) CLUSTER
+Macromolecule #61: IRON/SULFUR CLUSTER
+Macromolecule #62: POTASSIUM ION
+Macromolecule #63: 1,2-DIACYL-SN-GLYCERO-3-PHOSPHOCHOLINE
+Macromolecule #64: CARDIOLIPIN
+Macromolecule #65: NADPH DIHYDRO-NICOTINAMIDE-ADENINE-DINUCLEOTIDE PHOSPHATE
+Macromolecule #66: S-[2-({N-[(2S)-2-hydroxy-3,3-dimethyl-4-(phosphonooxy)butanoyl]-b...
+Macromolecule #67: 1,2-Distearoyl-sn-glycerophosphoethanolamine
+Macromolecule #68: UBIQUINONE-10
+Macromolecule #69: ZINC ION
+Macromolecule #70: FLAVIN MONONUCLEOTIDE
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 3.5 mg/mL | |||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 7.4 Component:
Details: SEC-Q buffer (30 mM Tris pH 7.4, 200 mM NaCl, 1mM EDTA, 0.002% PMSF (w/v), 0.001% CHS (w/v), 0.007% LMNG (w/v)) | |||||||||||||||||||||
| Grid | Model: Quantifoil R0.6/1 / Material: COPPER / Mesh: 300 / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 120 sec. / Pretreatment - Atmosphere: AIR / Details: 25mA | |||||||||||||||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277.15 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: FEI FALCON IV (4k x 4k) / Digitization - Dimensions - Width: 4096 pixel / Digitization - Dimensions - Height: 4096 pixel / Number grids imaged: 1 / Number real images: 2702 / Average exposure time: 7.0 sec. / Average electron dose: 61.5 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 2.0 µm / Nominal defocus min: 0.8 µm / Nominal magnification: 130000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Refinement | Space: REAL / Protocol: RIGID BODY FIT |
|---|---|
| Output model |  PDB-8j9h: |
 Movie
Movie Controller
Controller






























 Z
Z Y
Y X
X