Yorodumi
Yorodumi+ Open data
Open data
 Loading...
Loading...
- Basic information
Basic information
| Entry |  | |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of Euglena gracilis super-complex I+III2+IV, composite | |||||||||||||||
 Map data Map data | ||||||||||||||||
 Sample Sample |
| |||||||||||||||
 Keywords Keywords | Electron transport chain / supercomplex / membrane protein / Euglena gracilis / ELECTRON TRANSPORT | |||||||||||||||
| Function / homology |  Function and homology information Function and homology informationcytochrome-c oxidase / quinol-cytochrome-c reductase / mitochondrial electron transport, cytochrome c to oxygen / quinol-cytochrome-c reductase activity / cytochrome-c oxidase activity / mitochondrial electron transport, ubiquinol to cytochrome c / NADH:ubiquinone reductase (H+-translocating) / ubiquinone binding / electron transport coupled proton transport / NADH dehydrogenase activity ...cytochrome-c oxidase / quinol-cytochrome-c reductase / mitochondrial electron transport, cytochrome c to oxygen / quinol-cytochrome-c reductase activity / cytochrome-c oxidase activity / mitochondrial electron transport, ubiquinol to cytochrome c / NADH:ubiquinone reductase (H+-translocating) / ubiquinone binding / electron transport coupled proton transport / NADH dehydrogenase activity / NADH dehydrogenase (ubiquinone) activity / ATP synthesis coupled electron transport / respiratory electron transport chain / mitochondrial membrane / electron transfer activity / oxidoreductase activity / mitochondrial inner membrane / copper ion binding / heme binding / membrane / metal ion binding Similarity search - Function Cytochrome C oxidase subunit II, transmembrane domain / Cytochrome C oxidase subunit II, transmembrane domain / Cytochrome oxidase subunit II transmembrane region profile. / Cytochrome c oxidase subunit III-like superfamily / Cytochrome c1, transmembrane anchor, C-terminal / Cytochrome c/quinol oxidase subunit II / : / Copper centre Cu(A) / Cytochrome C oxidase subunit II, transmembrane domain superfamily / CO II and nitrous oxide reductase dinuclear copper centers signature. ...Cytochrome C oxidase subunit II, transmembrane domain / Cytochrome C oxidase subunit II, transmembrane domain / Cytochrome oxidase subunit II transmembrane region profile. / Cytochrome c oxidase subunit III-like superfamily / Cytochrome c1, transmembrane anchor, C-terminal / Cytochrome c/quinol oxidase subunit II / : / Copper centre Cu(A) / Cytochrome C oxidase subunit II, transmembrane domain superfamily / CO II and nitrous oxide reductase dinuclear copper centers signature. / Cytochrome c1 / Cytochrome C1 family / Cytochrome c oxidase, subunit I, copper-binding site / Heme-copper oxidase catalytic subunit, copper B binding region signature. / Cytochrome c oxidase-like, subunit I domain / Cytochrome oxidase subunit I profile. / Cytochrome b/b6, C-terminal / Cytochrome b(C-terminal)/b6/petD / Cytochrome b/b6 C-terminal region profile. / Cytochrome b/b6, C-terminal domain superfamily / Cytochrome b/b6/petB / Cytochrome C oxidase subunit II, periplasmic domain / Cytochrome c oxidase subunit I / Cytochrome c oxidase-like, subunit I superfamily / Cytochrome C and Quinol oxidase polypeptide I / Cytochrome c oxidase subunit II-like C-terminal / Cytochrome oxidase subunit II copper A binding domain profile. / Cytochrome b/b6, N-terminal / Cytochrome b/b6-like domain superfamily / Cytochrome b/b6 N-terminal region profile. / Di-haem cytochrome, transmembrane / Peptidase M16, N-terminal / Insulinase (Peptidase family M16) / Metalloenzyme, LuxS/M16 peptidase-like / NADH:ubiquinone oxidoreductase / NADH:quinone oxidoreductase/Mrp antiporter, membrane subunit / NADH:quinone oxidoreductase/Mrp antiporter, TM / Cytochrome c-like domain superfamily / Cupredoxin Similarity search - Domain/homology Cytochrome b / NADH-ubiquinone oxidoreductase chain 4 / Cytochrome c1, heme protein / Ubiquinol-cytochrome-c reductase complex core protein 2, mitochondrial / Ubiquinol-cytochrome-C reductase complex subunit IX, mitochondrial / Cytochrome c oxidase subunit 1 / Putative NADH dehydrogenase subunit 6 / Cytochrome c oxidase subunit 2 Similarity search - Component | |||||||||||||||
| Biological species |  Euglena gracilis (euglena) Euglena gracilis (euglena) | |||||||||||||||
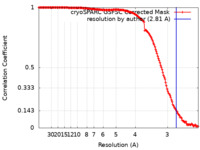
| Method | single particle reconstruction / cryo EM / Resolution: 2.81 Å | |||||||||||||||
 Authors Authors | Wu MC / Tian HT / He ZX / Hu YQ / Zhou L | |||||||||||||||
| Funding support |  China, 1 items China, 1 items
| |||||||||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2024 Journal: Nat Commun / Year: 2024Title: Euglena's atypical respiratory chain adapts to the discoidal cristae and flexible metabolism. Authors: Zhaoxiang He / Mengchen Wu / Hongtao Tian / Liangdong Wang / Yiqi Hu / Fangzhu Han / Jiancang Zhou / Yong Wang / Long Zhou /  Abstract: Euglena gracilis, a model organism of the eukaryotic supergroup Discoba harbouring also clinically important parasitic species, possesses diverse metabolic strategies and an atypical electron ...Euglena gracilis, a model organism of the eukaryotic supergroup Discoba harbouring also clinically important parasitic species, possesses diverse metabolic strategies and an atypical electron transport chain. While structures of the electron transport chain complexes and supercomplexes of most other eukaryotic clades have been reported, no similar structure is currently available for Discoba, limiting the understandings of its core metabolism and leaving a gap in the evolutionary tree of eukaryotic bioenergetics. Here, we report high-resolution cryo-EM structures of Euglena's respirasome I + III + IV and supercomplex III + IV. A previously unreported fatty acid synthesis domain locates on the tip of complex I's peripheral arm, providing a clear picture of its atypical subunit composition identified previously. Individual complexes are re-arranged in the respirasome to adapt to the non-uniform membrane curvature of the discoidal cristae. Furthermore, Euglena's conformationally rigid complex I is deactivated by restricting ubiquinone's access to its substrate tunnel. Our findings provide structural insights for therapeutic developments against euglenozoan parasite infections. | |||||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_35720.map.gz emd_35720.map.gz | 311.1 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-35720-v30.xml emd-35720-v30.xml emd-35720.xml emd-35720.xml | 127.7 KB 127.7 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_35720_fsc.xml emd_35720_fsc.xml | 14.7 KB | Display |  FSC data file FSC data file |
| Images |  emd_35720.png emd_35720.png | 105.4 KB | ||
| Filedesc metadata |  emd-35720.cif.gz emd-35720.cif.gz | 24.2 KB | ||
| Others |  emd_35720_half_map_1.map.gz emd_35720_half_map_1.map.gz emd_35720_half_map_2.map.gz emd_35720_half_map_2.map.gz | 318.8 MB 318.8 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-35720 http://ftp.pdbj.org/pub/emdb/structures/EMD-35720 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-35720 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-35720 | HTTPS FTP |
-Related structure data
| Related structure data |  8iufMC  8iujC  8j9hC  8j9iC  8j9jC C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_35720.map.gz / Format: CCP4 / Size: 343 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_35720.map.gz / Format: CCP4 / Size: 343 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
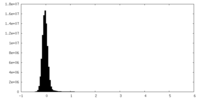
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.2 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: #2
| File | emd_35720_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
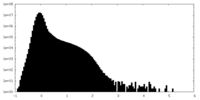
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #1
| File | emd_35720_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
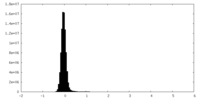
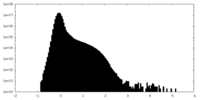
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
+Entire : Euglena gracilis respirasome supercomplex I+III2+IV
| Entire | Name: Euglena gracilis respirasome supercomplex I+III2+IV |
|---|---|
| Components |
|
+Supramolecule #1: Euglena gracilis respirasome supercomplex I+III2+IV
| Supramolecule | Name: Euglena gracilis respirasome supercomplex I+III2+IV / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#89 |
|---|---|
| Source (natural) | Organism:  Euglena gracilis (euglena) Euglena gracilis (euglena) |
| Molecular weight | Theoretical: 2.5 MDa |
+Supramolecule #2: Euglena gracilis respirasome supercomplex I+III2+IV
| Supramolecule | Name: Euglena gracilis respirasome supercomplex I+III2+IV / type: complex / ID: 2 / Parent: 1 / Macromolecule list: #1-#89 |
|---|---|
| Source (natural) | Organism:  Euglena gracilis (euglena) Euglena gracilis (euglena) |
+Macromolecule #1: NDUFS1a
| Macromolecule | Name: NDUFS1a / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Euglena gracilis (euglena) Euglena gracilis (euglena) |
| Molecular weight | Theoretical: 43.888836 KDa |
| Sequence | String: MRRVLARFGP HVPRSFHTTV SRLQEAVPAS ILNAPVGLQP SQTVTCWIDH ILCEFQYPAD ITVFELARRN GINIPHFCYN RNLPIAGNC RMCMCHRVSD KKYAIACNEI AEPNAKYITV DDNLKNIRQY ILEFILANHS LDCPICDQGG ECDLQDLAEL Y GYDTSRYD ...String: MRRVLARFGP HVPRSFHTTV SRLQEAVPAS ILNAPVGLQP SQTVTCWIDH ILCEFQYPAD ITVFELARRN GINIPHFCYN RNLPIAGNC RMCMCHRVSD KKYAIACNEI AEPNAKYITV DDNLKNIRQY ILEFILANHS LDCPICDQGG ECDLQDLAEL Y GYDTSRYD YSDIKHEPDD MPINFLIKSD MNRCIHCTKC VRFLDNFSDD GKEGELGLMG RDPQTICVFR DDGNPQSYVA DI LSANVIE ICPVGALTGR ETNHETRPWE ITRLDAINIF DGTLSAINVE VKEGTELYRV NASKDPQNPD MLLNNEFITD RAR EAPQGN EFKRMTANYA ISLDNKKLLL HHALRLYAID PLFRSKALFL LADIMNEDRH LPETKQLGH |
+Macromolecule #2: NDUFS1b
| Macromolecule | Name: NDUFS1b / type: protein_or_peptide / ID: 2 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Euglena gracilis (euglena) Euglena gracilis (euglena) |
| Molecular weight | Theoretical: 59.671816 KDa |
| Sequence | String: MASGSEVLRQ FLTIRKNSYK YAPAFQRLHA LVNGANSAAK LRARHQKRLG INVVLGEKSD LGLCQLADTL ADRLKLADLG VSARPAKSP AVYYGHLAAQ QHRYAVPSEL KYTESSYSSR NVYIWLWTDV QQEAPDLHTQ IFTGPTSNCN VYSFGHVHNA R AGVKPVGG ...String: MASGSEVLRQ FLTIRKNSYK YAPAFQRLHA LVNGANSAAK LRARHQKRLG INVVLGEKSD LGLCQLADTL ADRLKLADLG VSARPAKSP AVYYGHLAAQ QHRYAVPSEL KYTESSYSSR NVYIWLWTDV QQEAPDLHTQ IFTGPTSNCN VYSFGHVHNA R AGVKPVGG MEEFVGWLEG RTNLFSRTPK LETRLSNVYV LYSDNFLEMF PTNYGDIFKK IEELLGDQTF VSFSYLSRHP VS YNAVQTY AFPPVTQLLK RNDQYRLNVL TNVQRQDYSE NESRGRFTAR LMCHSTLLRA DQPMNELVIA QKTPAEDNAA LAY IDKFGD YKSAINSIFI SEFSDKLQLM HPHQLLTYAF ALLAWPRALA RLLPLTSIPK ADEEKTFKAT HSQFLERLIR DFDN DPTRL SLIHALSLGR PALVEDLRLR LWPYTVVPGT AFNVVKAKAL LQRLNATPEY SPDGPYYEFQ TPAAPVPSAA PTPAP QRVA LKSDSIFAID CEFVRHSMPL RGHINEVNRK QHLSWCKLAP ESK |
+Macromolecule #3: ND2b
| Macromolecule | Name: ND2b / type: protein_or_peptide / ID: 3 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Euglena gracilis (euglena) Euglena gracilis (euglena) |
| Molecular weight | Theoretical: 15.756794 KDa |
| Sequence | String: MNNNLQIENY TNKNKIVISP ISYIGNNHPY KMYTIINLCI SSSLLITNYT IAKTSIFLYL IYIFNNNIYF IIIMLFFVLY PIIFIVLIH PFIIISVNNH LINKANNKGI IINNFI(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) ...String: MNNNLQIENY TNKNKIVISP ISYIGNNHPY KMYTIINLCI SSSLLITNYT IAKTSIFLYL IYIFNNNIYF IIIMLFFVLY PIIFIVLIH PFIIISVNNH LINKANNKGI IINNFI(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK) (UNK)(UNK) |
+Macromolecule #4: COXEG1
| Macromolecule | Name: COXEG1 / type: protein_or_peptide / ID: 4 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Euglena gracilis (euglena) Euglena gracilis (euglena) |
| Molecular weight | Theoretical: 27.278852 KDa |
| Sequence | String: MLSRALCSRN VPMSLKALNR PGSAGKLPMQ LMKYASAVAP ISHEGTLVRI SQVKKLSELQ LHFNDSHLGE SELAAKVLGK LRKLEAEVL ARNQAFNEAH PLVFDPKRAF NDEIFLCCSL CCIIFLIFLF NQYEEFAHEL SFDIREQFGL GFYMLLGLHG S HVIFGTIM ...String: MLSRALCSRN VPMSLKALNR PGSAGKLPMQ LMKYASAVAP ISHEGTLVRI SQVKKLSELQ LHFNDSHLGE SELAAKVLGK LRKLEAEVL ARNQAFNEAH PLVFDPKRAF NDEIFLCCSL CCIIFLIFLF NQYEEFAHEL SFDIREQFGL GFYMLLGLHG S HVIFGTIM LALLTLWGAQ GSVGPQSHAL RFTSLYVHLV DLVFIILVLA IYSANASPEL YGGIVPNILE ARTFVSVDAA GN PQIKEF |
+Macromolecule #5: COXEG3
| Macromolecule | Name: COXEG3 / type: protein_or_peptide / ID: 5 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Euglena gracilis (euglena) Euglena gracilis (euglena) |
| Molecular weight | Theoretical: 16.394043 KDa |
| Sequence | String: MMQRIPFKKP NQIRGYFTRV HKYNHVPVPF ILNVGMSISI VTSFVYFTYT SLWVRPEYDR VVDPSKAYVN PVWVDYWLKL RDEKRIQGA LERSILEEEP EKAAEKILEW ARTSAQNKIL EDLKLLKPAL SPATIAQFEK |
+Macromolecule #6: COXEG4
| Macromolecule | Name: COXEG4 / type: protein_or_peptide / ID: 6 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Euglena gracilis (euglena) Euglena gracilis (euglena) |
| Molecular weight | Theoretical: 19.359492 KDa |
| Sequence | String: MSTVLSILGK RFQRSALTPK MNPFIRIRCQ GPIEEFQRGF IGEFHAFALP GACMLVASCL GTFHIIRCLV VNPELSLAKV IPEILQPFT NPNAQLKAAD GKDDDDSQVP KQWGMWGRHP NYGVLHVPFL DALNKEALAR GKDGVNMGAE YNLVFTKSMA D QVVDLILD DVQKRV |
+Macromolecule #7: COXEG5
| Macromolecule | Name: COXEG5 / type: protein_or_peptide / ID: 7 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Euglena gracilis (euglena) Euglena gracilis (euglena) |
| Molecular weight | Theoretical: 19.464051 KDa |
| Sequence | String: MPSSMAWTIG WGFYAAWIMK ETWNLRSSSV GWTPITLMEA YKTKERYLRS KAMMERYNSE LEAVDDSNIT EEDAKKFELE KATPSISIW EQFRSNPYWK EVEEEISTDV RKTMLEKHPD YALLLEAVKK SGYSKLWHLP GPWMNEHYND GLHGRFLGWT P KAAHH |
+Macromolecule #8: COXEG6
| Macromolecule | Name: COXEG6 / type: protein_or_peptide / ID: 8 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Euglena gracilis (euglena) Euglena gracilis (euglena) |
| Molecular weight | Theoretical: 8.728172 KDa |
| Sequence | String: VFPSITKPLG LFKNLPRQHR AARDASIWLA ILTAGPFGIF IAFKYYADWY DKKLLMEYYK DSIVYGETYG KGKYV |
+Macromolecule #9: COXEG7
| Macromolecule | Name: COXEG7 / type: protein_or_peptide / ID: 9 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Euglena gracilis (euglena) Euglena gracilis (euglena) |
| Molecular weight | Theoretical: 35.651516 KDa |
| Sequence | String: MLRQVVRRSN PLRMQVRGSA WNFQELMESR IPDYKGRPNR SGAELEQVKA ALPKIEFMTS YEFDVLTKTR SNLTKEYSYQ RDMRLKVTE LMLDEAPHEL EGLAVEGDAA LKQLAELKAL QTLTEYAGDL LEGQNQIVQR VNDFVDSNPV YLLDQPLREE A RWNLLPEM ...String: MLRQVVRRSN PLRMQVRGSA WNFQELMESR IPDYKGRPNR SGAELEQVKA ALPKIEFMTS YEFDVLTKTR SNLTKEYSYQ RDMRLKVTE LMLDEAPHEL EGLAVEGDAA LKQLAELKAL QTLTEYAGDL LEGQNQIVQR VNDFVDSNPV YLLDQPLREE A RWNLLPEM DHKTRSLVRT ELRDWLPAEY RQTRAVDLQQ VAAFSPPVKA DMFRAIEARA KDAEAEIRSL PPAEQAGLLA LV KDNVAKS KAFIDPTYDI TPEAINACND VDALRAMAHR VTEYSGDARL LAIYGKAAQL TGDTAAQAIL KEAKDLVF |
+Macromolecule #10: COXEG8
| Macromolecule | Name: COXEG8 / type: protein_or_peptide / ID: 10 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Euglena gracilis (euglena) Euglena gracilis (euglena) |
| Molecular weight | Theoretical: 24.49532 KDa |
| Sequence | String: MLRVLTPAIA RPGLRCFFKD GFRDNASLEL VYRVVLKSPA VSQKLIEFYA KSLDQLSVES LSALKGTTVG IPLQPYLGDP HRVLLAYSL LPHTVETEAD GNPVVETKIG DEEQKIKIID SEVISFLAKE ILGKLGLETT PQAARQYLDS LVEGAEALYA K IAPVEPSP ...String: MLRVLTPAIA RPGLRCFFKD GFRDNASLEL VYRVVLKSPA VSQKLIEFYA KSLDQLSVES LSALKGTTVG IPLQPYLGDP HRVLLAYSL LPHTVETEAD GNPVVETKIG DEEQKIKIID SEVISFLAKE ILGKLGLETT PQAARQYLDS LVEGAEALYA K IAPVEPSP LEKAIAEINE EIKSGTPWDT LKNRADPKEL HALKFAQLPH PITKKVEGKF KYF |
+Macromolecule #11: COXEG9
| Macromolecule | Name: COXEG9 / type: protein_or_peptide / ID: 11 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Euglena gracilis (euglena) Euglena gracilis (euglena) |
| Molecular weight | Theoretical: 31.754857 KDa |
| Sequence | String: MMNKGRILLG TNPGDIALNS KRFTVGKFVA WACGGWGLKD WIFPSLFIGR GDGPDFDRIV KHTLQSSSAI EKVNWFDSPF ACYTEWFVE HFPGFFDSRY RFEMSAKTIL ANKYPIKDFP VVDMRSWRSS RLFDLFEVPH PEHTFVFGGP VLLNTEAKRA E RLEQEWHG ...String: MMNKGRILLG TNPGDIALNS KRFTVGKFVA WACGGWGLKD WIFPSLFIGR GDGPDFDRIV KHTLQSSSAI EKVNWFDSPF ACYTEWFVE HFPGFFDSRY RFEMSAKTIL ANKYPIKDFP VVDMRSWRSS RLFDLFEVPH PEHTFVFGGP VLLNTEAKRA E RLEQEWHG KDGTFVDVHP LNVATESHTE VSVIGGIKVY NGVWQGGKDS WKRDSAKPEL TAPFHSPIWY RNMFIVKNAD QL VEHFGEN LSDETWQEVR KEHLAFHERF HKDYSFA |
+Macromolecule #12: COXEG10
| Macromolecule | Name: COXEG10 / type: protein_or_peptide / ID: 12 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Euglena gracilis (euglena) Euglena gracilis (euglena) |
| Molecular weight | Theoretical: 10.043338 KDa |
| Sequence | String: MNHERPWVFL NKVTGKWGAC GWQPFWKATS QSLNYIPDNF IALRTNGSWV RNLWQSSKVL DRGLNASGYP TVGTTDWVRI PLDAPLEA |
+Macromolecule #13: ND4L
| Macromolecule | Name: ND4L / type: protein_or_peptide / ID: 13 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Euglena gracilis (euglena) Euglena gracilis (euglena) |
| Molecular weight | Theoretical: 19.673094 KDa |
| Sequence | String: MLKAIQPMSL RSRVVSRALS SGYKQFGGLT FKEVQNNEVK QNPTTFMQRP LAQIASNGLF VAWPGQFETV FDLLTSQIGP YCVIGLYLG ARGCFKPEMA WTDRLIHVEA STFLLYGVFF ITFASTPLLY WAWFFMLFSN SLKTLMFVHL SNPWYLVLDQ P MQVKFSLK RLK |
+Macromolecule #14: COX5b-2
| Macromolecule | Name: COX5b-2 / type: protein_or_peptide / ID: 14 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Euglena gracilis (euglena) Euglena gracilis (euglena) |
| Molecular weight | Theoretical: 19.736387 KDa |
| Sequence | String: MFRRGLVLAA SRSKSLLDSV HVFRPEFAQG KFRIDLDQPA KQNKLQQQIY TLTDDEREMY EDEPYIGVDH LYEAHKGSKE NPVVVEAIG VHGNDVFTGC LGGCHKDAAD AVAYYTIVPP NTLAVCIDCG IHFVARINEQ LTFWPDGTQP WEKVDFKAVE G FLYKHYKY GSPLMI |
+Macromolecule #15: COX5c
| Macromolecule | Name: COX5c / type: protein_or_peptide / ID: 15 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Euglena gracilis (euglena) Euglena gracilis (euglena) |
| Molecular weight | Theoretical: 23.788312 KDa |
| Sequence | String: YFFSRARMSL QLGHMRVGAL WQSNANSRGL PNLLKEVMAA EPIYPYTHPG LSAKQILWNT KFNLKLEKDL ISIAESAVQS QTVNKVVIP TMLTGDVNPK SPASVVAKDK YQLAKQMAEF QYKKSYFEKP WGYMFAYYCD EDRARLCGGI GYVGHHGGGD A EILSYTEY ...String: YFFSRARMSL QLGHMRVGAL WQSNANSRGL PNLLKEVMAA EPIYPYTHPG LSAKQILWNT KFNLKLEKDL ISIAESAVQS QTVNKVVIP TMLTGDVNPK SPASVVAKDK YQLAKQMAEF QYKKSYFEKP WGYMFAYYCD EDRARLCGGI GYVGHHGGGD A EILSYTEY VSTVVYPTIF YMAITFVVSL FLMYYYIYGT FFSSTRNRFD |
+Macromolecule #16: COX6a
| Macromolecule | Name: COX6a / type: protein_or_peptide / ID: 16 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Euglena gracilis (euglena) Euglena gracilis (euglena) |
| Molecular weight | Theoretical: 13.079197 KDa |
| Sequence | String: MSTNKNVFFP PALHLKENSI FQYKFKNLAV RHDAARLGII LAGPTLFYWF TVYYFKGMPN GLPPVLLNPF VNRNYRGQKR DMPWGTDCA FLDTKCHEEK DTWAKRGPFF IIA |
+Macromolecule #17: COX6b-1
| Macromolecule | Name: COX6b-1 / type: protein_or_peptide / ID: 17 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Euglena gracilis (euglena) Euglena gracilis (euglena) |
| Molecular weight | Theoretical: 32.995148 KDa |
| Sequence | String: MEGFVDKIDD NKYLGKWETI LTDGRTHLPK HITFHDAAAI SARWNQQYVN DSGPVYYRHW LACQQTYGAG NEDCRKLRWW AQQITHPLH LAEWDDWWKD EHYDLQIGQH WNRICGEEFE EASNLLKDLK EKREGLAAKF RDLLKTKTAE DPMGKILHEV A QLEEPSKT ...String: MEGFVDKIDD NKYLGKWETI LTDGRTHLPK HITFHDAAAI SARWNQQYVN DSGPVYYRHW LACQQTYGAG NEDCRKLRWW AQQITHPLH LAEWDDWWKD EHYDLQIGQH WNRICGEEFE EASNLLKDLK EKREGLAAKF RDLLKTKTAE DPMGKILHEV A QLEEPSKT PVADLVEAGT LSKEAVEAAA ALKIKELKAL RDDATWAEVK GSLLNGVTTT CSTLKKTSKV VAELKAQAEL ER NKTSAVK LDIPHMRVNY EKPGLYEYDT WFGKFLPRTP QFGFAESDEE |
+Macromolecule #18: COX7a
| Macromolecule | Name: COX7a / type: protein_or_peptide / ID: 18 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Euglena gracilis (euglena) Euglena gracilis (euglena) |
| Molecular weight | Theoretical: 20.078975 KDa |
| Sequence | String: MLSRLLHGPG SDKGHGYFHP DHGLVDSAAQ NIRGPYWHAE NMQFMQQTTQ SGEKLPVPLT ETAAQSSKLP VLSAQDGKTL QKIQLTDGS KVPKAATLVK WNPKSVHQWH SDDILRVDMT KHPEWVRTMR MFHGTVWRHR PEFRHPWFSR GSRASGIIML V FGAVAYGE ITFGKRLGGV |
+Macromolecule #19: COX7c
| Macromolecule | Name: COX7c / type: protein_or_peptide / ID: 19 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Euglena gracilis (euglena) Euglena gracilis (euglena) |
| Molecular weight | Theoretical: 19.897588 KDa |
| Sequence | String: MQKAVVGNLL RQLSLTRPRG HGSNLYNRVH GNLPQLYVEQ LYNTDEILDT VPHSSDPVHH MFPKCAAASP LGFRPYDNNK LWDAFVLWA IVYGVTTIFC VLHILKYPQI WKHLFETLTF QYTYQREIGE EYVWQYGGGG LNPAKWRFVK SPLEELGLVT E YRSDLDYP DDL |
+Macromolecule #20: NDUFA1
| Macromolecule | Name: NDUFA1 / type: protein_or_peptide / ID: 20 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Euglena gracilis (euglena) Euglena gracilis (euglena) |
| Molecular weight | Theoretical: 15.659606 KDa |
| Sequence | String: MPGGGGWSNM VPIIILNGVV WAALGRASLA CSPPEFHKRT KNDTEFNKYL HLRFNKAVQN PESVAGQAVK AGCAPEFRPF DSPANPLVV VYGWKDEIQP RPNPGSLAQS FDDRGLSWYQ SHFSNRVVDD PKHNSLPFPG YY |
+Macromolecule #21: NDUFA2
| Macromolecule | Name: NDUFA2 / type: protein_or_peptide / ID: 21 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Euglena gracilis (euglena) Euglena gracilis (euglena) |
| Molecular weight | Theoretical: 21.331168 KDa |
| Sequence | String: MAQVWRSRLS CHFRKLRVRY PAAKLPEAAA INWATYLDVP SPANLPAADL NKALEAMRRP NPALASSRGV REFVQRVVPE LEAENPFCP LIVDKFDPEV ASQFPSESTD PTLHAHFLDG TQVNVPLANK SAAEIEDILA DLVKLAGLLQ PQAPLEGDNL P VEDTIYAA ...String: MAQVWRSRLS CHFRKLRVRY PAAKLPEAAA INWATYLDVP SPANLPAADL NKALEAMRRP NPALASSRGV REFVQRVVPE LEAENPFCP LIVDKFDPEV ASQFPSESTD PTLHAHFLDG TQVNVPLANK SAAEIEDILA DLVKLAGLLQ PQAPLEGDNL P VEDTIYAA ASRPRFPNYS RHAKQARLGD ESTEM |
+Macromolecule #22: NDUFA3
| Macromolecule | Name: NDUFA3 / type: protein_or_peptide / ID: 22 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Euglena gracilis (euglena) Euglena gracilis (euglena) |
| Molecular weight | Theoretical: 14.950273 KDa |
| Sequence | String: MDRYEFQKIR RQPPTLHWEA GNRFENIQRL RWENAALLKD PKLTWFRREM LMRPAFFHCT LFAGAVAVGY PFVAYFYEKV FPDRQDFRS TMTLLRAVGG LEEQEYYIME RAKAIERAKA RAAVQG |
+Macromolecule #23: NDUFA5
| Macromolecule | Name: NDUFA5 / type: protein_or_peptide / ID: 23 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Euglena gracilis (euglena) Euglena gracilis (euglena) |
| Molecular weight | Theoretical: 20.961854 KDa |
| Sequence | String: MLRAGLRCAS AAAVIGRRGY ATKVKDFTGL AGYPVEPKWR GKLLALYAET LEAVQGIPAD HPYRRSVENL TRHYARVVEE AEDYEVVET TVGLGQVEQL IRMATNELQL VRDFTVWRTW EIDERVLDKM DVDLRTSIGG HYTKRAYEVL DEVEDERRLL E KLQQEVDL EKARVAAKAA EGPKTL |
+Macromolecule #24: NDUFA6
| Macromolecule | Name: NDUFA6 / type: protein_or_peptide / ID: 24 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Euglena gracilis (euglena) Euglena gracilis (euglena) |
| Molecular weight | Theoretical: 48.78698 KDa |
| Sequence | String: MPQMRVATKG VRPFSVRAAG GQYVLPDHGR YGQVVRPARL EEFELNPHQN PSRDRDWSVE IRGFYRDLLK SIPTMKQRFR LVIPNDVVR QNIRKRFEQG PKLTDPAALR HRALMVSADL EEYFREDFLD SQVQGKYNNM DPRTLLNQEI AAAASETQTA H RFFNEGTN ...String: MPQMRVATKG VRPFSVRAAG GQYVLPDHGR YGQVVRPARL EEFELNPHQN PSRDRDWSVE IRGFYRDLLK SIPTMKQRFR LVIPNDVVR QNIRKRFEQG PKLTDPAALR HRALMVSADL EEYFREDFLD SQVQGKYNNM DPRTLLNQEI AAAASETQTA H RFFNEGTN VLLETGIGGE DVTENRVYIT REQAYRKGLA SLRGDAAVRH LLPAVDPANQ TTLQALAAEN DLQALVDLLG HL PAAKTAE AYVQRCEAFH KEAGLRHQKA SGGAVLAAWE KFKDEEVNST VLLHPAYKAL IADPSRNPLL RGAADWVRLV EAG GLSTTE PDSAADKLLK VAQHLYYSDQ LPEGFAQDLG VSYLADLKGV DRRLDLLLDE EIAYRQELLL KIYAHTVESI KATA SNPTD PAAVKKHLDA HDWSAFVVPT EGVKSSYEAL AL |
+Macromolecule #25: NDUFA7
| Macromolecule | Name: NDUFA7 / type: protein_or_peptide / ID: 25 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Euglena gracilis (euglena) Euglena gracilis (euglena) |
| Molecular weight | Theoretical: 16.358675 KDa |
| Sequence | String: MTTLKLNLKE RWLAMVHKRI QQWWQVGGDE RWVHTDPANY WWGAPAGLAS AREERQFFPS YYQRDLRRIK RKSFIMNRPM MKTNALALA DGQLPPRPHL NLADPSPFWW QSHPKDRGDR TMIYQSRSED AIEHYPR |
+Macromolecule #26: NDUFA8
| Macromolecule | Name: NDUFA8 / type: protein_or_peptide / ID: 26 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Euglena gracilis (euglena) Euglena gracilis (euglena) |
| Molecular weight | Theoretical: 25.916332 KDa |
| Sequence | String: MTAPWNSSYT AKDWKKFPEI ESHAALQEKL DREVSSKDVV NVEKVVDDIL SHPHRKHFGT LNIPNEQTKK EHEGIFGMGI PENVLFSFG HWYRMSQPLE KAREEYLVCK HNAPKRAAPT ECLEEAKNMF NLYLHMSEIP FRTCPKQTAD YTYCVETQGH R KPGMDYGA ...String: MTAPWNSSYT AKDWKKFPEI ESHAALQEKL DREVSSKDVV NVEKVVDDIL SHPHRKHFGT LNIPNEQTKK EHEGIFGMGI PENVLFSFG HWYRMSQPLE KAREEYLVCK HNAPKRAAPT ECLEEAKNMF NLYLHMSEIP FRTCPKQTAD YTYCVETQGH R KPGMDYGA RSFTSFVRYC LPEQEAFEEC LGKTYGCKFP PAPAHGQFYQ RSARFTNLPW DNIYA |
+Macromolecule #27: NDUFA9
| Macromolecule | Name: NDUFA9 / type: protein_or_peptide / ID: 27 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Euglena gracilis (euglena) Euglena gracilis (euglena) |
| Molecular weight | Theoretical: 54.983121 KDa |
| Sequence | String: MARPPAEVTM LIKNAGDLLT KVKLENPPTR LLLDPKTIKL ATQDPTVKGK VKDLMLKGVK VEPSTAARVE HTFIPAPKQT ENQYSKPLL GYRLRELRTK VLSNEVYSTP RPRPLRGVVA TVFGGNGFLG NQVVAQLAQY GATVICPTRI NNEEHPVVMN T RDFRQIKS ...String: MARPPAEVTM LIKNAGDLLT KVKLENPPTR LLLDPKTIKL ATQDPTVKGK VKDLMLKGVK VEPSTAARVE HTFIPAPKQT ENQYSKPLL GYRLRELRTK VLSNEVYSTP RPRPLRGVVA TVFGGNGFLG NQVVAQLAQY GATVICPTRI NNEEHPVVMN T RDFRQIKS LGDQGQVFPV VYNPTVFDEV AQCVERSQVV FNCIGGFYPA MNQSQSFGPE ALFANLPRNI ARACAMKGVQ RL VHTSHIN ADVSSPIPFF KYKALGEEAV LDEFPNGIII RPADIFGDRD NFTTLMVNLL KGSNWPIMST NTYLLEGNEY VEC QPVWVV DVARAMVRAA MREYTFGQTY QLPGPDRYKL IEVMRYIEAI TQLQPSHVRV YSPLEAQLRF DRPGGENHRS WIDL HLREN VVPKPGVKTW QDLEIDNSIL TKMENITGDW MSKAPYRDMP TGFDEELTDL SLPRVWGDYD KKLIAFPAVS AVAAV LYAL AILFP |
+Macromolecule #28: NDUFAB1-alpha
| Macromolecule | Name: NDUFAB1-alpha / type: protein_or_peptide / ID: 28 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Euglena gracilis (euglena) Euglena gracilis (euglena) |
| Molecular weight | Theoretical: 14.936051 KDa |
| Sequence | String: MLARFGRVLS VAYRTYPVGK SLVRATPFIE AQKRWGGGHG HGPVTPLDSA IHFLDREDVT KRVLKIVAAF DKVTKEVKAD SHFVNDLGL DSLDVVEVVM SLEQEFNVDM SNQDAEKIHT VKDAIDYFCA HPASM |
+Macromolecule #29: NDUFAB1-beta
| Macromolecule | Name: NDUFAB1-beta / type: protein_or_peptide / ID: 29 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Euglena gracilis (euglena) Euglena gracilis (euglena) |
| Molecular weight | Theoretical: 14.899051 KDa |
| Sequence | String: MLARFGRVLS VAYRTYPVGK SLVRATPFIE AQKRWGGGHG HGPVTPLDSA IHFLDREDVT KRVLKIVAAF DKVTKEVKAD SHFVNDLGL DSLDVVEVVM ALEWEFQVEM HSSEAEMIAS VADAIDYFCL HPAAH |
+Macromolecule #30: NDUFA12
| Macromolecule | Name: NDUFA12 / type: protein_or_peptide / ID: 30 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Euglena gracilis (euglena) Euglena gracilis (euglena) |
| Molecular weight | Theoretical: 33.338133 KDa |
| Sequence | String: MIRTLLLRHS APAVITGRPQ WWTQAIAVPP TQAEMELFQP KEVVHTKPYK PHPWFKDFGQ GRRHIVGPPE RGEFWRFRKF YAVMREKTK ELGVRGALRF LVRKLRTQRE AWYEKGYEED ILVGEDEMGN KYWQSSYTTA VQSRWVEYGT GSTFTKDASV V APEWYQWL ...String: MIRTLLLRHS APAVITGRPQ WWTQAIAVPP TQAEMELFQP KEVVHTKPYK PHPWFKDFGQ GRRHIVGPPE RGEFWRFRKF YAVMREKTK ELGVRGALRF LVRKLRTQRE AWYEKGYEED ILVGEDEMGN KYWQSSYTTA VQSRWVEYGT GSTFTKDASV V APEWYQWL HGAPDPEVQE LRPRHPAALT KGLTGDYWYR MKHSESQYAF GRKYWPRGNP HPKNTKYDDF LLRKRRLSKR RG FMEFDPF VLPAERLRKR AKWAPNPVSD RRHSAYSKNL PLGA |
+Macromolecule #31: NDUFA13
| Macromolecule | Name: NDUFA13 / type: protein_or_peptide / ID: 31 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Euglena gracilis (euglena) Euglena gracilis (euglena) |
| Molecular weight | Theoretical: 22.683824 KDa |
| Sequence | String: MAEKAPEVHH PQDKLPEWFD PNSQQFTRNW RQAMPPKGGY PAVRHRYLSP GKGVSMRTAW AIIGSLTLFG LYQRRVDRAF ILDLGDDFH ARNCAAVPYM EAENQLKDLI SLHYEKEYRN FVAPGLKLDP SQFFNHPIPG AVGEYIRKTK PQRSTKNRVA W MNLELCTE ...String: MAEKAPEVHH PQDKLPEWFD PNSQQFTRNW RQAMPPKGGY PAVRHRYLSP GKGVSMRTAW AIIGSLTLFG LYQRRVDRAF ILDLGDDFH ARNCAAVPYM EAENQLKDLI SLHYEKEYRN FVAPGLKLDP SQFFNHPIPG AVGEYIRKTK PQRSTKNRVA W MNLELCTE PISQLSGAPQ SGPDKYERML GFTVYEGPAF |
+Macromolecule #32: NDUFA11
| Macromolecule | Name: NDUFA11 / type: protein_or_peptide / ID: 32 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Euglena gracilis (euglena) Euglena gracilis (euglena) |
| Molecular weight | Theoretical: 32.617457 KDa |
| Sequence | String: MFQPVWQPIL MVGSPDIILH SAERRALAWD HPNRFSALRN ALYQARLLEQ PRPENRIALL GQDLLEDTIY TTVGAYLFAG VSCIQRLGG HVPFTPSFTG QNIWTMPKWA SRLLHQVRMM RYFSAYWAVG MTYFTTYNIL TGFMGFPVNE YHNYQPQASV L SVIPTALI ...String: MFQPVWQPIL MVGSPDIILH SAERRALAWD HPNRFSALRN ALYQARLLEQ PRPENRIALL GQDLLEDTIY TTVGAYLFAG VSCIQRLGG HVPFTPSFTG QNIWTMPKWA SRLLHQVRMM RYFSAYWAVG MTYFTTYNIL TGFMGFPVNE YHNYQPQASV L SVIPTALI YAALHPNRRP ERLWVGKATP FVGRFFLSGI VGAALAVFAA RRFAHATVSE LYHPSGSDSY FETLRNSAPS AD LVADMPY IPFYKEARCS PGLPVKSPYY DPEYVAKAKE EVKRKLDSLY |
+Macromolecule #33: NDUFB2
| Macromolecule | Name: NDUFB2 / type: protein_or_peptide / ID: 33 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Euglena gracilis (euglena) Euglena gracilis (euglena) |
| Molecular weight | Theoretical: 17.202416 KDa |
| Sequence | String: RSSDLVRRGA LIASLVRNRT IRPKPLQVRT GSGGHHSHSV DPNDPFDGVE NELFLPPQAY YYFSRILFVT TLIVWLKTYR DKAIDQFSF KKWVDSLPEM TIKEEEIYLD EWEDWEAPQH DVARLFPRYT KKFRKADDWN FGATVW |
+Macromolecule #34: NDUFB3
| Macromolecule | Name: NDUFB3 / type: protein_or_peptide / ID: 34 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Euglena gracilis (euglena) Euglena gracilis (euglena) |
| Molecular weight | Theoretical: 6.831087 KDa |
| Sequence | String: MVFWNHRRYQ FHPFINAWGK NMRHAFFPNL FWSR(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) |
+Macromolecule #35: NDUFB4
| Macromolecule | Name: NDUFB4 / type: protein_or_peptide / ID: 35 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Euglena gracilis (euglena) Euglena gracilis (euglena) |
| Molecular weight | Theoretical: 19.511402 KDa |
| Sequence | String: MAERTFRSLG RLPAPSAPLP HDKIAWPEKI RFLPEGSIMM PDGYSLDPNQ HFEDPYHWSV LRRAWEPKQD VFVGKDSHGN VVKQYNSPQ VRHLHPGFKL FYLYRKGAAR RFQVNAQNIA YAAIFAGTCL ALCEFGLRVQ NWAGGVNTLE KLQSTERKVK M NGKVIVAP DSF |
+Macromolecule #36: NDUFB5
| Macromolecule | Name: NDUFB5 / type: protein_or_peptide / ID: 36 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Euglena gracilis (euglena) Euglena gracilis (euglena) |
| Molecular weight | Theoretical: 15.815024 KDa |
| Sequence | String: FWQAEHKVYP PSKIHPGVHK GYIRTGVVHN LMRLLA(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)WAPVTALA SFATWR YRA EVAEWLDNLS NVDYPKWLAE REIAIRRVET EIFWRIEGHA QQRSQVASGQ GRAMLNKAAK ASYEESEASR LHGAGVS DL LK |
+Macromolecule #37: NDUFB6
| Macromolecule | Name: NDUFB6 / type: protein_or_peptide / ID: 37 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Euglena gracilis (euglena) Euglena gracilis (euglena) |
| Molecular weight | Theoretical: 10.926597 KDa |
| Sequence | String: DIDLGPHPYR PDFAGHAPNP KHWRRNTFFL SCGILLIYYQ CCKLEFYHGR FLEPPIRPFP LQGYFRNVYL DDPNYDDKKK LMKEDFAKL KA |
+Macromolecule #38: NDUFB7
| Macromolecule | Name: NDUFB7 / type: protein_or_peptide / ID: 38 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Euglena gracilis (euglena) Euglena gracilis (euglena) |
| Molecular weight | Theoretical: 12.247089 KDa |
| Sequence | String: EVFDNELVSY EGVDWEKIPN RFKRRRMPLS DEVMDEWKIP YRERDYCVHK LLELRKCVQK TFYRGECDHH RHEYHMCQHL ELRRRKAIK DLREELGR |
+Macromolecule #39: NDUFB8
| Macromolecule | Name: NDUFB8 / type: protein_or_peptide / ID: 39 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Euglena gracilis (euglena) Euglena gracilis (euglena) |
| Molecular weight | Theoretical: 20.488459 KDa |
| Sequence | String: MLATRLRRPA VGLARTTIRM QARTRFMQME YWEVPMEADH PEYRPVVHDQ FLPFVGAPGV HWNPTPYPVG YDLWWNSFEA LAMDIHDMG HQESIGESLE KYLLGAGLCF AIFNFLILFD CHFWFSEVNA GHPLYPKQFH YLLRGGDPTD EIWLAQQQRE K ELMEHGKA LVKGEAKA |
+Macromolecule #40: NDUFB9
| Macromolecule | Name: NDUFB9 / type: protein_or_peptide / ID: 40 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Euglena gracilis (euglena) Euglena gracilis (euglena) |
| Molecular weight | Theoretical: 18.347 KDa |
| Sequence | String: MKRSLLKLKD LKKVKVTQPQ SSIVLKPVPI GEPIIWSRVQ HLYWRALHVI EDFPWMGPSQ TSEQFDIMLS VKEEIQKNAN AAGALKDRL LEDFEAHIKR RDWMQRRVHY FANGGPGYQT FASERHTLEQ QGVWKAGTPI PFPKSDYNDD GPVEKGPYY |
+Macromolecule #41: NDUFB10
| Macromolecule | Name: NDUFB10 / type: protein_or_peptide / ID: 41 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Euglena gracilis (euglena) Euglena gracilis (euglena) |
| Molecular weight | Theoretical: 17.471033 KDa |
| Sequence | String: MGQHFGKLGW DADDVYKVTT FEKHNYTVEE PRTPAFKAPR RPFPICDRPP MPDQEGLYET HADLYEMKRW VMWEWSVYCK ELSLVREEL KWCVAREGSN AHRECHDLAR QYTDMTRKLK MEERFSWLNH FKKPLVKDVL FDLKQ |
+Macromolecule #42: NDUFB11
| Macromolecule | Name: NDUFB11 / type: protein_or_peptide / ID: 42 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Euglena gracilis (euglena) Euglena gracilis (euglena) |
| Molecular weight | Theoretical: 13.048724 KDa |
| Sequence | String: GGVSRPVRSK RETDFNSSDP WETPFGRIEH DDEDIPYTTT VYQPRRPTEW STDVIWHAVL LMFFVWFTTL QGILVPQNKE NDMAWEEAL RRRRLFREKQ V(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) |
+Macromolecule #43: Cytochrome c oxidase subunit 1
| Macromolecule | Name: Cytochrome c oxidase subunit 1 / type: protein_or_peptide / ID: 43 / Number of copies: 1 / Enantiomer: LEVO / EC number: cytochrome-c oxidase |
|---|---|
| Source (natural) | Organism:  Euglena gracilis (euglena) Euglena gracilis (euglena) |
| Molecular weight | Theoretical: 55.759504 KDa |
| Sequence | String: MINNIMHMIN KYTLTTSHKI IGILYGYMGY IAGILGYIIS MLIRMELNTQ GLAIVRKVKE VTIYNNWITI HGLIMLFVFI MPVGIGFYG NYLIPMLIGT SELSMPRMNG ISFWMLIVGV VIFVISNVLM SKPISSGWTL YPPLSTRDAD NIGVNIDLSL L VVHVLGIS ...String: MINNIMHMIN KYTLTTSHKI IGILYGYMGY IAGILGYIIS MLIRMELNTQ GLAIVRKVKE VTIYNNWITI HGLIMLFVFI MPVGIGFYG NYLIPMLIGT SELSMPRMNG ISFWMLIVGV VIFVISNVLM SKPISSGWTL YPPLSTRDAD NIGVNIDLSL L VVHVLGIS STIGSVNYIT TNKYNRHVGL TFMNINIYNF SIIVTSILLI GSLPILGVAI TGLLLDRNIN STIYDVIGDP VL YQHLFWF FGHPEVYVII LPVFGLTSLI LTSIIHKDIF GREGMMYCII SIGVVGYFVW AHHMFTVGLD IDSRSYFSIA TSI ISIPTS VKMFSYINTW ASGRGFRGNN SSWSFFSFLI CFCFGGFTGL LLSSGSLDIM LHDTYFVVGH FHTVLSLAAT FGLL IAHYF FLPIIFSYSI FESFSFYHTF LLLVGALLVF YPMHLAGLSG MARRVPEYAD IFIPFMTVGF HGTFLLIFST LTFIR SYFQ FLSHINHSNY L UniProtKB: Cytochrome c oxidase subunit 1 |
+Macromolecule #44: Cytochrome c oxidase subunit 2
| Macromolecule | Name: Cytochrome c oxidase subunit 2 / type: protein_or_peptide / ID: 44 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Euglena gracilis (euglena) Euglena gracilis (euglena) |
| Molecular weight | Theoretical: 22.443406 KDa |
| Sequence | String: MRYGNREIES IILFFDQTII LYTSIINALI VGIIIIIRKW YNRKGICNQY IYHIKIEVIW TILPIIFLIV IVLHSVTVIY NLEINKGTN NKYINVIGNQ WYWIYNNIES RISSLGRIIL VDQPLFIKAN NNTHLIISSL DVIHSFALPT LGIKVDAIPG R INNISING ...String: MRYGNREIES IILFFDQTII LYTSIINALI VGIIIIIRKW YNRKGICNQY IYHIKIEVIW TILPIIFLIV IVLHSVTVIY NLEINKGTN NKYINVIGNQ WYWIYNNIES RISSLGRIIL VDQPLFIKAN NNTHLIISSL DVIHSFALPT LGIKVDAIPG R INNISING LTQGLYVGYC SELCGSGHAF MPINLIVY UniProtKB: Cytochrome c oxidase subunit 2 |
+Macromolecule #45: Putative NADH dehydrogenase subunit 6
| Macromolecule | Name: Putative NADH dehydrogenase subunit 6 / type: protein_or_peptide / ID: 45 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Euglena gracilis (euglena) Euglena gracilis (euglena) |
| Molecular weight | Theoretical: 19.414947 KDa |
| Sequence | String: MILNTNINYG GRNSRLCYLI PLMVHTIFLY LAYLIYYNIN LINIYLLLII IIFINIEWPL VIIYEYIYLT NYNIQNNLLY NIATFIFLE ILLFIGFYWL YINNIIHYPN NLPYNNNIAV NLLDHLNHNN NTDICLYIIL HNLLLLLYVT LILQLVHRYV Q L UniProtKB: Putative NADH dehydrogenase subunit 6 |
+Macromolecule #46: NDUFC2
| Macromolecule | Name: NDUFC2 / type: protein_or_peptide / ID: 46 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Euglena gracilis (euglena) Euglena gracilis (euglena) |
| Molecular weight | Theoretical: 22.010127 KDa |
| Sequence | String: MDRSSVTPRQ VFLEPVELPF AKKDASLEIF TESQLKSLLE PADASLFKWK NLAKWKKEVR TLLPQEFQSW DPISNQETIV DLNPYLVGG IFGGFNAMAH NYLSCTVWWK VPWRFPMWMA AWAGVSAYLF EFRREVEYEN RKSRLLVNNY YNRVRTVIVE R EKRKNGPD HQFKDLPWMD REEQRSG |
+Macromolecule #47: COX4
| Macromolecule | Name: COX4 / type: protein_or_peptide / ID: 47 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Euglena gracilis (euglena) Euglena gracilis (euglena) |
| Molecular weight | Theoretical: 20.42748 KDa |
| Sequence | String: MGGDAHAHGG HGLHADPIFN GSKVVRSIQM MRSYHRLPVG PEPPHLKIRG APLHPWSMYT DKGGFFFGVG TQLPRNFFPK FLATTSAIM GVTYGLVWLY NAAGPRAKTQ TRKWKELESE DPRFPFQEIP EIYLPSDIDR DPNADCWRIL NQVPRKEKLL V EITDPLRN FDFKVAKIKE N |
+Macromolecule #48: NDUEG1
| Macromolecule | Name: NDUEG1 / type: protein_or_peptide / ID: 48 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Euglena gracilis (euglena) Euglena gracilis (euglena) |
| Molecular weight | Theoretical: 53.351969 KDa |
| Sequence | String: MRRVGVYALR CARGHASSAL AAAPASETLT VGSLYSKKLR EYPHKVLWRR KTGFENSSWN QCRRSHEALA TNFSEIEVGD RALFITDCD SLMVNVALAI QKRGAQVCIV PAQGLTSNKL ETYIEQLRPK VIFLGKDKVS VPDPLGGDEA PQKMTLDLYS M IWKIFPIS ...String: MRRVGVYALR CARGHASSAL AAAPASETLT VGSLYSKKLR EYPHKVLWRR KTGFENSSWN QCRRSHEALA TNFSEIEVGD RALFITDCD SLMVNVALAI QKRGAQVCIV PAQGLTSNKL ETYIEQLRPK VIFLGKDKVS VPDPLGGDEA PQKMTLDLYS M IWKIFPIS NVVEGLPLQS KRYPFVKSVV LCSQVDAINQ HDMITNIKYF ETPRDQDYYE SPLVHIAMHI APDFPALSVV DS AGKWTTH SHTTLLNAGR LFAAKYFGKA EHVLLLPGSE STPGGILTLY AALHAHAVLC YGEDQLVTKG DCKRVTRAVH QHE ATAVVG TKAQWDFLLK YGDAATKKED ATKEWFKSLK WAAVFLNDVE DAGDLASKIR SSFGIDRVEC IRGSPETLNL TSDG SGKLF DSLSVVIKDK DGVATRGDAE GFVWIKGPHV SAQYWNHIGL MSAPKDPNGY VKLPYVGKKK PDGGFTITTV QPVRE LVL |
+Macromolecule #49: NDUEG2
| Macromolecule | Name: NDUEG2 / type: protein_or_peptide / ID: 49 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Euglena gracilis (euglena) Euglena gracilis (euglena) |
| Molecular weight | Theoretical: 51.015891 KDa |
| Sequence | String: MVIGTFFKTG FEKGLPLHEQ VVRHLLPLVP KARKGFWPYY FAVNERVVLP RRAGAALNSR LRIPGKNRRE CLPTSASSPL ELAQLRKAT DKPVEDVKPQ VFVSTSSPSD AVPLHNESVH SKWLEALDEV NKTASTFSDA FEIQNESLSK EIFHRLAVPA S LKAGNIFA ...String: MVIGTFFKTG FEKGLPLHEQ VVRHLLPLVP KARKGFWPYY FAVNERVVLP RRAGAALNSR LRIPGKNRRE CLPTSASSPL ELAQLRKAT DKPVEDVKPQ VFVSTSSPSD AVPLHNESVH SKWLEALDEV NKTASTFSDA FEIQNESLSK EIFHRLAVPA S LKAGNIFA HDGAFGSNSA DDIKFTAVTH DPTAALFLRH MVNPVPQVDP VDFPNLFSVF HIHDYEFTDP RIVEEFDGVK KE QLGITSP RFVLYDLAER NVYVSGSSQD LRDAIVCLGG LVAFHLYGSL TLACNSFIDK DGKLTLVFGS EANLNSPQLF GAH HSLWTP NGVSRAWNGV TVEGAKAQFA SDLVEVTAKG PRLTAPLPLQ LGGTARPRGA NLLAGAAAGT PEPPLAVDPK LPWR PNVVS AAGAKFVFVG KEEAKLSVDD AAALFADSHA AYPLGFSTKK KLAAKFKELA ATAPGASFVT TPNVSSL |
+Macromolecule #50: NDUEG3
| Macromolecule | Name: NDUEG3 / type: protein_or_peptide / ID: 50 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Euglena gracilis (euglena) Euglena gracilis (euglena) |
| Molecular weight | Theoretical: 63.09698 KDa |
| Sequence | String: SRNVLTKGWR FHAHAGTFQS ALRFEEFQLK KSVGDVVLRL QCAPVTGFDL DQVRGLQGKV PLPAVGGLSG VGVVTEGSGI FKEGDRAVL LGANGAWSQY AVSSANHLLS VPATIPVEYA SLLASGPFAA YRILKAAHLK AGDLVLVNGA HTAIGLAALQ I MSRNVLTK ...String: SRNVLTKGWR FHAHAGTFQS ALRFEEFQLK KSVGDVVLRL QCAPVTGFDL DQVRGLQGKV PLPAVGGLSG VGVVTEGSGI FKEGDRAVL LGANGAWSQY AVSSANHLLS VPATIPVEYA SLLASGPFAA YRILKAAHLK AGDLVLVNGA HTAIGLAALQ I MSRNVLTK GWRFHAHAGT FQSALRFEEF QLKKSVGDVV LRLQCAPVTG FDLDQVRGLQ GKVPLPAVGG LSGVGVVTEG SG IFKEGDR AVLLGANGAW SQYAVSSANH LLSVPATIPV EYASLLASGP FAAYRILKAA HLKAGDLVLV NGAHTAIGLA ALQ IAKAWG IDAVGVAHGA PALQVEKLKQ MGLNVVSSFA LDPKQVFGTS QPKFAISLVG GNAAAYVTHL IGSDGHIITC PLAS DEPHI LPNVDLVNKN LTIQTFSPWK SLLSATATEN EQMVSELCDL IAAHKLKANA VVRHEFGNLL DAIREAEHGT HNAVI LHEG TEKTWDNKNH DIYMEIDDKL QANWDAAAAA QDPYLKTGRD QPWQVLAEAE EVALPDELRV KLAAVTTEAE LLAVLD TLT LKERHLLGLP ATQAITVSAE ELKKMVSEFA S |
+Macromolecule #51: NDUEG4
| Macromolecule | Name: NDUEG4 / type: protein_or_peptide / ID: 51 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Euglena gracilis (euglena) Euglena gracilis (euglena) |
| Molecular weight | Theoretical: 41.313383 KDa |
| Sequence | String: MQSLKRAGTL SRPFLRCFYD AGAPQIFRSN VPGRPLPWRQ ERQVPPNPSQ SKWQWEPEHI PTAEEYEAFP EVITLYGGDG LLRSSVIQE LVQSPRVSTI RVGTPWPDEF ASKLPGEWQS KVVAEFVDIL DRHSVLAAAE GSQALVNMMD IPYECELTYY Q AHVGSAQM ...String: MQSLKRAGTL SRPFLRCFYD AGAPQIFRSN VPGRPLPWRQ ERQVPPNPSQ SKWQWEPEHI PTAEEYEAFP EVITLYGGDG LLRSSVIQE LVQSPRVSTI RVGTPWPDEF ASKLPGEWQS KVVAEFVDIL DRHSVLAAAE GSQALVNMMD IPYECELTYY Q AHVGSAQM ISHAANTCMC SRVIHVSSLA SRVDSWSRYS ESKFRGEDMS LACFPWTTIL RFGPLVGKNS PALKQFASYM KY APIYPCV AKDTKIQPTF VGDAAKAILA ALGNPSTRQL QFDLGGPEVF KHADFIKEVM RLTKASRPVV PVPGVIGDSI VAL LQWLPD PLVTRDMVYL IRSHHIANHD SMRTWKDLLP EHKLKTMAEA LQ |
+Macromolecule #52: NDUEG5
| Macromolecule | Name: NDUEG5 / type: protein_or_peptide / ID: 52 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Euglena gracilis (euglena) Euglena gracilis (euglena) |
| Molecular weight | Theoretical: 29.182729 KDa |
| Sequence | String: MALAKGWRFS AHGGTWKAVL KLEDFPLTKT AGAAVLKVQA APVTPRDLDR IRGLYGALPL PAVAGTSGVG IVTQAGGAFK EGDRAVLAA ANPAGSYATL AAVDPAHLIK VPAALPVDVA ATLAVGPFAA YQILKLSGLK SGDSLALDGE ATLLGKSVAL L AKSRGITV ...String: MALAKGWRFS AHGGTWKAVL KLEDFPLTKT AGAAVLKVQA APVTPRDLDR IRGLYGALPL PAVAGTSGVG IVTQAGGAFK EGDRAVLAA ANPAGSYATL AAVDPAHLIK VPAALPVDVA ATLAVGPFAA YQILKLSGLK SGDSLALDGE ATLLGKSVAL L AKSRGITV VSGAAAGKDI KFALSLQGGR SASSLLGALG HGGQLLLHVA PSDEATVLDG ALVADKSVTI RSFAPAAVSA KE AEAMVEE VVELVKGNAL GLKVVRHDLA KLLEAVEEVT AGPSDTVHIL TLS |
+Macromolecule #53: NDUEG6
| Macromolecule | Name: NDUEG6 / type: protein_or_peptide / ID: 53 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Euglena gracilis (euglena) Euglena gracilis (euglena) |
| Molecular weight | Theoretical: 43.669457 KDa |
| Sequence | String: MWRTSGCALR SRASFTTAVT AHASGVKAID PFKILSLPDS ATRDDLRNQF FELAKSNHPD VGGDKAKFQA IQDAYEDAIR IADQKHPVA PWDGISPMTY AQAWQGKDYW RKLWEEHWAA RLAHMYKHNA ELTTLEANKK WREAQYMQVK DWMVLAKDVL D PKTKAEWQ ...String: MWRTSGCALR SRASFTTAVT AHASGVKAID PFKILSLPDS ATRDDLRNQF FELAKSNHPD VGGDKAKFQA IQDAYEDAIR IADQKHPVA PWDGISPMTY AQAWQGKDYW RKLWEEHWAA RLAHMYKHNA ELTTLEANKK WREAQYMQVK DWMVLAKDVL D PKTKAEWQ AGCELARDML LWTQANKKNY RRYFLSNQNV AVNMRQVYDE HEYWRQYENV QWAQWDAFFA RASAWALEHE EQ IRSVNST EGPLAAKFDY LFHGRLQYSS MSLEERLSRR AQEEKAYTRQ YWIAELMKAM RFSFRWVERF SRAFFPVLIL VVI AGYITD FQLIIRWLNI TRSETGALEV HNRKMDMVDW LLAGTPTPQN IEGTI |
+Macromolecule #54: NDUEG7
| Macromolecule | Name: NDUEG7 / type: protein_or_peptide / ID: 54 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Euglena gracilis (euglena) Euglena gracilis (euglena) |
| Molecular weight | Theoretical: 26.818773 KDa |
| Sequence | String: QKLLPKISCK YLALFLGLAG ATTALGFGLA PPTELVQGKI KSAELQVGDL LTYKDGGHAK CHIKEHWAVY LGTGEEIAQL VGQDAAALK LQPEERYIGH RLANTHHEEP VVRIDPLSTA RPPQGGALYK AEAPALGEPE DREAIVKGIL ARRAEKDGSL A AKNCQHFS ...String: QKLLPKISCK YLALFLGLAG ATTALGFGLA PPTELVQGKI KSAELQVGDL LTYKDGGHAK CHIKEHWAVY LGTGEEIAQL VGQDAAALK LQPEERYIGH RLANTHHEEP VVRIDPLSTA RPPQGGALYK AEAPALGEPE DREAIVKGIL ARRAEKDGSL A AKNCQHFS TWARYQYPYS NDSLETSSRF IVTRAGATAA AVFLGTLAVS QAPLFSTMFG AMGWMVGIFR PICNRQKVVT LQ KWEVEQ |
+Macromolecule #55: NDUEG8
| Macromolecule | Name: NDUEG8 / type: protein_or_peptide / ID: 55 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Euglena gracilis (euglena) Euglena gracilis (euglena) |
| Molecular weight | Theoretical: 23.994727 KDa |
| Sequence | String: MTTHRPPLVT DRMFPWTWNS RYYMMTPLAT YGREDIFANP HLPADKKLHA AAVEGGEFMK DAYAPPHKAV SLVRRVGRKA MVGAGLYFH LFLMDKFVSK GLLDRNTSPY FYMTLYEKAI YCLLPNKNKP NEEDLLKLFR EARLESNYKS VSMWTYRLRS E RVPKSYFD ...String: MTTHRPPLVT DRMFPWTWNS RYYMMTPLAT YGREDIFANP HLPADKKLHA AAVEGGEFMK DAYAPPHKAV SLVRRVGRKA MVGAGLYFH LFLMDKFVSK GLLDRNTSPY FYMTLYEKAI YCLLPNKNKP NEEDLLKLFR EARLESNYKS VSMWTYRLRS E RVPKSYFD KPPVSYDADR RTSGFVEWAF WMVGLESFGK SVVPEAW |
+Macromolecule #56: NDUEG9
| Macromolecule | Name: NDUEG9 / type: protein_or_peptide / ID: 56 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Euglena gracilis (euglena) Euglena gracilis (euglena) |
| Molecular weight | Theoretical: 18.272891 KDa |
| Sequence | String: MSATFWKLTF GIPLSGDYDP TDIPNAEAEL LTHVIIKTIQ LCTVLGAVVA APVAAAAQPA ARTLAGLGAT AARYGKAGAL IGVPLGAFM HYARLQGITG IDPIYDRAYR LRHNANQHRA DRSFLRGAAL GAAVGYVTPI GVAGGYTAFS IAGLFLSGLY N TQLEAAAK AAEAAAAAAE |
+Macromolecule #57: NDUEG10
| Macromolecule | Name: NDUEG10 / type: protein_or_peptide / ID: 57 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Euglena gracilis (euglena) Euglena gracilis (euglena) |
| Molecular weight | Theoretical: 14.140555 KDa |
| Sequence | String: MSKQNHGLHN RLIPIFVKEN AFNGFPLVNK GKWGSWLWFR NTQLKNLYHP LGAVLKWPRS FLMLNGTETA FWMVFFWAAA NTWLYSRQR KEFIKDCDYG (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK) (UNK)(UNK)(UNK)(UNK) ...String: MSKQNHGLHN RLIPIFVKEN AFNGFPLVNK GKWGSWLWFR NTQLKNLYHP LGAVLKWPRS FLMLNGTETA FWMVFFWAAA NTWLYSRQR KEFIKDCDYG (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) |
+Macromolecule #58: NDUEG11
| Macromolecule | Name: NDUEG11 / type: protein_or_peptide / ID: 58 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Euglena gracilis (euglena) Euglena gracilis (euglena) |
| Molecular weight | Theoretical: 11.472202 KDa |
| Sequence | String: (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)SL QCHDTRNRRD KCMVQHGFCA QQI EDHFEC LEQYYQKEKV ...String: (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)SL QCHDTRNRRD KCMVQHGFCA QQI EDHFEC LEQYYQKEKV DQLRREFRSQ SFQMVKQMKL EDLGHFDRYG NLR |
+Macromolecule #59: NDUEG11
| Macromolecule | Name: NDUEG11 / type: protein_or_peptide / ID: 59 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Euglena gracilis (euglena) Euglena gracilis (euglena) |
| Molecular weight | Theoretical: 11.323098 KDa |
| Sequence | String: MHRIATRVAM QQRFYGVARP DVQNRVIQTV QPFVDICCRP RTEVKPTSHF VLDLNLPAVH LKQVVLALEG EFAAPVPDAQ FKNIVSVAD AVEYFAGHPQ AK |
+Macromolecule #60: NDUEG13
| Macromolecule | Name: NDUEG13 / type: protein_or_peptide / ID: 60 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Euglena gracilis (euglena) Euglena gracilis (euglena) |
| Molecular weight | Theoretical: 17.731408 KDa |
| Sequence | String: MAHDKPHNEP PKRLLSKQEK TDMAVRENIE KPSWIKENGF QRWCYEMVRG VGRHPWPRYF ELTDFFGDKR HGDAIPRSWA AVKKLERGQ VIGPAVCVTL TLWVFVAGTI SKRNNTVWCD SIQEKYGFKQ PYSREFRTQH DGVLKPKLPA IY |
+Macromolecule #61: NDUFX
| Macromolecule | Name: NDUFX / type: protein_or_peptide / ID: 61 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Euglena gracilis (euglena) Euglena gracilis (euglena) |
| Molecular weight | Theoretical: 37.196953 KDa |
| Sequence | String: MTTIHAYTAT QKTVDGPSKK DWRGGRAAAI TMVFRPLFRS FVNPFSLARL MAFRPTSVLA LRTCQLRLGG GHGHGHGHGH GHGGHGGKP NKFENHIYRD ASGQPWFDLH KWELGRKPGK EDLEHPGRHE SFSQTKRLVS LTFIDKDGNP HNIKCEPGMN M ADAALQAG ...String: MTTIHAYTAT QKTVDGPSKK DWRGGRAAAI TMVFRPLFRS FVNPFSLARL MAFRPTSVLA LRTCQLRLGG GHGHGHGHGH GHGGHGGKP NKFENHIYRD ASGQPWFDLH KWELGRKPGK EDLEHPGRHE SFSQTKRLVS LTFIDKDGNP HNIKCEPGMN M ADAALQAG VSLDWLDGEG NQGFSAWSHM VISSPQFEML VPPDHREDKY LWDLEQWGTS HRNSRFVKFL FVQPWMEGMV VT HPFFADV VTPNFTDPFT GKIEDKDQAV APTYFVSKVV DPEKPDYKFP NPFELLWAGI DTYEQLWDYT RQSWLKRLEA RRQ KQQQA |
+Macromolecule #62: NDUCA1
| Macromolecule | Name: NDUCA1 / type: protein_or_peptide / ID: 62 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Euglena gracilis (euglena) Euglena gracilis (euglena) |
| Molecular weight | Theoretical: 48.726906 KDa |
| Sequence | String: MAKLTWLKNL LNPYSRVPPS TFLRFFPRLL NKFGSAERDI NAEFPGTVHK HIKTYQERFM EQGAGDRIAT KWNPKPWEKA YMGQPDHPM TKAEQAKKED FMVGIHWDRS AGGRWTPNDK FPLFDYEFPI HPGRIILRWL YKQGKEPVNM QRSILVTDDF A TPSVYPFG ...String: MAKLTWLKNL LNPYSRVPPS TFLRFFPRLL NKFGSAERDI NAEFPGTVHK HIKTYQERFM EQGAGDRIAT KWNPKPWEKA YMGQPDHPM TKAEQAKKED FMVGIHWDRS AGGRWTPNDK FPLFDYEFPI HPGRIILRWL YKQGKEPVNM QRSILVTDDF A TPSVYPFG WHAPSAILIG DACISNDAAV FDHCVLRADR AAIWVGPKSH VLEGCTLTTA PPTPDRPALG SVLIGENTVV GA GSSLNAC WIGDHCIIGS GCTIGFGARI DDGAVVGAGS VVEDDQYIPA GEVWVGRPAR YLRKTGDVDT FTAVAENDTL RSL HLAYSE YETTHGNVWA ESDKVCDNLE EEVAHRLQAH DVARAMVSKN FDAKLLKLPK SLVADLMDIV SDDDHPNPKP TVSA QARQH FSSQWDFNRK QEQRPVFTGN YNSPTMSRDM A |
+Macromolecule #63: NDUCA2
| Macromolecule | Name: NDUCA2 / type: protein_or_peptide / ID: 63 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Euglena gracilis (euglena) Euglena gracilis (euglena) |
| Molecular weight | Theoretical: 28.857973 KDa |
| Sequence | String: MPLPQKSTLV RLGAAIRECG QALDRWGSFL QGRYGHLEKL QRTRRINGFH NFFPEVKGVR FIAPSASVIG QVTVSPGSSI WYNSVVRGD RGKVTIGEDT HILERVVIRS GILSVRDVKI GKDVIIEPGA IISPCQIEDG AYIGANAVLM EGCKIGKGVV V GPGAVVTE ...String: MPLPQKSTLV RLGAAIRECG QALDRWGSFL QGRYGHLEKL QRTRRINGFH NFFPEVKGVR FIAPSASVIG QVTVSPGSSI WYNSVVRGD RGKVTIGEDT HILERVVIRS GILSVRDVKI GKDVIIEPGA IISPCQIEDG AYIGANAVLM EGCKIGKGVV V GPGAVVTE FAELTQPGVY QGVPAKSATA LTTEAAEAIT TRRAEFAKLA EEHEEMNTKL IEKQTEERVI LKDILENQLN EG NEFTMRS HHVARAPNVS PGNIAAGSAA |
+Macromolecule #64: NDUCA3
| Macromolecule | Name: NDUCA3 / type: protein_or_peptide / ID: 64 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Euglena gracilis (euglena) Euglena gracilis (euglena) |
| Molecular weight | Theoretical: 28.0169 KDa |
| Sequence | String: (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)GAA IRECGQALDR WGSFLQGRYG HLEKLQRTRR INGFHNFFPE VK GVRFIAP SASVIGQVTV SPGSSIWYNS VVRGDRGKVT IGEDTHILER VVIRSGILSV RDVKIGKDVI IEPGAIISPC QIE DGAYIG ...String: (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)GAA IRECGQALDR WGSFLQGRYG HLEKLQRTRR INGFHNFFPE VK GVRFIAP SASVIGQVTV SPGSSIWYNS VVRGDRGKVT IGEDTHILER VVIRSGILSV RDVKIGKDVI IEPGAIISPC QIE DGAYIG ANAVLMEGCK IGKGVVVGPG AVVTEFAELT QPGVYQGVPA KSATALTTEA AEAITTRRAE FAKLAEEHEE MNTK LIEKQ TEERVILKDI LEDQLNEGNE FTMRSHHVAR APNVSPGNIA AGSA |
+Macromolecule #65: ND1
| Macromolecule | Name: ND1 / type: protein_or_peptide / ID: 65 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Euglena gracilis (euglena) Euglena gracilis (euglena) |
| Molecular weight | Theoretical: 79.682836 KDa |
| Sequence | String: MLNSNIYIII YGGIIMYSIM IIIQMFLYNF SNKIYIEVEI NKYILSKNNI DIYWIICNCT IIIIITTLNH IINKIGIYNM IEYNICYWL IGTGLGLYIS PFIVFGYKFF VYIMDLNNYS LNIYHNNNKM NDIQQIYNGT NYNDTMIFFI KDINNIFTIY R SINFFMNW ...String: MLNSNIYIII YGGIIMYSIM IIIQMFLYNF SNKIYIEVEI NKYILSKNNI DIYWIICNCT IIIIITTLNH IINKIGIYNM IEYNICYWL IGTGLGLYIS PFIVFGYKFF VYIMDLNNYS LNIYHNNNKM NDIQQIYNGT NYNDTMIFFI KDINNIFTIY R SINFFMNW LYQMIYYGVR MWLVFVLHSF SLGSFGELIT VITDNNLIFN VFYIGLLGLG VILYLIVIFY LGIQIYVYIS FS LSFLHST ILLFLVNYIP HYNNKSIFNT FTNKSIYLIP VSLVDLININ IIFYILLLYT LLLFFIPLFL ASINYTYHYI YKY YNYNYN FINNNTNTNN NNKFINISYN FFIILIILNI INISSIYNIL QILLIMLIVL SLSSLLTVLE RKGLASSQRR IGPS YNGWF GLVQIVQDGI KLIYKDYNRY NNINNKYIMI SCILNFIYSY LLFIFIYIDL ILYINISYII FMIIIILMIN HITII ICGI VINNSKWTIL SSIRLILLYF MYDIIFLLIL LYLSPINNLG INLLYNNNNL NLNNYIESQF YYINLYKYPL LLYIYI FIV LIEAGRIPVD LIESESELIS GYSIEYSGFL YALFASAEYS IILFHSILLS LLFFSYYSFN ILFIHITILF FIFVIIR ST LPRFKYTNLF NLTYYYILPF ILTYLLLL |
+Macromolecule #66: ND2a
| Macromolecule | Name: ND2a / type: protein_or_peptide / ID: 66 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Euglena gracilis (euglena) Euglena gracilis (euglena) |
| Molecular weight | Theoretical: 35.864434 KDa |
| Sequence | String: MLYNNNLIII ILMISIIIGI SLQNILVNDI SELRWINRFN LDNFIIIYII LLYNNIILIL GIISLIISTN KNTTNNKVQL IHMIIIIIN TIYICNNNNN TIINIILMII TIDILSVLNI ILIQKGEGIW YYFLYQSLMT ILIWWVLILD LSSLLSFFYY Y KLGSGIGG ...String: MLYNNNLIII ILMISIIIGI SLQNILVNDI SELRWINRFN LDNFIIIYII LLYNNIILIL GIISLIISTN KNTTNNKVQL IHMIIIIIN TIYICNNNNN TIINIILMII TIDILSVLNI ILIQKGEGIW YYFLYQSLMT ILIWWVLILD LSSLLSFFYY Y KLGSGIGG YYIPSLYSSI IYYNINLMIY IGTTNIILMY NPIFLFNNFN HNYFLIISNF LFILYILYIW IFNGYLFINL WL YSISFST IILANIYYLF TSIDFIYYNL FYYIYYFTIS SIIIWFIFIL SLYFINNYNN HIN |
+Macromolecule #67: ND3
| Macromolecule | Name: ND3 / type: protein_or_peptide / ID: 67 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Euglena gracilis (euglena) Euglena gracilis (euglena) |
| Molecular weight | Theoretical: 34.504223 KDa |
| Sequence | String: MINNIIDIYI FNNNIINKSL INIIDIIILN IGILLILVEG SIRGIIGLII VGISINIYLI NNGYILISLL LIISIISISI IISLIKYNT MSIYKFNNNK YIQLFFILYL YIILINNNNY NNNINNIEII GYILIFESNI LLSLLILIMF MFIIILVKFN N KISINYNI ...String: MINNIIDIYI FNNNIINKSL INIIDIIILN IGILLILVEG SIRGIIGLII VGISINIYLI NNGYILISLL LIISIISISI IISLIKYNT MSIYKFNNNK YIQLFFILYL YIILINNNNY NNNINNIEII GYILIFESNI LLSLLILIMF MFIIILVKFN N KISINYNI IYFNVINYII LNIYILFIIA LLICYIPIII RYKSKIYKWS SDIYEVGIGV NMSNSIYNSI SNYNIGNYFI YF YIYLFIF LDIIINIIIL IYFININLFT YYFTLFFLIL AISYSLLFSY NFSNNF |
+Macromolecule #68: NADH-ubiquinone oxidoreductase chain 4
| Macromolecule | Name: NADH-ubiquinone oxidoreductase chain 4 / type: protein_or_peptide / ID: 68 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Euglena gracilis (euglena) Euglena gracilis (euglena) |
| Molecular weight | Theoretical: 56.417312 KDa |
| Sequence | String: MIYKIIILRY ISILIIMIIQ LMKMYNKEYR LISIISIISI LSIMNWPQSI KEINIYIINL IIGYEINLSY NYIGIYMSLI LDILIWISI LYERYYIKNN NTYNIIIYYI INNIFFSSND IITISLIYEF QTIPLLLIIN NSYNKINIYN KKGIGISNIL L ILYSIISG ...String: MIYKIIILRY ISILIIMIIQ LMKMYNKEYR LISIISIISI LSIMNWPQSI KEINIYIINL IIGYEINLSY NYIGIYMSLI LDILIWISI LYERYYIKNN NTYNIIIYYI INNIFFSSND IITISLIYEF QTIPLLLIIN NSYNKINIYN KKGIGISNIL L ILYSIISG LLLYNSMYNI YLLNHTSNIS HNINILNYSY NHILINYILI NILISGSMKL SIFPFHIWLG KVHVEAPTIG SI LLAGISL KTGFYLHYLF IYLYQYIYIN YLIYFIYILF IGIIINNINI FYQIDTKRWI ALYSIIHMNL YYIIMLILLI SNN NNLYYI IFINILIYGM IGHSLISGGL FLIIGYIYDI TNNKNLYLIN NNIISSYIYF ILLLLLLANS SFPLFVLFIF ELLA FTSLS IFNILLSIFL LILSCSNLLS SLYIYYKYFY THYYSPIITF TSFDFILIIL AIPILIIIIL MGFNLLLPIH FLI UniProtKB: NADH-ubiquinone oxidoreductase chain 4 |
+Macromolecule #69: ND5
| Macromolecule | Name: ND5 / type: protein_or_peptide / ID: 69 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Euglena gracilis (euglena) Euglena gracilis (euglena) |
| Molecular weight | Theoretical: 68.375359 KDa |
| Sequence | String: MLIISIIKWL IISIIMNLSG NIIGINGLIY IIMLILVLSP IISWYVLYNI IIINIYNNKL IYLLSYNFIN YYNYNIDLEI GISIYEMIT VILLINVSYM INIYILKYLY KDKNVIRFVC IIMLFTYNMI LLIISNDLIM LFIGWEMIGI ISLLLINYYN N RIEATKAG ...String: MLIISIIKWL IISIIMNLSG NIIGINGLIY IIMLILVLSP IISWYVLYNI IIINIYNNKL IYLLSYNFIN YYNYNIDLEI GISIYEMIT VILLINVSYM INIYILKYLY KDKNVIRFVC IIMLFTYNMI LLIISNDLIM LFIGWEMIGI ISLLLINYYN N RIEATKAG LKAVVYNRIG DVFLLLSIIL SINMYNSNSI LLYNILISYM YYNINYININ LIIGMSFIIC AWSKSTQLGF QP WLLDAME GPTPVSALLH SATLVTAGII LLYKNRYILY YNSSLAILLL ILGGISCLLN SFSSINYLDI KRIVAYSTCT HIS LMIMIL GIDILINISE ISLLHLFYHG WSKSLIFMLC GYMISIIHSQ DLRFFGNLFQ HIPILFVIIN ISLLTILGFP GSYL SYSKD IILEFGLISI YGYNIILLFI IIILLSQGYS LGILLYLIYN YSYYNSTHNI YNFYSNKNNY IYIFAFLYLI IIIIY LPFL LYDILIYNNI SIMHHISYID PFSLIAFLGF ILSYYNYNYN HTLYIFNIHN NRLYIDKLLS SFMSIFSIHI IYYFQF ILE YGFIMHYLHI TNIIIFLIFL I |
+Macromolecule #70: MPP-beta
| Macromolecule | Name: MPP-beta / type: protein_or_peptide / ID: 70 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Euglena gracilis (euglena) Euglena gracilis (euglena) |
| Molecular weight | Theoretical: 53.520293 KDa |
| Sequence | String: MTTPSLSSIL RHTRPIFKET LRAARPTLQN ALPNGFRIAS ESKDGDTCTV GVWIDAGSRW ETEKNNGVAH FLEHMNFKGT GKRSRQDIE FGMEKMGAHL NAYTSREHTC YYVKCFKKDV PEAVDILADI LLNSKRTEQD LDAERQTIVQ EKEDVEARID E VLMDHLHS ...String: MTTPSLSSIL RHTRPIFKET LRAARPTLQN ALPNGFRIAS ESKDGDTCTV GVWIDAGSRW ETEKNNGVAH FLEHMNFKGT GKRSRQDIE FGMEKMGAHL NAYTSREHTC YYVKCFKKDV PEAVDILADI LLNSKRTEQD LDAERQTIVQ EKEDVEARID E VLMDHLHS AAFEGSGLGL SILGPLENIQ KSITKGMIDD FVKTHYTGPR MALVGSGAVD HGQLCDLASK YFGALPTGQP KP SGFTRFL GGDKRETNQL NPLTHVAVAF QTPGISHPDA IKIKVLEQLL GSYSRDKGEA AYSCFARAIV MDFYDPKVGQ FFR PNKAGH NPIHSLNAFW APYSDVGLLG FYAIAEPGKS YGHEWENILH YAMRELIRVS RNISEEEFER AKNQLKLQTM LQLD GTTNI ADDIGRQVLS FGARVPLASF FEQLDAISRE DLIRVAHEYF YDKDPVVAVI GDTDNVPEYD ALRAVTYSVD LGRA |
+Macromolecule #71: Ubiquinol-cytochrome-c reductase complex core protein 2, mitochondrial
| Macromolecule | Name: Ubiquinol-cytochrome-c reductase complex core protein 2, mitochondrial type: protein_or_peptide / ID: 71 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Euglena gracilis (euglena) Euglena gracilis (euglena) |
| Molecular weight | Theoretical: 51.13266 KDa |
| Sequence | String: MKSVVRSKGT QALFRRFSSA LGDSINPNQV GVGDNVIRVN GRLFEVDKVQ EKGLKTSVLD NGTKVITLDN GGSVAQLTFL YKDGPVYEN IFNAGISSFM KHALTKDGLT SSEYITKTFL QKAGIIVHEP TVVNKSAIAF TVEGFRDTLA QPAVADKFWQ S LLFPRFSP ...String: MKSVVRSKGT QALFRRFSSA LGDSINPNQV GVGDNVIRVN GRLFEVDKVQ EKGLKTSVLD NGTKVITLDN GGSVAQLTFL YKDGPVYEN IFNAGISSFM KHALTKDGLT SSEYITKTFL QKAGIIVHEP TVVNKSAIAF TVEGFRDTLA QPAVADKFWQ S LLFPRFSP ENVKEVKRLV ELESKETKRD SPFAYLQDIL HKTAFKGSPL GHTSFVPAYN LGYIDSNKLF DRWDAHYGFG NI AVIATNI EHEAVLAAIT DSAWVARAHN KVGGVAAPAS KYSGGEGYDV VHRAKEFDDQ FTDVYSTYTA YAFKAPGRSN LKE HAASLV IAQALSNAVS PVLNTSFAPK RLEVFYQAYD TVGLIGLSSV QASNAQLKAF KAALSKIGTL SEADLAVHKS AALL TAYGN VESWRATQAT LIDSFNTTGQ PLSPLEIVSA IKAVSADTVK SVVATMLGSP ATLVHHGDSP CAPTLDALQ UniProtKB: Ubiquinol-cytochrome-c reductase complex core protein 2, mitochondrial |
+Macromolecule #72: Cytochrome b
| Macromolecule | Name: Cytochrome b / type: protein_or_peptide / ID: 72 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Euglena gracilis (euglena) Euglena gracilis (euglena) |
| Molecular weight | Theoretical: 42.476266 KDa |
| Sequence | String: MNKISLYRSY STTILLSPAY SLGFCASIFI VIQIISGYIL ASNYIASTNE SFNIIHNVIM RELDTGWLIR FNHINGCAFL FIVIYMHIY RSLYHNSITK TSVWIVGIIM YILICGIAFT GYSLVYGQMS LWAIVVICSL VTAIPFIGNK LLILIWGGNI V SSVTLQRI ...String: MNKISLYRSY STTILLSPAY SLGFCASIFI VIQIISGYIL ASNYIASTNE SFNIIHNVIM RELDTGWLIR FNHINGCAFL FIVIYMHIY RSLYHNSITK TSVWIVGIIM YILICGIAFT GYSLVYGQMS LWAIVVICSL VTAIPFIGNK LLILIWGGNI V SSVTLQRI FCIHYLLPLL LILFIIIHLY NLHNVNSTGD NYFINNRYDR INFYPLLLIR DVFIGSNILI IYNIFVYYYS DL FGHPDNY VPANPLVTPS EIMPEFYLLP FYALIRAIPH KVLGIIIMVL FLLSLTNLYP IYFIRFYNNI NILQRSLLLL LLL DLVIAS KLCLLINHYE SFYLLLILSI LCVLSHHIYN TSFNFSNSIS NL UniProtKB: Cytochrome b |
+Macromolecule #73: Cytochrome c1, heme protein
| Macromolecule | Name: Cytochrome c1, heme protein / type: protein_or_peptide / ID: 73 / Number of copies: 2 / Enantiomer: LEVO / EC number: quinol-cytochrome-c reductase |
|---|---|
| Source (natural) | Organism:  Euglena gracilis (euglena) Euglena gracilis (euglena) |
| Molecular weight | Theoretical: 27.889436 KDa |
| Sequence | String: GVDSHPPALP WPHFQWFQGL DWRSVRRGKE VYEQVFAPCH SLSFIKYRHF EAFMSKEEVK NMAASFEVDD DPDEKGEARK RPGKRFDTV VQPYKNEQEA RYANNGALPP DLSVITNARH GGVDYIYALL TGYGRPVPGG VQLSTTQWYN PYFHGGIIGM P PPLTDDMI ...String: GVDSHPPALP WPHFQWFQGL DWRSVRRGKE VYEQVFAPCH SLSFIKYRHF EAFMSKEEVK NMAASFEVDD DPDEKGEARK RPGKRFDTV VQPYKNEQEA RYANNGALPP DLSVITNARH GGVDYIYALL TGYGRPVPGG VQLSTTQWYN PYFHGGIIGM P PPLTDDMI EYEDGTPASV PQMAKDVTCF LEWCSNPWWD ERKLLGYKTI ATLAVIAVSS GYYNRFLSGL WRSRRLAFRP FN YSK UniProtKB: Cytochrome c1, heme protein |
+Macromolecule #74: UQCRFS1
| Macromolecule | Name: UQCRFS1 / type: protein_or_peptide / ID: 74 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Euglena gracilis (euglena) Euglena gracilis (euglena) |
| Molecular weight | Theoretical: 27.825922 KDa |
| Sequence | String: MFKQLSRMYS TSPAVANAVG GLATSQIRDL VGNPAKSGKP VVKMPPYLKP SQLRPSRGAY SDEFVRENYI KAADPEDPYN YRSRAFTYA AKIPMWAGLL AGTRVTVVYL MSQFMPSKSS LALANIEVDI GDIPEGKTVT IMWQGKPVFL RKRTDAEIED M RAVPMDAL ...String: MFKQLSRMYS TSPAVANAVG GLATSQIRDL VGNPAKSGKP VVKMPPYLKP SQLRPSRGAY SDEFVRENYI KAADPEDPYN YRSRAFTYA AKIPMWAGLL AGTRVTVVYL MSQFMPSKSS LALANIEVDI GDIPEGKTVT IMWQGKPVFL RKRTDAEIED M RAVPMDAL KDPQTDEERL GEGRWAVFMA VCTHLGCVPV IDQGSYNAYF CPCHGSHYDH SGRIRKGPAP LNLEVPPFKL LD DSTLFIG NADAA |
+Macromolecule #75: UQCRH
| Macromolecule | Name: UQCRH / type: protein_or_peptide / ID: 75 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Euglena gracilis (euglena) Euglena gracilis (euglena) |
| Molecular weight | Theoretical: 8.207571 KDa |
| Sequence | String: GKMAAEQVSV SELKLKFEEE CAHHHCHDIV KKVEVCEESI KGKEGKNCLF QHARLWECVD HCASPKVFAL LK |
+Macromolecule #76: UQCRB
| Macromolecule | Name: UQCRB / type: protein_or_peptide / ID: 76 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Euglena gracilis (euglena) Euglena gracilis (euglena) |
| Molecular weight | Theoretical: 27.404115 KDa |
| Sequence | String: MNDAPILAKL GLSKVNTYYF QRRKHIFALP ITIPNFPLAA LGIFRSTDMF EERTLERYSR DEREKTLGKA ALLLKEAKEK GNYSELVRL DPTDPRHPYY FEHPWQSALK MDTVSLTPYQ QYLRWHCLTY RAMHEWDRQG LLYDDLMQPK ALATDPFLEE A ILRLPYNQ ...String: MNDAPILAKL GLSKVNTYYF QRRKHIFALP ITIPNFPLAA LGIFRSTDMF EERTLERYSR DEREKTLGKA ALLLKEAKEK GNYSELVRL DPTDPRHPYY FEHPWQSALK MDTVSLTPYQ QYLRWHCLTY RAMHEWDRQG LLYDDLMQPK ALATDPFLEE A ILRLPYNQ RVERERRLSR AYDLALRREY LPDEDCIHPE DDVAYLHPYY HMVVDEHREQ HENPVDVYSR |
+Macromolecule #77: UQCRQ
| Macromolecule | Name: UQCRQ / type: protein_or_peptide / ID: 77 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Euglena gracilis (euglena) Euglena gracilis (euglena) |
| Molecular weight | Theoretical: 9.948402 KDa |
| Sequence | String: EGHSAYVALE QRVNTRQKGI IQRHLSPYDV DIRGNGFRQI YRLGMQKIRY FGLIGTPCLL VAGAYRGIWV FCERLEDERY WSNAL |
+Macromolecule #78: UQCR9
| Macromolecule | Name: UQCR9 / type: protein_or_peptide / ID: 78 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Euglena gracilis (euglena) Euglena gracilis (euglena) |
| Molecular weight | Theoretical: 6.939355 KDa |
| Sequence | String: MLANFSRFFY KTFLQSNATL IPLTLVFTY(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK) ...String: MLANFSRFFY KTFLQSNATL IPLTLVFTY(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK) |
+Macromolecule #79: UQCR10
| Macromolecule | Name: UQCR10 / type: protein_or_peptide / ID: 79 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Euglena gracilis (euglena) Euglena gracilis (euglena) |
| Molecular weight | Theoretical: 17.577297 KDa |
| Sequence | String: MALEIKREWH VVPATKKFAD VPTEWPAKEH ERFALPYGPS EGRIVSFMRA SRIRNRLLFT LNPTGYTEDT YVTKEALRRQ ALAESKNLW YLPERPVVAD QLETIVNRST FFVVKGHKNK GLIGIAGARH NPLVWLPTLA LGGVWAWRWA SAGTI |
+Macromolecule #80: Ubiquinol-cytochrome-C reductase complex subunit IX, mitochondrial
| Macromolecule | Name: Ubiquinol-cytochrome-C reductase complex subunit IX, mitochondrial type: protein_or_peptide / ID: 80 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Euglena gracilis (euglena) Euglena gracilis (euglena) |
| Molecular weight | Theoretical: 11.300364 KDa |
| Sequence | String: MQTHVRRVAL QALRPCLRAG LMAPKFPVRF ATTAVSGELL TKTPYTRPGY AAQWTCLVVL FLKNQLLMRL FFAFVAYVVA MKVFGARFH VDHDEDATPA E UniProtKB: Ubiquinol-cytochrome-C reductase complex subunit IX, mitochondrial |
+Macromolecule #81: NDUFS2
| Macromolecule | Name: NDUFS2 / type: protein_or_peptide / ID: 81 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Euglena gracilis (euglena) Euglena gracilis (euglena) |
| Molecular weight | Theoretical: 45.251949 KDa |
| Sequence | String: MPARADARTM QHFTLNFGPA HPAAHGVLRL ILEMNHEYII RADPHIGFLH RGTEKLMEMK PVIQVTPYMD RLDYMSAMAN EHAFIMAVE KLCEIPVPLR GSYIRVIFLE ITRLLNHLLA VACAAADVGA LTPILWTFEE REKLMTFYEC VSGS(2MR)MH PN FFRPGGVASD ...String: MPARADARTM QHFTLNFGPA HPAAHGVLRL ILEMNHEYII RADPHIGFLH RGTEKLMEMK PVIQVTPYMD RLDYMSAMAN EHAFIMAVE KLCEIPVPLR GSYIRVIFLE ITRLLNHLLA VACAAADVGA LTPILWTFEE REKLMTFYEC VSGS(2MR)MH PN FFRPGGVASD LPYGFLDQLY EFINQFAARL DEVEDLLTQN NIWRERLKAG YMDLDRALSY GFSGPMLRAT GVAWDLRV T APYEVYPYLD VEVPYGVNSD SYDRYLIRMQ EMRNSIRIIH QCINDIVPGD YRLHDAKFAT IPFRDCKDSM ENLIAHFKY YSEGFRIPEG FVYCGTESPK GELGVYLQSN GDSLPYRVRI RAPGFYHLAA LPALAQDTMF PDLVTVIGTL DIVFGEVDR |
+Macromolecule #82: NDUFS3
| Macromolecule | Name: NDUFS3 / type: protein_or_peptide / ID: 82 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Euglena gracilis (euglena) Euglena gracilis (euglena) |
| Molecular weight | Theoretical: 32.466703 KDa |
| Sequence | String: MFRRVATHVV PKLSTLLRAR NSIKFVTTQE VPPVHQGREV KSTPQNKGKD EFVRYLSSVL YPMASEIMWD DVTNDVTINV YPQYIRPVA KMLRDHQQFQ YKVLVDVTCV DTMTNLKRFD VVYNFLSVVF RHRVMVRASV DDASGIDSLT PLFHSAMWAE R EAWDMFGV ...String: MFRRVATHVV PKLSTLLRAR NSIKFVTTQE VPPVHQGREV KSTPQNKGKD EFVRYLSSVL YPMASEIMWD DVTNDVTINV YPQYIRPVA KMLRDHQQFQ YKVLVDVTCV DTMTNLKRFD VVYNFLSVVF RHRVMVRASV DDASGIDSLT PLFHSAMWAE R EAWDMFGV FFVGHPDLRR MLSDYGFEGH PLRKDFPLGG YTEVYWDAPN STIKYASPPI FREEYRDYEN IETEWDDLYH RC NTAWAYD RMVKQGMESC DADYTYDDPA AHKKPQQTQG |
+Macromolecule #83: NDUFS4
| Macromolecule | Name: NDUFS4 / type: protein_or_peptide / ID: 83 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Euglena gracilis (euglena) Euglena gracilis (euglena) |
| Molecular weight | Theoretical: 23.736867 KDa |
| Sequence | String: AGLAARPLHT SAPRLIFPFR RPFKSEPEDE TKIPDCYVNG KPIADGQGYQ AIGLAEHFER IRRNVNIRRR VPSRFTQMVG GRRARTFSI HFERGSVERE WNQGGRLGVE PFAMMRRKMK NFPTVEDAEA FALSAGWKIN SSNPGNMRFN PRVGGNRREF K QRYEFAKD ...String: AGLAARPLHT SAPRLIFPFR RPFKSEPEDE TKIPDCYVNG KPIADGQGYQ AIGLAEHFER IRRNVNIRRR VPSRFTQMVG GRRARTFSI HFERGSVERE WNQGGRLGVE PFAMMRRKMK NFPTVEDAEA FALSAGWKIN SSNPGNMRFN PRVGGNRREF K QRYEFAKD SYMAKDSGKG AQYPQDPHSS GTRGTSSFQA KVTKRVVQQE |
+Macromolecule #84: NDUFS5
| Macromolecule | Name: NDUFS5 / type: protein_or_peptide / ID: 84 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Euglena gracilis (euglena) Euglena gracilis (euglena) |
| Molecular weight | Theoretical: 14.244042 KDa |
| Sequence | String: MDRYGKKDES TWWVLHPFHE KTYTEDLGKV STVEPSVCDE YIELFKRCAA KSLDPEAQCR SYQEDLKECR LGAKQRKFLS QYEFATNSF TVNEKNDFRN RWHETFFRGS LL(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK) (UNK)(UNK) |
+Macromolecule #85: NDUFS6
| Macromolecule | Name: NDUFS6 / type: protein_or_peptide / ID: 85 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Euglena gracilis (euglena) Euglena gracilis (euglena) |
| Molecular weight | Theoretical: 17.101748 KDa |
| Sequence | String: MATTSRLSKV TVNIMPASRA LGVKPHRHRK QWQMPEVSFL KDVFSDPLMT EQKITERFIK AEFGPNRIDV RRAVPPIVVP KTFKHAWCD GNWGNRSTHG HPRISIRMKA GKVSQCKYCE NKFVRDDFPA FFAKREQVAA EEKYHDED |
+Macromolecule #86: NDUFS7
| Macromolecule | Name: NDUFS7 / type: protein_or_peptide / ID: 86 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Euglena gracilis (euglena) Euglena gracilis (euglena) |
| Molecular weight | Theoretical: 22.837303 KDa |
| Sequence | String: PQVKLLATWN PYNDPSLESV LQGDRKQTHM QRYAERQTLL GEELTPGQTH IRPGDNSVSW VSTTVSQVFK WGRSGSVWPL TFGLACCAI EMMQAYASRY DLDRYGTVPR GSPRQSDLII VAGTLTNKMA PCLRKVYDQM ATPCYVISMG SCANGGGYYH F SYAVVRGC ...String: PQVKLLATWN PYNDPSLESV LQGDRKQTHM QRYAERQTLL GEELTPGQTH IRPGDNSVSW VSTTVSQVFK WGRSGSVWPL TFGLACCAI EMMQAYASRY DLDRYGTVPR GSPRQSDLII VAGTLTNKMA PCLRKVYDQM ATPCYVISMG SCANGGGYYH F SYAVVRGC NRVVPVDIYV PGCPPTAEAL VYGILLLQRK IKMG(UNK)(UNK)(UNK)(UNK)(UNK) |
+Macromolecule #87: NDUFS8
| Macromolecule | Name: NDUFS8 / type: protein_or_peptide / ID: 87 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Euglena gracilis (euglena) Euglena gracilis (euglena) |
| Molecular weight | Theoretical: 24.370025 KDa |
| Sequence | String: MQHLKLARRG LPLQQLRSFH WTAPPLLRQP HPDVPKDVSG LGMGLAFMNQ FLLKEIMKAM WIVTIYTFRQ PSVTIHYPYE RNTKSTRFR GEHALLIYPD GDERCITCQL CEVTCPAQAI AIDGVEDDDG SRMASRFDLD MHKCIYCGLC QEACPVDAIV E TPHAEFCS ...String: MQHLKLARRG LPLQQLRSFH WTAPPLLRQP HPDVPKDVSG LGMGLAFMNQ FLLKEIMKAM WIVTIYTFRQ PSVTIHYPYE RNTKSTRFR GEHALLIYPD GDERCITCQL CEVTCPAQAI AIDGVEDDDG SRMASRFDLD MHKCIYCGLC QEACPVDAIV E TPHAEFCS TEYDSLLYDK ARLLENGIRW TSAIEYMVAK ERSDFPTNSQ RFKS |
+Macromolecule #88: NDUFV1
| Macromolecule | Name: NDUFV1 / type: protein_or_peptide / ID: 88 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Euglena gracilis (euglena) Euglena gracilis (euglena) |
| Molecular weight | Theoretical: 57.940191 KDa |
| Sequence | String: MLLRGVRCGA LAGKQRRSII QVPDKAPKTP AGFLQDKDRI FQNIYNDHGQ GLKNAQKLGD WYRTKDLLAK GPEWIIDQIK KSGLRGRGG AGFPSGLKWS FMPKKPPADG RPSYVVINAD ESEPGTCKDR EIMRMEPHKL VEGTLIAGFA MRARAAYIYV R GEFYNETL ...String: MLLRGVRCGA LAGKQRRSII QVPDKAPKTP AGFLQDKDRI FQNIYNDHGQ GLKNAQKLGD WYRTKDLLAK GPEWIIDQIK KSGLRGRGG AGFPSGLKWS FMPKKPPADG RPSYVVINAD ESEPGTCKDR EIMRMEPHKL VEGTLIAGFA MRARAAYIYV R GEFYNETL SVEKAIKEAY DAGLLGDNAC GSGYKFDIVL CRGAGAYICG EETGLIESLE GKSGKPRLKP PFPANIGLWG CP TIVTNVE TVSVSPTILR RGPEWFASFG RKNNSGTKLF CISGHVNTPC TVEESMSIPL RELLEKHCGG VRGGWDNLLA VIP GGSSVP VLPKSICDDV LMDFDALMDE GSALGTAAVI VMDKSTDLIK ALARISMFYK NESCNQCTPC REGAGVMANI MDRI WKGNG FGPEVDMIKD IADNVELRSI CALGAAAAWP IQGLHKHFRE EMIQRIEDFT DISYNLHKDE MFGHGAAFDW RRTPK TFHG TLDSFQDSTY RPPAANNNAY RGQYPHVPIY EKEKSEADKI AA |
+Macromolecule #89: NDUFV2
| Macromolecule | Name: NDUFV2 / type: protein_or_peptide / ID: 89 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Euglena gracilis (euglena) Euglena gracilis (euglena) |
| Molecular weight | Theoretical: 25.189109 KDa |
| Sequence | String: (UNK)(UNK)(UNK)(UNK)HIEWHN VRNAASDPFD FLPELYPEVY KILAKFPETK QQSAIIPLLH LAQRQNGGWV PLPG MLKIA EICEVPQKRV FECATFYCMF NHKPVGKYHI QVCVTTPCMI RGADEILHGL CHHLGVGEEE VTADGLFSIC EMECM GACV HAPMICISDY ...String: (UNK)(UNK)(UNK)(UNK)HIEWHN VRNAASDPFD FLPELYPEVY KILAKFPETK QQSAIIPLLH LAQRQNGGWV PLPG MLKIA EICEVPQKRV FECATFYCMF NHKPVGKYHI QVCVTTPCMI RGADEILHGL CHHLGVGEEE VTADGLFSIC EMECM GACV HAPMICISDY SNPPNYRYDY IEDLTLEQAI EVVDNLKKGI YPKVGSQQGR ASAIPLAGRT SLFDPPRGPF CRDLDK A |
+Macromolecule #90: poly(UNK)
| Macromolecule | Name: poly(UNK) / type: protein_or_peptide / ID: 90 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Euglena gracilis (euglena) Euglena gracilis (euglena) |
| Molecular weight | Theoretical: 1.039273 KDa |
| Sequence | String: (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK) |
+Macromolecule #91: FE2/S2 (INORGANIC) CLUSTER
| Macromolecule | Name: FE2/S2 (INORGANIC) CLUSTER / type: ligand / ID: 91 / Number of copies: 4 / Formula: FES |
|---|---|
| Molecular weight | Theoretical: 175.82 Da |
| Chemical component information |  ChemComp-FES: |
+Macromolecule #92: IRON/SULFUR CLUSTER
| Macromolecule | Name: IRON/SULFUR CLUSTER / type: ligand / ID: 92 / Number of copies: 6 / Formula: SF4 |
|---|---|
| Molecular weight | Theoretical: 351.64 Da |
| Chemical component information |  ChemComp-FS1: |
+Macromolecule #93: POTASSIUM ION
| Macromolecule | Name: POTASSIUM ION / type: ligand / ID: 93 / Number of copies: 1 / Formula: K |
|---|---|
| Molecular weight | Theoretical: 39.098 Da |
+Macromolecule #94: 1,2-DIACYL-SN-GLYCERO-3-PHOSPHOCHOLINE
| Macromolecule | Name: 1,2-DIACYL-SN-GLYCERO-3-PHOSPHOCHOLINE / type: ligand / ID: 94 / Number of copies: 41 / Formula: PC1 |
|---|---|
| Molecular weight | Theoretical: 790.145 Da |
| Chemical component information |  ChemComp-PC1: |
+Macromolecule #95: 1,2-Distearoyl-sn-glycerophosphoethanolamine
| Macromolecule | Name: 1,2-Distearoyl-sn-glycerophosphoethanolamine / type: ligand / ID: 95 / Number of copies: 9 / Formula: 3PE |
|---|---|
| Molecular weight | Theoretical: 748.065 Da |
| Chemical component information |  ChemComp-3PE: |
+Macromolecule #96: O-[(S)-hydroxy{[(2S)-2-hydroxy-3-(octadec-9-enoyloxy)propyl]oxy}p...
| Macromolecule | Name: O-[(S)-hydroxy{[(2S)-2-hydroxy-3-(octadec-9-enoyloxy)propyl]oxy}phosphoryl]-L-serine type: ligand / ID: 96 / Number of copies: 1 / Formula: S12 |
|---|---|
| Molecular weight | Theoretical: 523.597 Da |
| Chemical component information | 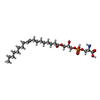 ChemComp-S12: |
+Macromolecule #97: CARDIOLIPIN
| Macromolecule | Name: CARDIOLIPIN / type: ligand / ID: 97 / Number of copies: 31 / Formula: CDL |
|---|---|
| Molecular weight | Theoretical: 1.464043 KDa |
| Chemical component information |  ChemComp-CDL: |
+Macromolecule #98: ZINC ION
| Macromolecule | Name: ZINC ION / type: ligand / ID: 98 / Number of copies: 3 / Formula: ZN |
|---|---|
| Molecular weight | Theoretical: 65.409 Da |
+Macromolecule #99: NADPH DIHYDRO-NICOTINAMIDE-ADENINE-DINUCLEOTIDE PHOSPHATE
| Macromolecule | Name: NADPH DIHYDRO-NICOTINAMIDE-ADENINE-DINUCLEOTIDE PHOSPHATE type: ligand / ID: 99 / Number of copies: 1 / Formula: NDP |
|---|---|
| Molecular weight | Theoretical: 745.421 Da |
| Chemical component information |  ChemComp-NDP: |
+Macromolecule #100: S-[2-({N-[(2S)-2-hydroxy-3,3-dimethyl-4-(phosphonooxy)butanoyl]-b...
| Macromolecule | Name: S-[2-({N-[(2S)-2-hydroxy-3,3-dimethyl-4-(phosphonooxy)butanoyl]-beta-alanyl}amino)ethyl] tetradecanethioate type: ligand / ID: 100 / Number of copies: 2 / Formula: ZMP |
|---|---|
| Molecular weight | Theoretical: 568.704 Da |
| Chemical component information |  ChemComp-ZMP: |
+Macromolecule #101: HEME-A
| Macromolecule | Name: HEME-A / type: ligand / ID: 101 / Number of copies: 2 / Formula: HEA |
|---|---|
| Molecular weight | Theoretical: 852.837 Da |
| Chemical component information |  ChemComp-HEA: |
+Macromolecule #102: COPPER (II) ION
| Macromolecule | Name: COPPER (II) ION / type: ligand / ID: 102 / Number of copies: 3 / Formula: CU |
|---|---|
| Molecular weight | Theoretical: 63.546 Da |
| Chemical component information |  ChemComp-CU: |
+Macromolecule #103: MAGNESIUM ION
| Macromolecule | Name: MAGNESIUM ION / type: ligand / ID: 103 / Number of copies: 1 / Formula: MG |
|---|---|
| Molecular weight | Theoretical: 24.305 Da |
+Macromolecule #104: 1,2-DILAUROYL-SN-GLYCERO-3-PHOSPHATE
| Macromolecule | Name: 1,2-DILAUROYL-SN-GLYCERO-3-PHOSPHATE / type: ligand / ID: 104 / Number of copies: 3 / Formula: PX2 |
|---|---|
| Molecular weight | Theoretical: 535.671 Da |
| Chemical component information |  ChemComp-PX2: |
+Macromolecule #105: UBIQUINONE-10
| Macromolecule | Name: UBIQUINONE-10 / type: ligand / ID: 105 / Number of copies: 1 / Formula: U10 |
|---|---|
| Molecular weight | Theoretical: 863.343 Da |
| Chemical component information |  ChemComp-U10: |
+Macromolecule #106: PROTOPORPHYRIN IX CONTAINING FE
| Macromolecule | Name: PROTOPORPHYRIN IX CONTAINING FE / type: ligand / ID: 106 / Number of copies: 4 / Formula: HEM |
|---|---|
| Molecular weight | Theoretical: 616.487 Da |
| Chemical component information |  ChemComp-HEM: |
+Macromolecule #107: HEME C
| Macromolecule | Name: HEME C / type: ligand / ID: 107 / Number of copies: 2 / Formula: HEC |
|---|---|
| Molecular weight | Theoretical: 618.503 Da |
| Chemical component information |  ChemComp-HEC: |
+Macromolecule #108: FLAVIN MONONUCLEOTIDE
| Macromolecule | Name: FLAVIN MONONUCLEOTIDE / type: ligand / ID: 108 / Number of copies: 1 / Formula: FMN |
|---|---|
| Molecular weight | Theoretical: 456.344 Da |
| Chemical component information |  ChemComp-FMN: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.3 mg/mL | ||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 7.4 Component:
Details: SEC buffer (30 mM Tris pH 7.4, 100 mM NaCl, 0.002% PMSF, 0.1% GDN (w/v), 1mM EDTA) | ||||||||||||||||||
| Grid | Model: Quantifoil R1.2/1.3 / Material: COPPER / Mesh: 300 / Support film - Material: CARBON / Support film - topology: CONTINUOUS / Support film - Film thickness: 2 / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 15 sec. / Pretreatment - Atmosphere: AIR / Pretreatment - Pressure: 0.039 kPa / Details: 15mA | ||||||||||||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277.15 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: FEI FALCON IV (4k x 4k) / Digitization - Dimensions - Width: 4096 pixel / Digitization - Dimensions - Height: 4096 pixel / Number grids imaged: 1 / Number real images: 9807 / Average exposure time: 7.8 sec. / Average electron dose: 51.5 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 1.8 µm / Nominal defocus min: 0.6 µm / Nominal magnification: 105000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Refinement | Space: REAL / Protocol: RIGID BODY FIT |
|---|---|
| Output model |  PDB-8iuf: |
+ About Yorodumi
About Yorodumi
-News
-Feb 9, 2022. New format data for meta-information of EMDB entries
New format data for meta-information of EMDB entries
- Version 3 of the EMDB header file is now the official format.
- The previous official version 1.9 will be removed from the archive.
Related info.: EMDB header
EMDB header
External links: wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
-Aug 12, 2020. Covid-19 info
Covid-19 info

URL: https://pdbj.org/emnavi/covid19.php
New page: Covid-19 featured information page in EM Navigator.
Related info.: Covid-19 info /
Covid-19 info /  Mar 5, 2020. Novel coronavirus structure data
Mar 5, 2020. Novel coronavirus structure data
+Mar 5, 2020. Novel coronavirus structure data
Novel coronavirus structure data
- International Committee on Taxonomy of Viruses (ICTV) defined the short name of the 2019 coronavirus as "SARS-CoV-2".
- In the structure databanks used in Yorodumi, some data are registered as the other names, "COVID-19 virus" and "2019-nCoV". Here are the details of the virus and the list of structure data.
Related info.: Yorodumi Speices /
Yorodumi Speices /  Aug 12, 2020. Covid-19 info
Aug 12, 2020. Covid-19 info
External links: COVID-19 featured content - PDBj /
COVID-19 featured content - PDBj /  Molecule of the Month (242):Coronavirus Proteases
Molecule of the Month (242):Coronavirus Proteases
+Jan 31, 2019. EMDB accession codes are about to change! (news from PDBe EMDB page)
EMDB accession codes are about to change! (news from PDBe EMDB page)
- The allocation of 4 digits for EMDB accession codes will soon come to an end. Whilst these codes will remain in use, new EMDB accession codes will include an additional digit and will expand incrementally as the available range of codes is exhausted. The current 4-digit format prefixed with “EMD-” (i.e. EMD-XXXX) will advance to a 5-digit format (i.e. EMD-XXXXX), and so on. It is currently estimated that the 4-digit codes will be depleted around Spring 2019, at which point the 5-digit format will come into force.
- The EM Navigator/Yorodumi systems omit the EMD- prefix.
Related info.: Q: What is EMD? /
Q: What is EMD? /  ID/Accession-code notation in Yorodumi/EM Navigator
ID/Accession-code notation in Yorodumi/EM Navigator
External links: EMDB Accession Codes are Changing Soon! /
EMDB Accession Codes are Changing Soon! /  Contact to PDBj
Contact to PDBj
+Jul 12, 2017. Major update of PDB
Major update of PDB
- wwPDB released updated PDB data conforming to the new PDBx/mmCIF dictionary.
- This is a major update changing the version number from 4 to 5, and with Remediation, in which all the entries are updated.
- In this update, many items about electron microscopy experimental information are reorganized (e.g. em_software).
- Now, EM Navigator and Yorodumi are based on the updated data.
External links: wwPDB Remediation /
wwPDB Remediation /  Enriched Model Files Conforming to OneDep Data Standards Now Available in the PDB FTP Archive
Enriched Model Files Conforming to OneDep Data Standards Now Available in the PDB FTP Archive
-Yorodumi
Thousand views of thousand structures
- Yorodumi is a browser for structure data from EMDB, PDB, SASBDB, etc.
- This page is also the successor to EM Navigator detail page, and also detail information page/front-end page for Omokage search.
- The word "yorodu" (or yorozu) is an old Japanese word meaning "ten thousand". "mi" (miru) is to see.
Related info.: EMDB /
EMDB /  PDB /
PDB /  SASBDB /
SASBDB /  Comparison of 3 databanks /
Comparison of 3 databanks /  Yorodumi Search /
Yorodumi Search /  Aug 31, 2016. New EM Navigator & Yorodumi /
Aug 31, 2016. New EM Navigator & Yorodumi /  Yorodumi Papers /
Yorodumi Papers /  Jmol/JSmol /
Jmol/JSmol /  Function and homology information /
Function and homology information /  Changes in new EM Navigator and Yorodumi
Changes in new EM Navigator and Yorodumi
 Movie
Movie Controller
Controller




































 X (Sec.)
X (Sec.) Y (Row.)
Y (Row.) Z (Col.)
Z (Col.)